Abstract
Newcastle disease virus (NDV) is not only one of the most economically important pathogen of poultry but also has a potential as anticancer virotherapy. The role of NDV V protein in virus-production kinetics was investigated using DF-1 cell-based production system. The presence of an anti-interferon (IFN)-alpha antibody resulted in enhanced NDV production kinetics in a dose-dependent manner by blocking binding of NDV-induced IFN to its receptor. To prepare DF-1 cell whose cellular IFN signaling is blocked efficiently, stable cell lines expressing either lentogenic or velogenic NDV V protein known as an IFN antagonist were established. The overexpression of NDV V protein enhanced NDV production kinetics and expedited the rate of NDV production, while it had no effect on Japanese encephalitis virus production. NDV V protein functions as an IFN antagonist by inhibiting the increase in type I IFNs by NDV infection. The IFN signals in cells expressing NDV V protein were weakened by decreased activation or expression of the dsRNA-activated enzymes. These IFN antagonist activities enhance rapid virus replication and spread in the early phase of viral infection and will be useful in improving the production of viral vaccine strains.
Similar content being viewed by others
Introduction
Newcastle disease (ND) is a contagious viral disease caused by specified viruses of the avian paramyxovirus type I (APMV-1) serotype of the genus Avulavirus belonging to the family Paramyxoviridae (Alexander 2008; Mayo 2002). The virus can infect most birds, including chickens, ducks, and doves. Since an outbreak and spread of virulent Newcastle disease virus (NDV) can cause massive economic losses in the poultry industry at a national level (Alexander 2000), many countries make great efforts to keep the virus under control.
NDV is group V virus according to the International Committee on Taxonomy of Viruses classification. Group V contains viruses possessing negative-sense single-stranded RNA (ssRNA) genomes (Malicki and Toka 1996). The size of the RNA genome is approximately 15,200 nucleotides. The RNA genome consists of six structural genes that code for nucleocapsid protein (NP), phosphoprotein (P), matrix protein (M), fusion protein (F), hemagglutinin-neuraminidase (HN), and an RNA-dependent RNA polymerase (L), respectively. The six genes are arranged in that order, from the 3′ terminus to the 5′ terminus on the genomic RNA (Kattenbelt et al. 2006). NDV strains are usually classified into three groups on the basis of their virulence: velogenic (high virulence), mesogenic (moderate virulence), and lentogenic (low virulence; Beard and Hanson 1984). The virulence of NDV depends on proteases specificity of the F protein cleavage site. The F proteins have multiple basic residues and F117 in the cleavage site is usually cleaved well by host intracellular protease(s) and showed higher virulence (Alexander 2008). The HN protein is related to the tropism of the virus. HN recognizes and binds to target receptors containing sialic acid on the host cell surface and releases the newly assembled virus particles from the infected cells (Fields and Knipe 2001; Huang et al. 2004). HN also affects the virulence of NDV by promoting the fusion activity of the F protein.
Two auxiliary proteins are coded from the P gene open reading frame (ORF) of NDV by RNA editing (Mebatsion et al. 2001). One is V protein, translated from a transcript in which one pseudotemplate G nucleotide is introduced at the RNA-editing site. The V protein shares a common N terminus with the phosphoprotein P, while it has unique C terminus containing seven highly conserved cysteine residues that binds two zinc atoms (Hagmaier et al. 2007). The NDV V protein is related with viral pathogenesis since it functions as an interferon (IFN) antagonist (Huang et al. 2003). The other protein is the W protein, which is translated from a transcript in which two pseudotemplate G nucleotides are present at the RNA-editing site. The introduction of two pseudotemplate G nucleotides into the mRNA is a rare event and the W ORF terminates in a stop codon shortly after the RNA-editing site (Fields and Knipe 2001). The functions and roles of W in viral pathogenesis and replication are not well understood.
Since the development of a recombinant NDV (Peeters et al. 1999), it has been possible to integrate mutated NDV genes or foreign genes into the NDV genome. This ability has accelerated NDV researches related to tropism, virulence, and the functions of viral genes (Engel-Herbert et al. 2003). Recombinant NDV containing foreign gene(s) may be useful for the development of a viral vector or a vaccine for other viruses. Glycoproteins from other viruses could be integrated efficiently into the envelope of NDV in recombinant NDV. For example, vaccines for severe acute respiratory syndrome (DiNapoli et al. 2007; Yang et al. 2004) and avian influenza (Ge et al. 2007) have been developed using recombinant NDV.
Additionally, because NDV is oncolytic, that is, the virus specifically infects and lyses cancer cells (Puhler et al. 2008; Sinkovics and Horvath 2000), NDV has received considerable attention in the fields of cancer research and drug development.
Although the NDV has many advantages, its production still depends on traditional egg-based production. However, in the egg-production system, the chances of contamination and the difficulties in quality control are disadvantageous. Recently, efforts to produce NDV using cell-based production system, which is safer and more effective than the egg-based production system, have been undertaken. For future applications in clinical settings and therapeutic drug development, a cell culture-based production system should be used to comply with the required Good Manufacturing Practice/Laboratory Practice regulations.
However, the productivity of NDV in cell-based production systems is relatively low and usually only reaches a titer of ∼106 PFU/ml, which is approximately 100-fold lower than that of the egg-based production system. To overcome this limitation, DF-1 host cells were genetically modified in this study. The well-known IFN antagonist, NDV V protein, was stably overexpressed in DF-1 cells to weaken the cellular innate immune system. Since the IFN system plays a major role in cellular innate immunity, especially at the early phase of infection, disruption of the IFN system should enhance NDV production in the cell-based production system.
Materials and methods
Cells and viruses
DF-1 (chicken embryo fibroblast; American Type Culture Collection [ATCC], CRL12203®, Manassas, VA, USA) and Vero (African green monkey kidney; ATCC, CCL-81®) cells were maintained in Minimum Essential Medium (MEM)-α (GIBCO®) supplemented with 10% and 5% fetal bovine serum (FBS; PAA Laboratories, Ontario, Canada), respectively. The lentogenic NDV strain, LaSota from DaeSung Microbiological Labs. Co., Ltd. (Uiwang, South Korea) was propagated in DF-1 cells with basal MEM-α containing 400 ng/ml trypsin. The Japanese encephalitis virus (JEV) Beijing1 strain was provided by Boryung Pharmaceutical Co., Ltd. (Seoul, South Korea). The RNA genome of a velogenic NDV strain, Kr-102/89, was obtained from the Korean National Veterinary Research and Quarantine Service.
Nucleotide sequence accession numbers
The GenBank accession numbers for the V gene of the LaSota and Kr-102/89 strains are AF077761 and FJ534731, respectively.
Construction of V gene expression plasmids
The LaSota RNA genome was extracted using a NucleoSpin® RNA Virus kit (Macherey-Nagel GmbH, Duren, Germany). The P gene complementary DNAs (cDNAs) of the LaSota and Kr-102/89 strains were prepared using M-MLV Reverse Transcriptase (iNtRON Biotechnology, Seongnam, Korea) with the LaSota P-S primer, 5′-GGGAATTCATGGCCAC CTTTACAGATGCAG-3′, and the Kr-102/89 P-S primer, 5′-GAGAATTCATGGCCACCTTTA CAGATGCTG-3′, respectively.
The P gene ORFs of the LaSota and Kr-102/89 strains were prepared by polymerase chain reaction (PCR) using the following primers: LaSota P-S, LaSota P-AS, 5′-AGTCTAGATTAGCCATTTAGAGCAAGGCG-3′; Kr-102/89 P-S, Kr-102/89 P-AS, 5′-GATCTAGATCAACCATTCAGCGCAAGGCG-3′. The prepared P gene ORFs were cloned into the mammalian cell expression vector pcDNA3.1(+) (Invitrogen, CA, USA).
The V gene ORF of the LaSota strain was prepared by two-step PCR using the LaSota P gene ORF as a template and the following primers: LaSota P-S, LaSota V-Ne-AS, 5′-GGCTCGA CCATGGGCCCCTTTTTAGCATTG-3′ for the N-terminal region and LaSota V-Ne-S, 5′-CAATGCTAAAAAGGGGCCCATGGTCGAGCC-3′, LaSota V-AS, 5′-AGTCTAGAT TACTTACTCTCTGTGATATCGCCTCC-3′ for the C-terminal region. In the first-round PCR, the N-terminal and C-terminal regions were amplified. In the second-round PCR, the full-length V gene ORF was amplified using the N-terminal and C-terminal fragments as templates and the LaSota P-S, LaSota V-AS primers (nontemplate, additional G and C are italicized; Fig. 2). The V gene ORF of the Kr-102/89 strain was prepared using the same method as the LaSota strain with the following primers: Kr-102/89 P-S, Kr-102/89 V-Ne-AS, 5′-GCTCGACCGTGGCCCCCTTTTTAGCATTAG-3′; Kr-102/89 V-Ne-S, 5′-CTAATGCTAAAAAGGGGGCCACGGTCGAGC-3′; and Kr-102/89 V-AS, 5′-GATCTAGATCAACCATTCAGCGCAAGGCG-3′. The N-terminal FLAG-tagging plasmid, pcDNA3.1(+)-FLAG-4T-1 was kindly provided by Hyangshuk Rhim, the Catholic University Medical College of Korea. The V gene ORFs were cloned into pcDNA3.1(+)-FLAG-4T-1. All constructs were confirmed by sequencing.
Transfection and generation of stable cell lines
One day before transfection, DF-1 cells were seeded in 12-well plates at a density of 2.0 × 105 cells per well. Cells were transfected with 1.6 µg of plasmid using 4 µl Lipofectamine2000™ (Invitrogen). One day after transfection, the medium was replaced with culture medium containing 2.5 mg/ml G418 (Duchefa, Haarlem, The Netherlands). Stable cell lines expressing NDV V protein were maintained in culture medium containing 600 µg/ml of G418. The expression levels of FLAG-tagged V protein in stable cell lines were examined by Western blotting analysis. The stable cells were lysed by RIPA buffer [50 mM Tris–HCl (pH 7.6), 150 mM NaCl, 1% (v/v) NP-40, 1% (w/v) sodium deoxycholate, 0.1% (w/v) sodium dodecyl sulfate (SDS), 1 mM phenylmethanesulphonylfluoride, 1 mM Na3VO4, 1 mM NaF, 0.15 µM aprotinin, 1 µM leupeptin, and 1 µM pepstatin] for Western blotting analysis. Western blotting was performed according to the instructions provided in the Millipore protein blotting handbook. The FLAG-tagged V protein and actin were detected by using a monoclonal antibody against FLAG tag (A4977) purchased from Sigma-Aldrich (Saint Louis, MO, USA) and a monoclonal antibody against actin (sc-1616) purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA), respectively.
NDV production kinetics
Control cells or DF-1 cells expressing LaSota V or Kr-102/89 V protein were seeded in six-well plates at a density of 2.5 × 105 cells per well. Two days later, the cell monolayers were infected with the NDV LaSota strain at a multiplicity of infection (MOI) of 0.1 and incubated at 37°C in virus-production medium (basal MEM-α containing 400 ng/ml trypsin). Supernatants were harvested daily, and virus titers were determined by plaque assays on Vero cells, as described previously (Kournikakis and Fildes 1988; Harper 1989). Briefly, Vero cells were seeded in 12-well plates at a density of 2.0 × 105 cells per well. The following day, monolayers of Vero cells were infected with 50 µl of serial tenfold dilutions of the virus samples. One hour after adsorption, the cells were covered with a basal MEM-α agar overlay containing 0.02% (w/v) DEAE-dextran, 0.1% (w/v) glucose, 0.7% (w/v) bacto agar, 30 mM MgSO4, and 4 µg/ml trypsin. The cells were incubated at 37°C for 3 days. Two days after virus infection, a 0.03% (w/v) neutral red agar overlay was added to each well to stain viable cells.
JEV production kinetics
Cells were prepared by the same method used above. The cell monolayers were infected with the Beijing1 strain of JEV at a MOI of 0.1 and incubated at 37°C in basal MEM-α. Supernatants were harvested daily, and virus titers were determined by plaque assays on Vero cells, as described previously (Huang et al. 2003). Briefly, Vero cells were seeded in 12-well plates at a density of 2.0 × 105 cells per well. The following day, monolayers of Vero cells were infected with 50 µl of serial tenfold dilutions of the virus samples. One hour after adsorption, the cells were covered with a basal MEM-α overlay containing 0.8% (w/v) methylcellulose, 5% FBS, 2 mM glutamine, 0.225% (w/v) sodium bicarbonate (NaHCO3), and 10 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid. The cells were incubated at 37°C for 3 days. The cells were then fixed with 5% (v/v) formaldehyde and stained with 1% (w/v) crystal violet solution for counting plaques.
Measurement of viable cell density
The viable cell density during virus production was measured by trypan blue exclusion. A 0.4% (w/v) trypan blue solution was added to cell suspensions and the cells were counted using a hemocytometer (Marienfeld, Germany) by light microscopy (Olympus CK40, Japan).
Observations of syncytia formation by the infected cells
To observe the cell morphology and formation of syncytia, Giemsa staining was performed. The cells were fixed with absolute methanol for 5 min and stained with 0.04% (w/v) Giemsa in phosphate buffer for 30 min. The cells were washed with tap water and then observed by light microscopy (Olympus CKX41, Japan).
NDV LaSota production in cell culture-based production system to examine integration of V protein into NDV particles
Control cells or DF-1 cells expressing LaSota V or Kr-102/89 V protein were seeded in six-well plates at a density of 2.5 × 105 cells per well. Two days later, the viable cell density was measured by trypan blue method. Then, cell monolayers were infected with the NDV LaSota strain at a MOI of 0.1 in virus-production medium and incubated at 37°C. Two days later, supernatants were harvested and then centrifuged at 16,000×g for 10 min at 4°C to remove cellular debris. NDV LaSota in the supernatants was resolved by SDS–polyacrylamide gel electrophoresis. The FLAG-tagged NDV V protein and NDV F protein were detected by using a monoclonal antibody against FLAG tag and a polyclonal antibody against the NDV LaSota strain prepared from rabbit (AbFRONTIER, Seoul, Korea), respectively.
RNA extraction from the cells and real-time RT-PCR
Cells were grown to 80–90% confluence in a 100-mm culture dish. Cells were infected with the LaSota NDV at a MOI of 3 in basal MEM-α containing 400 ng/ml trypsin. The cells were harvested 24 h later. Total RNA was extracted from the cells using an RNeasy® Mini kit (Qiagen GmbH, Hilden, Germany), and 3 µg of total RNA was used as a template for synthesizing cDNA strands. Reverse transcription was performed using an oligo(dT)20 primer (Invitrogen, USA) with the M-MLV reverse transcriptase (iNtRON Biotechnology). Real-time PCR was performed on a LightCycler® 480 (Roche Applied Science, Mannheim, Germany). Each reaction was performed with 100 ng cDNA, 0.5 µM primers (Table 1), 5% (v/v) dimethyl sulfoxide, and SYBR® Premix Ex Taq™ II (TaKaRa, Japan). Real-time PCR conditions were 1 min at 95°C for hot-start enzyme activation, and 45 cycles of 95°C denaturation for 10 s, 65°C annealing for 10 s, and 72°C extension for 10 s. The specificity of amplification was determined by melting curve analysis which consisted of 95°C denaturation for 1 min, 65°C annealing for 5 min, and heating to 95°C at the rate of 0.11°C/s. The T m and C p values were calculated using the analysis software provided by Roche Applied Science. Quantitative analysis on mRNA levels of type I and type II IFN genes were normalized by internal control, actin gene (Table 1).
Investigation of proteins related to IFN signaling
For investigating the expression levels of proteins related with IFN signaling, control cells and cells expressing LaSota V or Kr-102/89 V protein were infected with the NDV LaSota or JEV at a MOI of 3. At 24 h after infection, the cells were lysed by RIPA buffer for Western blotting analysis. The STAT proteins were detected by using anti-STAT1 (#9172) antibody purchased from Cell Signaling Technology (Boston, MA, USA) and an anti-STAT2 antibody (sc-476, Santa Cruz Biotechnology). The IFN-simulated gene products were detected by using anti-MxA (sc-50509), anti-OAS3 (sc-49870, Santa Cruz Biotechnology), and anti-phospho-eIF2α (#9171, Cell Signaling Technology) antibodies.
Statistical analysis
All statistical analyses were performed with the MINITAB® Release 14 software. All null hypotheses (H0) were tested using a 99% confidence interval (significance level, α = 0.01) unless noted otherwise.
Results
Blocking of IFN-α action enhances NDV production kinetics
Chicken IFN-α has 20-fold higher specific activity than IFN-β and is the main type I IFN produced by chicken embryonic fibroblast (Mo et al. 2001). To investigate the effect of blocking type I IFN activity on virus-production kinetics, an anti-IFN-α antibody was added to the virus-production medium. The antibody binding to IFN-α secreted from virus-infected cells may block the action of IFN-α by preventing IFN-α binding to the type I IFN receptors, IFNAR1 and IFNAR2. The presence of the antibody in the virus-production medium enhanced virus-production kinetics in a dose-dependent manner (Fig. 1a). At 2 and 3 days postinfection, the virus titer produced in the presence of 50 ng/ml of the antibody was threefold higher than that produced in the absence of the antibody. In contrast, the viable cell density decreased in a dose-dependent manner during virus production (Fig 1b).
Effect of anti-IFN-α antibody on NDV production kinetics in DF-1 cell culture. a Virus titers increased in the presence of anti-IFN-α antibody because the antibody disrupts the innate immune responses of DF-1 cells by binding to secreted IFN-α. A monoclonal antibody against chicken IFN-α (sc-57226; Santa Cruz Biotechnology) was added to NDV production medium. Values are from two independent experiments, each performed in triplicate. Bars show standard deviations. b Giemsa staining of DF-1 cells during virus production 2 days after infection. The greater the level of anti-IFN-α antibody in the production media, the more syncytia and fewer viable cells were observed. Bars indicate 500 µm. *p < 0.01, virus titers are statistically different
Preparation of cell lines expressing an IFN antagonist
Since blocking the action of IFN enhanced the virus-production kinetics, stable cell lines constitutively expressing an IFN antagonist were prepared. The NDV V protein is a well-known IFN antagonist and the V gene ORF was prepared from NDV genomic RNA. To compare the IFN antagonistic activity and the influence on virulence of V proteins originating from lentogenic and velogenic strains, the V gene ORFs of the LaSota (lentogenic) and Kr-102/89 (velogenic) strains were used in this experiment.
The V gene ORF was transcribed by addition of a nontemplate G residue into the G run within the conserved RNA-editing site (5′-AAAAAGGG-3′) in the P gene ORF (Steward et al. 1993). In NDV-infected cells, the proportions of P, V, and W mRNAs are 68%, 28%, and 4%, respectively (Mebatsion et al. 2003). Since the three mRNAs are almost identical, with the exception of the changes at the editing site, and the P mRNA is the most abundant, the V gene ORF was prepared by two-step PCR from the P gene ORF (see the Materials and methods section; Fig. 2). First, the P gene ORF was amplified from the viral RNA genome. The N-terminal and C-terminal fragments of the V gene were then amplified with overlapping primers containing a pseudotemplate G in the editing site. The full-length V gene ORF was amplified using these overlapping N and C terminus fragments as templates.
Preparation of V gene ORFs. a Schematic overview of two-step PCR used to prepare V gene ORF. In the first-round PCR, N-terminal and C-terminal fragments of the V gene ORF were amplified using the P gene ORF as a template. In the second-round PCR, full-length V ORF was amplified using the first-round PCR products as templates. b The result of the first-round PCR: lane M 100 bp DNA ladder (Fermentas), lane N N-terminal fragment of V gene ORF, lane C C-terminal fragment of V gene ORF. c The result of the second-round PCR: lane V full-length V gene ORF
The prepared V gene ORFs were confirmed by sequencing (Supplementary Fig. 1a). The V protein sequences from the two virus strains showed a high degree of similarity, with 80.3% identity between the full-length constructs and 78.1% identity in the C-terminal region (Supplementary Fig. 1b). The C terminus of the V proteins have conserved tryptophan residues that are required for binding to STAT proteins and seven conserved cysteine residues required for binding to DDB1, which is required for proteasomal degradation of STAT proteins (Nishio et al. 2005; Supplementary Fig. 1c).
The V gene ORFs were then cloned into a mammalian cell expression vector. It was previously reported that the tagging of the N terminus of paramyxovirus V protein with hemagglutinin or FLAG had no effect on the IFN antagonistic activity of V protein (Rodriguez et al. 2003; Ulane et al. 2005). Even the truncated version of NDV V, protein which has only the C-terminal region, showed the IFN antagonistic activity (Park et al. 2003). Therefore, the FLAG tag was added to the N terminus of NDV V protein in this study. The prepared stable cell lines were screened for expression levels of the FLAG-tagged V protein (Fig. 3). The cell lines exhibiting stable expression of V protein and normal cell physiology were selected for further experiments. These cell lines showed the stable expression of V protein during the whole period of virus production (Supplementary Fig. 2).
Screening of IFN antagonist-expressing cell lines. Total cell lysates were analyzed by Western blot with antibodies against the FLAG tag and actin. a Stable DF-1 cell lines expressing LaSota V protein. b Stable DF-1 cell lines expressing Kr-102/89 V protein. Lane M mock control, lanes 1–6 individual stable cell lines expressing either NDV LaSota V protein or Kr-102/89 V protein
NDV V protein-expressing cell lines show the enhanced LaSota production kinetics in DF-1 cell
To investigate the effect of the IFN antagonistic activity on virus production, virus-production kinetics was examined using a plaque assay. The virus-production kinetics was enhanced in cells expressing either LaSota or Kr-102/89 V protein (Fig. 4a). At 2 days postinfection, the virus titers of cells expressing NDV V protein were approximately tenfold higher than that of the control cell (Fig. 4c, h). The viable cell densities during the virus-production period were also examined. The viable cell densities decreased much more rapidly in cells expressing NDV V protein (Fig. 4b) compared to the control cell. By approximately 1 day postinfection, cells expressing NDV V protein began to form syncytia. The syncytia were widespread by 36 h postinfection (Fig. 4d). The maximum virus titers of cells expressing NDV V protein were almost twice that of control cell (Fig. 4i).
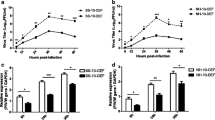
Expression of the IFN antagonist V protein enhances NDV production kinetics. a LaSota production kinetics between mock and IFN antagonist-expressing cell lines. b Viable cell densities during NDV production in mock and IFN antagonist-expressing cell lines. Values are from three independent experiments, each performed in duplicate. c Plaque formation of NDV LaSota produced from mock or IFN antagonist-expressing cell lines. Two days after infection, virus-production medium was diluted 10−3 and then used to inoculate Vero cell monolayers. Medium from IFN antagonist-expressing cell lines showed much higher virus titers. d Giemsa staining of the cells during virus production. At 40 h postinfection, IFN antagonist-expressing cell lines had lower visible cell densities and massive syncytia formation. Bars = 200 µm. e JEV production kinetics between mock and IFN antagonist-expressing cell lines. f Viable cell densities during JEV production in mock and IFN antagonist-expressing cell lines. Values are from three independent experiments, each performed in duplicate. g Giemsa staining of JEV-infected cells during virus production (40 h postinfection). Bars = 200 µm. h Relative virus titers during the virus-production period. The values were calculated by setting the titers of the mock control to 1. i Maximum virus titers. In LaSota production, virus titers from IFN antagonist-expressing cell lines were approximately twofold higher than the mock controls. *p < 0.01, statistically different virus titers. Bars show standard deviations
To investigate whether the overexpression of NDV V protein had similar effects on the production of other viruses, the production kinetics of JEV were examined. Both control and cells expressing NDV V protein showed very similar JEV production kinetics (Fig. 4e). The viable cell densities during JEV production also showed similar curves, and there were no distinct morphological changes (Fig. 4f, g). The relative virus titers during the production period and the maximum virus titers were very similar for both control and cells expressing NDV V protein (Fig. 4h, i). Therefore, the overexpression of NDV V protein had no effect on JEV production kinetics in the DF-1 cell.
