Abstract
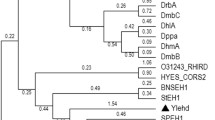
Previously, we reported that ten strains belonging to Erythrobacter showed epoxide hydrolase (EHase) activities toward various epoxide substrates. Three genes encoding putative EHases were identified by analyzing open reading frames of Erythrobacter litoralis HTCC2594. Despite low similarities to reported EHases, the phylogenetic analysis of the three genes showed that eeh1 was similar to microsomal EHase, while eeh2 and eeh3 could be grouped with soluble EHases. The three EHase genes were cloned, and the recombinant proteins (rEEH1, rEEH2, and rEEH3) were purified. The functionality of purified proteins was proved by hydrolytic activities toward styrene oxide. EEH1 preferentially hydrolyzed (R)-styrene oxide, whereas EEH3 preferred to hydrolyze (S)-styrene oxide, representing enantioselective hydrolysis of styrene oxide. On the other hand, EEH2 could hydrolyze (R)- and (S)-styrene oxide at an equal rate. The optimal pH and temperature for the EHases occurred largely at neutral pHs and 40–55 °C. The substrate selectivity of rEEH1, rEEH2, and rEEH3 toward various epoxide substrates were also investigated. This is the first representation that a strict marine microorganism possessed three EHases with different enantioselectivity toward styrene oxide.







Similar content being viewed by others
References
Arahira M, Nong VH, Udaka K, Fukazawa C (2000) Purification, molecular cloning and ethylene-inducible expression of a soluble-type epoxide hydrolase from soybean (Glycine max [L.] Merr.). Eur J Biochem 267:2649–2657
Arand M, Wagner H, Oesch F (1996) Asp333, Asp495, and His523 form the catalytic triad of rat soluble epoxide hydrolase. J Biol Chem 271:4223–4229
Arand M, Hemmer H, Durk H, Baratti J, Archelas A, Furstoss R, Oesch F (1999) Cloning and molecular characterization of a soluble epoxide hydrolase from Aspergillus niger that is related to mammalian microsomal epoxide hydrolase. Biochem J 344:273–280
Archelas A, Furstoss R (1997) Synthesis of enantiopure epoxides through biocatalytic approaches. Annu Rev Microbiol 51:491–525
Archelas A, Furstoss R (2001) Synthetic applications of epoxide hydrolases. Curr Opin Chem Biol 5:112–119
Beetham JK, Grant D, Arand M, Garbarino J, Kiyosue T, Pinot F, Oesch F, Belknap WR, Shinozaki K, Hammock BD (1995) Gene evolution of epoxide hydrolase and recommended nomenclature. DNA Cell Biol 14:61–71
Bhatnagar T, Manoj KM, Baratti JC (2001) A spectrophotometric method to assay epoxide hydrolase activity. J Biochem Biophys Methods 50:1–13
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Cao L, Lee J, Chen W, Wood TK (2006) Enantioconvergent production of (R)-1-phenyl-1,2-ethanediol from styrene oxide by combining the Solanum tuberosum and an evolved Agrobacterium radiobacter AD1 epoxide hydrolases. Biotechnol Bioeng 94:522–529
Falany CN, McQuiddy P, Kasper CB (1987) Structure and organization of the microsomal xenobiotic epoxide hydrolase gene. J Biol Chem 262:5924–5930
Giovannoni SJ, Stingl U (2005) Molecular diversity and ecology of microbial plankton. Nature 437:343–348
Janssen DB, Pries F, Ploeg J, Kazemier B, Terpstra P, Witholt B (1989) Cloning of 1,2-dichloroethane degradation genes of Xanthobacter autotrophicus GJ10 and expression and sequencing of the dhlA gene. J Bacteriol 171:6791–6799
Kaneko T, Nakamura Y, Sato S, Minamisawa K, Uchiumi T, Sasamoto S, Watanabe A, Idesawa K, Iriguchi M, Kawashima K, Kohara M, Matsumoto M, Shimpo S, Tsuruoka H, Wada T, Yamada M, Tabata S (2002) Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA110. DNA Res 9:189–197
Knehr M, Thomas H, Arand M, Gebel T, Zeller HD, Oesch F (1993) Isolation and characterization of a cDNA encoding rat liver cytosolic epoxide hydrolase and its functional expression in Escherichia coli. J Biol Chem 268:17623–17627
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the heal of bacteriophage T4. Nature 227:680–685
Misawa E, Chion CK, Archer IV, Woodland MP, Zhou NY, Carter SF, Widdowson DA, Leak DJ (1998) Characterisation of a catabolic epoxide hydrolase from a Corynebacterium sp. Eur J Biochem 253:173–183
Morisseau C, Beetham JK, Pinot F, Debernard S, Newman JWB, Hammock D (2000) Cress and potato soluble epoxide hydrolases: purification, biochemical characterization, and comparison to mammalian enzymes. Arch Biochem Biophys 378:321–332
Nardini M, Dijkstra BW (1999) α/β hydrolase fold enzymes: the family keeps growing. Curr Opin Struct Biol 9:732–737
Ollis DL, Cheah E, Cygler M, Dijkstra B, Frolow F, Franken SM, Harel M, Remington SJ, Silman I, Schrag J, Sussman JL, Verschueren KHG, Goldman A (1992) The α/β hydrolase fold. Protein Eng 5:197–211
Oppenheimer CH, ZoBell CE (1952) The growth and viability of sixty-three species of marine bacteria as influenced by hydrostatic pressure. J Mar Res 11:10–18
Rink R, Janssen DB (1998) Kinetic mechanism of the enantioselective conversion of styrene oxide by epoxide hydrolase from Agrobacterium radiobacter AD1. Biochemistry 37:18119–18127
Rink R, Fennema M, Smids M, Dehmel U, Janssen DB (1997) Primary structure and catalytic mechanism of the epoxide hydrolase from Agrobacterium radiobacter AD1. J Biol Chem 272:14650–14657
Rink R, Spelberg JHL, Pieters RJ, Kingma J, Nardini M, Kellogg RM, Dijkstra BW, Janssen DB (1999) Mutation of tyrosine residues involved in the alkylation half reaction of epoxide hydrolase from Agrobacterium radiobacter AD1 results in improved enantioselectivity. J Am Chem Soc 121:7417–7418
Rink R, Kingma J, Spelberg JH, Janssen DB (2000) Tyrosine residues serve as proton donor in the catalytic mechanism of epoxide hydrolase from Agrobacterium radiobacter. Biochemistry 39:5600–5613
Rui L, Cao L, Chen W, Reardon KF, Wood TK (2005) Protein engineering of epoxide hydrolase from Agrobacterium radiobacter AD1 for enhanced activity and enantioselective production of (R)-1-phenylethane-1,2-diol. Appl Environ Microbiol 71:3995–4003
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual (vol 2, 3rd edn.). Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, pp 8.65–8.71
Stapleton A, Beetham JK, Pinot F, Garbarino JE, Rockhold DR, Friedman M, Hammock BD, Belknap WR (1994) Cloning and expression of soluble epoxide hydrolase from potato. Plant J 6:251–258
