Abstract
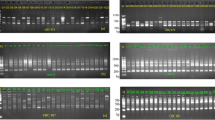
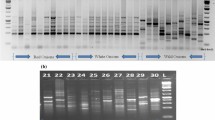
Wild populations of edible species are important source of genetic variability for cultivated lines that can undergo a drastic loss of diversity resulting from man’s selection. The development of tools aimed at the clear-cut and safe identification and assessment of genetic variability of the wild and cultivated strains is thus a fundamental goal of molecular genetic research. In this study, we used two polymerase chain reaction (PCR)-based fingerprinting methods—amplified fragment length polymorphism (AFLP) and restriction fragment length polymorphism (RFLP) of laccase and manganese peroxidase genes—to assess genetic differences among strains and independently evolving lineages belonging to the Pleurotus eryngii complex. Both laccase RFLP and AFLP have been proved to distinguish unambiguously the three taxa studied: Pleurotus ferulae, P. eryngii, and P. eryngii var. nebrodensis. AFLP also showed enough sensitivity to detect polymorphisms among the strains, proving to be an efficient DNA fingerprinting tool in studies of strain assignment. The divergent RFLP laccase and manganese peroxidase patterns are also discussed in relation to the role played by these genes in the interaction between these fungi and their host plants.



Similar content being viewed by others
References
Cailleux R, Diop A, Joly P (1981) Relations d’interfertilitè entre quelques reprèsententes des Pleurotes des Ombrellifères. Bull Trimest Soc Mycol Fr 97:97–124
Calvo-Bado L, Noble R, Challen M, Dobrovin-Pennington A, Elliott T (2000) Sexuality and genetic identity in the Agaricus section Arvenses. Appl Environ Microbiol 66:728–734
Chiu SW, Chen MJ, Chang ST (1995) Differentiating homothallic mushrooms Volvariella by AP-PCR and RFLPs. Mycol Res 99:333–336
Chiu SW, Ma AM, Lin FC, Moore D (1996) Genetic homogeneity of cultivated strains of shiitake (Lentinula edodes) used in China as revealed by the polymerase chain reaction. Mycol Res 100:1393–1399
De Gioia T, Sisto D, Rana GL, Figliuolo G (2005) Genetic structure of the Pleurotus eryngii species-complex. Mycol Res 109:71–80
Della Rosa V, Cappuccio I, Fanelli C, Urbanelli S (2004) Isolation and characterization of microsatellite markers in two basidiomycete species: Pleurotus eryngii and P. ferulae (DC.:Fr.). Mol Ecol Notes 4:271–273
Eggert C, Temp U, Eriksson K-EL (1996) The ligninolytic sytem of the white-rot fungus Pycnoporus cinnabarinus: purification and characterization of the laccase. Appl Environ Microbiol 62:1151–1158
Ellstrand NC, Elam DR (1993) Population genetic consequences of small population size: implications for plant conservation. Ann Rev Ecolog Syst 24:217–242
Felsenstein J (1993) PHYLIP (phylogeny interference package), version 3.5c. University of Washington, Seattle
Felsestein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution Int J Org Evolution 39:783–791
Fowler C, Hodgkin T (2004) Plant genetic resources for food and agriculture: assessing global availability. Annu Rev Environ Resour 29:143–179
Iraçabal B, Zervakis G, Labarère J (1995) Molecular systematics of the genus Pleurotus: analysis of restriction polymorphisms in ribosomal DNA. Microbiology 141:1479–1490
Hamrick JL, Godt MJW (1996) Allozyme diversity in plant species. In: Brown AHD, Clegg MT (eds) Plant population genetics, breeding and genetic resources. Sinauer, Sunderland, Massachusetts, USA, pp 43–63
Higgins DG, Thompson JD, Gibson TJ (1996) Using CLUSTAL for multiple sequence alignments. Methods Enzymol 266:383–402
Ito Y, Fushimi T, Yanagi SO (1998) Discrimination of species and strains of basidiomycete genus Coprinus by random amplified polymorphic DNA (RAPD) analysis. Mycoscience 39:361–365
Khush RS, Becker E, Wach M (1992) DNA amplification polymorphisms of the cultivated mushroom Agaricus bisporus. Appl Environ Microbiol 58:2971–2977
Martínez AT, Speranza M, Ruiz-Dueńas FJ, Ferriera P, Camarero S, Guillén F, