Abstract
The novel chitinolytic bacterium Serratia sp. KCK, which was isolated from kimchi juice, produced chitinase A. The gene coding for the chitinolytic enzyme was cloned on the basis of sequencing of internal peptides, homology search, and design of degenerated primers. The cloned open reading frame of chiA encodes for deduced polypeptide of 563 amino acid residues with a calculated molecular mass of 61 kDa and appears to correspond to a molecular mass of about 57 kDa, which excluded the signal sequence. The deduced amino acid sequence showed high similarity to those of bacterial chitinases classified as family 18 of glycosyl hydrolases. The chitinase A is an exochitinase and exhibits a greater pH range (5.0–10.0), thermostability with a temperature optimum of 40°C, and substrate range other than Serratia chitinases thus far described. These results suggested that Serratia sp. KCK chitinase A can be used for biotechnological applications with good potential.






Similar content being viewed by others

References
Adams DJ (2004) Fungal cell wall chitinases and glucanases. Microbiology 150:2029–2035
Bhushan B (2000) Production and characterization of a thermostable chitinase from a new alkalophilic Bacillus sp. BG-11. J Appl Microbiol 88:800–808
Cheigh HS, Park KY (1994) Biochemical, microbiological, and nutritional aspects of kimchi (Korean fermented vegetable products). Crit Rev Food Sci Nutr 34:175–203
Chernin LS, Fuente LDL, Sobolev V, Haran S, Vorgias CE, Oppenheim AB, Chet I (1997) Molecular cloning, structural analysis, and expression in Escherichia coli of a chitinase gene from Enterobacter agglomerans. Appl Environ Microbiol 63:834–839
Cho J, Lee D, Yang C, Jeon J, Kim J, Han H (2006) Microbial population dynamics of kimchi, a fermented cabbage product. FEMS Microbiol Lett 257:262–267
Cohen-Kupiec R, Chet I (1998) The molecular biology of chitin digestion. Curr Opin Biotechnol 9:270–277
Dahiya N, Tewari R, Singh, Hoondal GS (2006) Biotechnological aspects of chitinolytic enzymes: a review. Appl Microbiol Biotechnol 71:773–782
Frankowski J, Lorito M, Scala F, Schmid R, Berg G, Bahl H (2001) Purification and properties of two chitinolytic enzymes of Serratia plymuthica HRO-C48. Arch Microbiol 176:421–426
Golyshina OV, Golyshin PN, Timmis KN, Ferrer M (2005) The ‘pH optimum anomaly’ of intracellular enzymes of Ferroplasma acidiphilum. Environ Microbiol 8:416–425
Gooday GW (1990) Physiology of microbial degradation of chitin and chitosan. Biodegradation 1:177–190
Hobel CFV, Hreggvidsson GÓ, Marteinsson VT, Bahrani-Mougeot F, Einarsson JM, Kristjánsson JK (2005) Cloning, expression, and characterization of a highly thermostable family 18 chitinase from Rhodothermus marinus. Extremophiles 9:53–64
Imanaka T, Fukui T, Fujiwara S (2001) Chitinase from Thermococcus kodakaraensis KOD1. Methods Enzymol 330:319–329
Imoto T, Yagishita K (1971) A simple activity measurement of lysozyme. Agric Biol Chem 35:1154–1156
Kawase T, Yokokawa S, Saito A, Fujii T, Nikaidou N, Miyashita K, Watanabe T (2006) Comparison of enzymatic and antifungal properties between family 18 and 19 chitinase from Streptomyces coelicolor A3(2). Biosci Biotechnol Biochem 70:988–998
Keyhani NO, Roseman S (1999) Physiological aspects of chitin catabolism in marine bacteria. Biochim Biophys Acta 1473:108–122
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 277:680–685
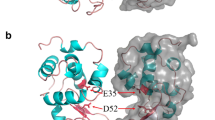
Lambert C, Leonard N, De Bolle X, Depiereux E (2002) ESyPred3D: Predition of protein 3D structures. Bioinformatics 18:1250–1256
Lan X, Zhang X, Hu J, Shimosaka M (2006) Cloning, expression and characterization of a chitinase from the chitinolytic bacterium Aeromonas hydrophila strain SUWA-9. Biosci Biotechnol Biochem 70:2437–2442