Strausberg RL, Feingold EA, Grouse LH, Derge JG, Klausner RD, Collins FS, Wagner L, Shenmen CM, Schuler GD, Altschul SF, Zeeberg B, Buetow KH, Schaefer CF, Bhat NK, Hopkins RF, Jordan H, Moore T, Max SI, Wang J, Hsieh F, Diatchenko L, Marusina K, Farmer A, Rubin GM, Hong L, Stapleton M, Soares MB, Bonaldo MF, Casavant TL, Scheetz TE, Brownstein MJ, Usdin TB, Toshiyuki S, Carninci P, Prange C, Raha SS, Loquellano NA, Peters GJ, Abramson RD, Mullahy SJ, Bosak SA, McEwan PJ, McKernan KJ, Malek JA, Gunaratne PH, Richards S, Worley KC, Hale S, Garcia AM, Gay LJ, Hulyk SW, Villalon DK, Muzny DM, Sodergren EJ, Lu X, Gibbs RA, Fahey J, Helton E, Ketteman M, Madan A, Rodrigues S, Sanchez A, Whiting M, Madan A, Young AC, Shevchenko Y, Bouffard GG, Blakesley RW, Touchman JW, Green ED, Dickson MC, Rodriguez AC, Grimwood J, Schmutz J, Myers RM, Butterfield YS, Krzywinski MI, Skalska U, Smailus DE, Schnerch A, Schein JE, Jones SJ, Marra MA (2002) Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences. Proc Natl Acad Sci USA 99:16899–16903
Thompson JD, Higgin DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tokunaga M, Larrow JF, Kakiuchi F, Jacobsen EN (1997) Asymmetric catalysis with water: efficient kinetic resolution of terminal epoxides by means of catalytic hydrolysis. Science 277:936–938
van Loo B, Spelberg JHL, Kingma J, Sonke T, Wubbolts MG, Janssen DB (2004) Directed evolution of epoxide hydrolase from A. radiobacter toward higher enantioselectivity by error-prone PCR and DNA shuffling. Chem Biol 11:981–990
van Loo B, Kingma J, Arand M, Wubbolts MG, Janssen DB (2006) Diversity and biocatalytic potential of epoxide hydrolases identified by genome analysis. Appl Environ Microbiol 72:2905–2917
Venter JC, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA, Wu D, Paulsen I, Nelson KE, Nelson W, Fouts DE, Levy S, Knap AH, Lomas MW, Nealson K, White O, Peterson J, Hoffman J, Parsons R, Baden-Tillson H, Pfannkoch C, Rogers YH, Smith HO (2004) Environmental genome shotgun sequencing of the Sargasso sea. Science 304:66–74
Visser H, Bont JAM, Verdoes JC (1999) Isolation and characterization of the epoxide hydrolase-encoding gene from Xanthophyllomyces dendrorhous. Appl Environ Microbiol 65:5459–5463
Visser H, Vreugdenhil S, Bont JAM, Verdoes JC (2000) Cloning and characterization of an epoxide hydrolase-encoding gene from Rhodotorula glutinis. Appl Microbiol Biotechnol 53:415–419
Weijers CAGM, Bont JAM (1999) Epoxide hydrolases from yeasts and other sources: versatile tools in biocatalysis. J Mol Catal, B Enzym 6:199–214
Yamada T, Morisseau C, Maxwell JE, Argiriadi MA, Christianson DW, Hammock BD (2000) Biochemical evidence for the involvement of tyrosine in epoxide activation during the catalytic cycle of epoxide hydrolase. J Biol Chem 275:23082–23088
Zou J, Hallberg BM, Bergfors T, Oesch F, Arand M, Mowbray SL, Jones TA (2000) Structure of Aspergillus niger epoxide hydrolase at 1.8 A resolution: implications for the structure and function of the mammalian microsomal class of epoxide hydrolases. Structure 15:111–122
Acknowledgement
This work was supported by KORDI in-house program (PE97803) and the Marine and Extreme Genome Research Center Program, Ministry of Marine Affairs and Fisheries, Republic of Korea. We would like to appreciate the Gordon and Betty Moore Foundation for the great effort on Microbial Genome Sequencing Project (Moore 155).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Woo, JH., Hwang, YO., Kang, S.G. et al. Cloning and characterization of three epoxide hydrolases from a marine bacterium, Erythrobacter litoralis HTCC2594. Appl Microbiol Biotechnol 76, 365–375 (2007). https://doi.org/10.1007/s00253-007-1011-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-1011-z