NDV V proteins disrupt the cellular innate immune response by inhibiting the expression of type I IFNs
To examine the mechanism by which the NDV V protein enhances NDV production kinetics, the expression level of IFNs and IFN signal-related proteins were investigated. In our other study, the expression of a certain NDV structural protein in DF-1 cell enhanced the NDV production kinetics (unpublished data). If NDV V protein acts just like a structural protein, there is a possibility that NDV V protein enhances virus-production kinetics as a structural protein does irrespective of its IFN antagonistic activity. Therefore, we tested whether the NDV V protein can be incorporated into NDV particles as a structural protein. The presence of FLAG-tagged V protein in NDV produced from cells expressing NDV V protein was examined. The result showed that the FLAG-tagged V protein was not incorporated into NDV particles (Fig. 5a), indicating that the NDV V protein does not function as a virus structural protein.
NDV V proteins disrupt innate cellular immunity by inhibiting the expression of type I IFNs. a NDV V proteins are not incorporated into virus particles. NDV LaSota was produced from the indicated cell lines. The asterisk indicates the cleaved F protein due to the presence of trypsin in the production medium. b Relative mRNA expression levels of type I and type II IFNs. NDV V proteins inhibit the LaSota-induced expression of type I IFNs. The values were calculated by setting the expression level of type I IFNs in each cell line to 1. c IFN signaling in cell lines expressing NDV V proteins. Cells were infected with the NDV LaSota or JEV at 3 MOI. At 24 h postinfection, cell lysates were analyzed by Western blotting with antibodies against STAT proteins and IFN-induced antiviral effector proteins
To examine how the NDV V protein inhibits cellular IFN signaling, the expression levels of the type I and type II IFNs were measured. In uninfected cells, the type I and type II IFNs were expressed in similar levels, in both control and cells expressing NDV V protein. However, when cells were infected with the NDV, the cells expressing NDV V protein exhibited reduced levels of type I IFNs (Fig. 5b).
The activation and expression levels of proteins involved in IFN signaling were investigated. Many paramyxovirus V proteins target cellular STAT proteins to inhibit IFN signaling (Horvath 2004). The V proteins form a complex with STAT, DDB1, and Cullin4A and cause the degradation of STAT proteins via the proteasome (Andrejeva et al. 2002). However, here, the STAT proteins were not degraded in both control and cells expressing NDV V protein, even when infected with NDV (Fig. 5c). Then, IFN-induced antiviral effectors were investigated. The protein kinase R (PKR), a double-stranded RNA (dsRNA)-activated cellular protein kinase, phosphorylates eIF-2α whose phosphorylation leads to a block in translation initiation (Basler and Garcia-Sastre 2002). When DF-1 cells were infected with NDV, phosphorylation of eIF-2α was increased, indicating that PKR was activated. While cells expressing NDV V protein showed the decreased activation of PKR in the NDV-infected state. Unlike NDV infection, when cells were infected with JEV, PKR was not activated by JEV infection irrespective of the expression of NDV V protein. The expression level of another dsRNA-activated enzyme, 2′,5′-oligoadenylate synthetases (OAS), was also reduced in cells expressing NDV V protein in the NDV-infected or JEV-infected state. In contrast to the dsRNA-activated enzymes, the GTPase family, Mx proteins, which also possess well-described antiviral activities, showed similar expression levels in both control and cells expressing NDV V protein irrespective of virus infection (Fig. 5c).
Discussion
NDV has proven useful for developing viral vaccines and agents for cancer therapy. However, NDV production in a cell-based system shows relatively lower productivity and production kinetics than other viruses such as JEV.
To overcome this problem, we tried to prepare stable cell line for NDV production through primary culture of egg cell, but this approach failed because of difficulty in developing stable cell lines. Then, a wide range of commonly used cell lines for virus production, such as DF-1, Vero, MDCK, BHK, and so on, were tested for NDV production. Our experiments showed that the DF-1 cell was the best choice for NDV production (data not shown).
To comply with regulatory policies, the NDV LaSota, which is a representative lentogenic strain used for producing live-attenuated ND vaccines, was used in this study. Since the F protein of the lentogenic strain is not cleaved by host cellular proteases, the addition of extracellular protease is inevitable for producing the lentogenic NDV strain in cell culture-based system. Generally speaking, the amount of extracellular protease added plays an important role in optimizing the virus-production kinetics. If excessive protease is present in the production medium, it will destroy host cell monolayers and reduce the virus-production kinetics. On the other hand, insufficient protease is not able to support the rapid NDV production since it takes more time to cleave F protein of NDV newly produced from the host cells. In this study, trypsin, acetyltrypsin, and fresh allantoic fluid were tested as external protease sources. All proteases tested in this study showed that the virus-production kinetics were almost the same when the proteases were used with their optimized concentration (Supplementary Fig. 3a). Regardless of kinds of proteases, excessive or insufficient protease is not effective on virus production (Supplementary Fig. 3b, c).
To develop a cell line to support more efficient production of NDV, we tried to disrupt the antiviral cellular defense system. IFN is a key component of the innate immune system. It shows antiviral activities and is used broadly in virus research and in the treatment of viral infection. Therefore, a decrease in IFN activity may cause an increase in virus production.
Indeed, the addition of an antibody against IFN to neutralize secreted IFN-enhanced virus-production kinetics (Fig. 1). This suggests that preparation of stable cell lines which constantly express virus IFN antagonist would have an effect on enhancement of NDV production kinetics.
The paramyxoviruses have accessory proteins acting as IFN antagonists (Gotoh et al. 2001). The accessory proteins are transcribed by RNA editing such as NDV V gene. Generally, these accessory proteins interfere with IFN signaling by (1) suppressing tyrosine phosphorylation of STAT proteins, (2) causing proteasome-mediated degradation of STAT proteins, (3) decreasing the level of STAT proteins, (4) associating with STAT proteins, and (5) inhibiting the production of type I IFNs (Gotoh et al. 2001, 2002).