Maidak BL, Olsen GJ, Larsen N, Overbeek R, McCaughey MJ, Woese CR (1997) The RDP (Ribosomal Database Project). Nucleic Acids Res 25:109–111
Nawani NN, Kapadnis BP (2001) One-step purification of chitinase from Serratia marcescens NK1, a soil isolate. J Appl Microbiol 90:803–808
Ohno T, Armand S, Hata T, Nikaidou N, Henrissat B, Mitsutomi M, Watanabe T (1996) A modular family 19 chitinase found in the prokaryotic organism Streptomyces griseus HUT 6037. J Bacteriol 178:5065–5070
Orikoshi H, Nakayama S, Miyamoto K, Hanato C, Yasuda M, Inamori Y, Tsujibo H (2005) Roles of four chitinases (ChiA, ChiB, ChiC, and ChiD) in the chitin degradation system of marine bacterium Alteromonas sp. strain 0-7. Appl Environ Microbiol 71:1811–1815
Perrakis A, Tews I, Dauter Z, Oppenheim AB, Chet I, Wilson KS, Vorgias CE (1994) Crystal structure of a bacterial chitinase at 2.3 Å resolution. Structure 2:1169–1180
Sakai K, Yokota A, Kurokawa H, Wakayama M, Moriguchi M (1998) Purification and characterization of three thermostable endochitinases of a novel Bacillus strain, MH-1, isolated from chitin-containing compost. Appl Environ Microbiol 64:3397–3402
Shigemasa Y, Minami S (1995) Application of chitin and chitosan for biomaterials. Biotechnol Genet Eng Rev 13:383–420
Suzuki K, Sugawara N, Suzuki M, Uchiyama T, Katouno F, Nikaidou N, Watanabe T (2002) Chitinases A, B, and C1 of Serratia marcescens 2170 produced by recombinant Escherichia coli: enzymatic properties and synergism on chitin degradation. Biosci Biotechnol Biochem 66:1075–1083
Svitil AL, Chadhain SMN, Moore JA, Kirchman DL (1997) Chitin degradation proteins produced by the marine bacterium Vibrio harveyi growing on different forms of chitin. Appl Environ Microbiol 63:408–413
Tanaka T, Fukui T, Imanaka T (2001) Different cleavage specificities of the dual catalytic domains in chitinase from the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J Biol Chem 276:35629–35635
Trudel J, Asselin A (1989) Detection of chitinase activity after polyacrylamide gel electrophoresis. Anal Biochem 178:362–366
Tsujibo H, Kubota T, Yamamoto M, Miyamoto K, Inamori Y (2003) Characterization of chitinase genes from an alkaliphilic actinomycete, Nocardiopsis prasina OPC-131. Appl Environ Microbiol 69:894–900
Vaidya RJ, Macmil SLA, Vyas PR, Chhatpar HS (2003) The novel method for isolating chitinolytic bacteria and its application in screening for hyperchitinase producing mutant of Alcaligenes xylosoxydans. Lett Appl Microbiol 36:129–134
Wang SL, Chang WT (1997) Purification and characterization of two bifunctional chitinase/lysozyme extracellulary produced by Pseudomonas aeruginosa K-187 in a shrimp and crab shell powder medium. Appl Environ Microbiol 63:380–386
Watanabe T, Kimura K, Sumiya T, Nikaidou N, Suzuki K, Suzuki M, Taiyoji M, Ferrer S, Regue M (1997) Genetic analysis of the chitinase system of Serratia marcescens 2170. J Bacteriol 179:7111–7117
Wiwat C, Siwayaprahm P, Bhumiratana A (1999) Purification and characterization of chitinase from Bacillus circulans No.4.1. Curr Microbiol 39:134–140
Xia G, Jin C, Zhou J, Yang S, Zhang S, Jin C (2001) A novel chitinase having a unique mode of action from Aspergillus fumigatus YJ-407. Eur J Biochem 268:4079–4085
Acknowledgements
We gratefully acknowledge MetaGenoMik Project of Federal Ministry for Science and Education (BMBF) and Fonds der Chemischen Industrie for generous support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, HS., Timmis, K.N. & Golyshin, P.N. Characterization of a chitinolytic enzyme from Serratia sp. KCK isolated from kimchi juice. Appl Microbiol Biotechnol 75, 1275–1283 (2007). https://doi.org/10.1007/s00253-007-0947-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-0947-3