To prepare cell lines expressing the IFN antagonist, the NDV V protein was used as an IFN antagonist in this study. To compare the antagonistic activity between lentogenic and velogenic NDV strains, the V proteins of the LaSota and Kr-102/89 strains were used.
The cells expressing either LaSota or Kr-102/89 V protein showed enhanced virus-production kinetics, especially in the early phase of NDV infection (Fig. 4). These increases were directly related with the decreased type I IFNs response in cells expressing the NDV V protein (Fig. 5). The NDV V protein showed its IFN antagonistic activity by inhibiting the induction of type I IFNs caused by NDV infection.
Unexpectedly, STAT proteins are not degraded in the IFN signal pathway of the DF-1 cell. Both STAT1 and STAT2 were not degraded in either control or cells expressing the NDV V protein even when infected with NDV. However, IFN-stimulated enzymes showed decreased activity or expression: both the activity of PKR and the expression of OAS decreased in cells expressing V protein. The decreased activity or expression of IFN-stimulated genes is thought to result from the inhibition of NDV-induced type I IFNs.
The type I IFNs are expressed in response to pathogen infection such as virus or bacteria. The expression of type I IFNs would not be induced much without virus infection. Actually, the relative expression level of type I IFNs in cells expressing NDV V protein was about 0.5-fold to 0.7-fold compared to the control cell without NDV infection (data not shown). Therefore, the NDV V protein has the inherent ability of inhibiting type I IFNs expression. This ability reached a peak when NDV infected the host cell (Fig. 5b). The exact mechanism of how NDV V protein inhibits the induction of type I IFNs by NDV infection is not yet understood. According to the previous reports (Yoneyama et al. 2004; Childs et al. 2007, 2009), NDV V proteins may inhibit RNA helicases such as mda-5 and RIG-I which have an essential role in dsRNA-induced innate immunity containing type I IFNs production. These RNA helicases are well known for their binding to dsRNA caused by RNA virus infection and stimulation of host cellular defense genes expression. Since the inhibitory effect of the NDV V protein on type I IFNs reaches a peak in the NDV-infected state, the NDV V protein might influence the action of RNA helicase. In addition, the activation of PKR and expression of OAS in response to the presence of dsRNA (Basler and Garcia-Sastre 2002) are decreased in cells expressing NDV V protein while other IFN-stimulated genes, Mx proteins, are not affected.
In this study, both lentogenic and velogenic NDV V proteins have the same IFN antagonistic effect although the expression levels of NDV V protein were somewhat different.
Because NDV expresses the NP gene from the 3′ terminus prior to the expression of P gene and V gene is expressed from the P gene ORF by RNA editing, it takes time to express V gene from the NDV genome. Therefore, the presence of NDV V protein in the host cell before virus infection, despite of its small amount, can expedite NDV replication at the early infection stage by saving time to express NDV V protein from the NDV genome and efficiently inhibiting host cellular innate immunity related to type I IFNs. In addition, NDV can spread more rapidly and efficiently through cell-to-cell fusion using the F protein. In the early phase of infection, the host cellular innate immune responses reduced by the overexpression of NDV V protein accelerate NDV replication and transcription of the F gene. The rapid increase in F protein accelerates cell fusion, making continued replication and transcription of NDV much easier. These synergistic effects enhance NDV productivity in cells expressing NDV V protein, especially early in the infection phase.
Unlike those previously reported (Huang et al. 2003), both STAT1 and STAT2 were not degraded by NDV V protein in DF-1 cell, which might be caused by the use of different cell lines. A certain cell may experience degradation of either STAT1 or STAT2, or both, or neither of them by a certain paramyxovirus V protein (Nishio et al. 2005). The degradation of STAT protein is determined by its corresponding binding to paramyxovirus V protein, which occurred in a cell line-specific manner. Huang used 2fTGH (human fibroblast) and Vero (monkey kidney) cells for his study, while our study used DF-1 (chicken embryonic fibroblast) cell.
Some recent reports have also suggested that paramyxovirus V proteins inhibit IFN signal without degradation of STAT proteins. Recombinant hPIV2 with a mutated V protein that cannot bind to STAT proteins showed similar virus growth kinetics in some cell lines (Nishio et al. 2005). This result suggests that the IFN antagonist activity of the V protein is mediated through mechanisms other than STAT protein degradation. V proteins of the measles, mumps, and Mapuera viruses appear to inhibit IFN without degradation of STAT proteins. In these viruses, V proteins inhibit the phosphorylation of STAT proteins by Jak1 or Tyk2, resulting in the inhibition of STAT dimers translocation to the nucleus and induction of IFN-stimulated genes (Caignard et al. 2007; Kubota et al. 2005). Therefore, the degradation of STAT proteins is not necessary for antagonization of host cellular innate immunity.
Unlike NDV infection, the overexpression of NDV V protein has no effect on virus-production kinetics in JEV infection. The PKR was not activated and OAS was not expressed much (although cells expressing NDV V protein showed lower expression level of OAS compared to the control cell) in JEV-infected state. We assume that JEV have indigenous mechanisms, not understood clearly yet, for evasion of host cellular innate immunity. The mechanisms might use another pathway which is different from that of NDV and they seem to have better antagonist activity since the PKR was not activated even in the JEV-infected state.
JEV is a positive sense ssRNA virus, and its viral genome can act as mRNA without transcription. Thus, JEV has a less chance of forming dsRNA than negative-sense ssRNA virus does. The NDV, negative-sense ssRNA virus, forms dsRNA to transcribe its six structural genes. Additionally, when the P gene is transcribed, RNA editing occurs in order to transcribe the V and W mRNAs. Because RNA editing generates a stuttering of the viral RNA polymerase complex at the editing site, the delay in transcription extends the presence of dsRNA. Because dsRNA functions as an indicator of virus infection, JEV has lower possibility of being detected by host RNA helicases. Therefore, the host cell may recognize the NDV-infected state easier than the JEV-infected state. In addition, since all JEV proteins are produced in the form of polyprotein and completed by cleavage of host cellular protease, the IFN antagonist of JEV is produced more rapidly than that of NDV. For these reasons, NDV V protein has no or minor effect on JEV replication.
Although cells expressing NDV V protein only showed twice increase of maximum titer, their production rate was much faster than that of the control cell. This increase of virus-production rate resulted from the attenuated host cellular innate immunity. As both production yield and production rate are important parameters in practice, we believe that cells expressing the IFN antagonist can contribute to the expedition of the virus-production process. The cells expressing the IFN antagonist may result to better productivity when they are combined with other strategies for enhancing virus production.
According to recent research (de Vries et al. 2008), various virus IFN antagonists have a wide range of IFN antagonistic effects on the enhancement of various virus-production kinetics. So, it is necessary to evaluate the IFN antagonistic activity from experiments in order to find the best IFN antagonist for target virus production. Also as seen in JEV production from cells expressing NDV V protein, it may or may not have effects on a specific virus production, depending on IFN antagonist. Therefore, we think that it is better to use an indigenous IFN antagonist of the target virus first and then perform evaluations for the effects of other virus IFN antagonists on target virus production. This strategy can help in determining the best IFN antagonist for target virus production and developing the recombinant host cell lines.
In conclusion, the overexpression of NDV V protein can inhibit the induction of type I IFN genes and activation of IFN-stimulated genes in response to dsRNA. The decreased cellular innate immune response and cell-to-cell fusion activity of NDV act synergistically to enhance NDV production kinetics in a cell-based production system. This strategy for enhancing NDV production may accelerate the advance of fields related to the development of viral vector, vaccine, and cancer therapy using NDV through rapid and efficient NDV production.
References
Alexander DJ (2000) Newcastle disease and other avian paramyxoviruses. Rev Sci Tech 19:443–462
Alexander DJ (2008) Manual of diagnostic tests and vaccines for terrestrial animals. Office International des Epizooties, Paris
Andrejeva J, Poole E, Young DF, Goodbourn S, Randall RE (2002) The p127 subunit (DDB1) of the UV-DNA damage repair binding protein is essential for the targeted degradation of STAT1 by the V protein of the paramyxovirus simian virus 5. J Virol 76:11379–11386
Basler CF, Garcia-Sastre A (2002) Viruses and the type I interferon antiviral system: induction and evasion. Int Rev Immunol 21:305–337
Beard CW, Hanson RP (1984) Newcastle diseases. In: Hofstad MS (ed) Diseases of poultry, 8th edn. Iowa State University Press, Ames, pp 452–470
Caignard G, Guerbois M, Labernardiere JL, Jacob Y, Jones LM, Wild F, Tangy F, Vidalain PO, Infectious Mapping Project I-MAP (2007) Measles virus V protein blocks Jak1-mediated phosphorylation of STAT1 to escape IFN-alpha/beta signaling. Virology 368:351–362
Childs KS, Stock N, Ross C, Andrejeva J, Hilton L, Skinner M, Randall RE, Goodbourn S (2007) mda-5, but not RIG-I, is a common target for paramyxovirus V proteins. Virology 1:190–200
Childs KS, Andrejeva J, Randall RE, Goodbourn S (2009) Mechanism of mda-5 inhibition by paramyxovirus V proteins. J Virol 83:1465–1473
de Vries W, Haasnoot J, van der Velden J, van Montfort T, Zorgdrager F, Paxton W, Cornelissen M, van Kuppeveld F, de Haan P, Berkhout B (2008) Increased virus replication in mammalian cells by blocking intracellular innate defense responses. Gene Ther 15:545–552
DiNapoli JM, Kotelkin A, Yang L, Elankumaran S, Murphy BR, Samal SK, Collins PL, Bukreyev A (2007) Newcastle disease virus, a host range-restricted virus, as a vaccine vector for intranasal immunization against emerging pathogens. Proc Natl Acad Sci U S A 104:9788–9793
Engel-Herbert I, Werner O, Teifke JP, Mebatsion T, Mettenleiter TC, Romer-Oberdorfer A (2003) Characterization of a recombinant Newcastle disease virus expressing the green fluorescent protein. J Virol Methods 108:19–28
Fields BN, Knipe DM (2001) Fields virology, 4th edn. Raven, New York, pp 1305–1340
Ge J, Deng G, Wen Z, Tian G, Wang Y, Shi J, Wang X, Li Y, Hu S, Jiang Y, Yang C, Yu K, Bu Z, Chen H (2007) Newcastle disease virus-based live attenuated vaccine completely protects chickens and mice from lethal challenge of homologous and heterologous H5N1 avian influenza viruses. J Virol 81:150–158
Gotoh B, Komatsu T, Takeuchi K, Yokoo J (2001) Paramyxovirus accessory proteins as interferon antagonists. Microbiol Immunol 45:787–800
Gotoh B, Komatsu T, Takeuchi K, Yokoo J (2002) Paramyxovirus strategies for evading the interferon response. Rev Med Virol 12:337–357
Hagmaier K, Stock N, Precious B, Childs K, Wang LF, Goodbourn S, Randall RE (2007) Mapuera virus, a rubulavirus that inhibits interferon signalling in a wide variety of mammalian cells without degrading STATs. J Gen Virol 88:956–966
Harper DR (1989) A novel plaque assay system for paramyxoviruses. J Virol Methods 25:347–350
Horvath CM (2004) Silencing STATs: lessons from paramyxovirus interferon evasion. Cytokine Growth Factor Rev 15:117–127
Huang Z, Krishnamurthy S, Panda A, Samal SK (2003) Newcastle disease virus V protein is associated with viral pathogenesis and functions as an alpha interferon antagonist. J Virol 77:8676–8685
Huang Z, Panda A, Elankumaran S, Govindarajan D, Rockemann DD, Samal SK (2004) The hemagglutinin-neuraminidase protein of Newcastle disease virus determines tropism and virulence. J Virol 78:4176–4184
Kattenbelt JA, Stevens MP, Gould AR (2006) Sequence variation in the Newcastle disease virus genome. Virus Res 116:168–184
Kournikakis B, Fildes J (1988) Titration of avirulent Newcastle disease virus by the plaque assay method. J Virol Methods 20:285–293
Kubota T, Yokosawa N, Yokota S, Fujii N, Tashiro M, Kato A (2005) Mumps virus V protein antagonizes interferon without the complete degradation of STAT1. J Virol 79:4451–4459
Malicki K, Toka FN (1996) Present status of classification of viruses of vertebrates. Arch Immunol Ther Exp 44:283–288
Mayo MA (2002) Virus taxonomy—Houston 2002. Arch Virol 147(1):1071–1076
Mebatsion T, Verstegen S, De Vaan LT, Romer-Oberdorfer A, Schrier CC (2001) A recombinant Newcastle disease virus with low-level V protein expression is immunogenic and lacks pathogenicity for chicken embryos. J Virol 75:420–428
Mebatsion T, de Vaan LT, de Haas N, Romer-Oberdorfer A, Braber M (2003) Identification of a mutation in editing-defective Newcastle disease virus recombinants that modulates P-gene mRNA editing and restores virus replication and pathogenicity in chicken embryos. J Virol 77:9259–9265
Mo CW, Cao YC, Lim BL (2001) The in vivo and in vitro effects of chicken interferon alpha on infectious bursal disease virus and Newcastle disease virus infection. Avian Dis 45:389–399
Nishio M, Tsurudome M, Ito M, Garcin D, Kolakofsky D, Ito Y (2005) Identification of paramyxovirus V protein residues essential for STAT protein degradation and promotion of virus replication. J Virol 79:8591–8601
Park MS, Garcia-Sastre A, Cros JF, Basler CF, Palese P (2003) Newcastle disease virus V protein is a determinant of host range restriction. J Virol 77:9522–9532
Peeters BP, de Leeuw OS, Koch G, Gielkens AL (1999) Rescue of Newcastle disease virus from cloned cDNA: evidence that cleavability of the fusion protein is a major determinant for virulence. J Virol 73:5001–5009
Puhler F, Willuda J, Puhlmann J, Mumberg D, Romer-Oberdorfer A, Beier R (2008) Generation of a recombinant oncolytic Newcastle disease virus and expression of a full IgG antibody from two transgenes. Gene Ther 15:371–383
Rodriguez JJ, Wang LF, Horvath CM (2003) Hendra virus V protein inhibits interferon signaling by preventing STAT1 and STAT2 nuclear accumulation. J Virol 77:11842–11845
Sinkovics JG, Horvath JC (2000) Newcastle disease virus (NDV): brief history of its oncolytic strains. J Clin Virol 16:1–15
Steward M, Vipond IB, Millar NS, Emmerson PT (1993) RNA editing in Newcastle-disease virus. J Gen Virol 74:2539–2547
Ulane CM, Kentsis A, Cruz CD, Parisien JP, Schneider KL, Horvath CM (2005) Composition and assembly of STAT-targeting ubiquitin ligase complexes: paramyxovirus V protein carboxyl terminus is an oligomerization domain. J Virol 79:10180–10189
Yang ZY, Kong WP, Huang Y, Roberts A, Murphy BR, Subbarao K, Nabel GJ (2004) A DNA vaccine induces SARS coronavirus neutralization and protective immunity in mice. Nature 428:561–564
Yoneyama M, Kikuchi M, Natsukawa T, Shinobu N, Imaizumi T, Miyagishi M, Taira K, Akira S, Fujita T (2004) The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat Immunol 5:730–737
Acknowledgements
This work was supported by the Small and Medium Business Administration of Korea (grant no. S0807222-E0843910-10100013) and Korea University Grant.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Comparison of the V gene and V protein sequences of lentogenic and velogenic NDV strains. a P and V gene mRNA sequences surrounding the RNA-editing sites. The introduction of one G (indicated as “+G”) in coding for the V gene ORF is shown. b Sequence alignment of lentogenic and velogenic V proteins. The start site of the V-specific C terminus is indicated by the closed triangle. c The cysteine-rich C terminus of the V proteins. The asterisks indicate conserved cysteine residues and the closed squares indicate conserved tryptophan residues (PDF 287 kb)
Supplementary Fig. 2
Constant expression of V protein during the whole NDV production period. Control cells and DF-1 cells expressing LaSota V or Kr-102/89 V protein were seeded in six-well plates at a density of 2.5 × 105 cells per well. Two days later, the cell monolayers were infected with the NDV LaSota at a MOI of 0.1 and incubated at 37°C in virus-production medium (basal MEM-α containing 400 ng/ml trypsin). Supernatants were harvested in 12-h intervals until the cells expressing NDV V protein were almost died (48 h after infection). The FLAG-tagged V protein and actin were detected by using monoclonal antibodies against FLAG tag and actin, respectively. FLAG-tagged NDV V protein was constantly expressed during the entire course of infection (GIF 30 kb)
Supplementary Fig. 3
Comparison of external proteases for NDV LaSota production. a NDV production kinetics with various external proteases in the production medium. DF-1 cells were seeded in six-well plates at a density of 2.5 × 105 cells per well. Two days later, the cell monolayers were infected with the NDV LaSota at a MOI of 0.1 and incubated at 37°C in basal MEM-α containing external protease. NDV production kinetics was almost the same regardless of the kind of protease when used with its optimized concentration. b NDV production kinetics upon amount of allantoic fluid. c NDV production kinetics upon amount of trypsin. As described in the text, when excessive protease is present in the production medium, the protease will destroy host cell monolayers and reduce the virus-production kinetics. On the other hand, insufficient protease is not able to support the rapid NDV production since it takes more time to cleave F protein of NDV newly produced from the cells (GIF 63 kb)
Rights and permissions
About this article
Cite this article
Jang, J., Hong, SH., Choi, D. et al. Overexpression of Newcastle disease virus (NDV) V protein enhances NDV production kinetics in chicken embryo fibroblasts. Appl Microbiol Biotechnol 85, 1509–1520 (2010). https://doi.org/10.1007/s00253-009-2189-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-009-2189-z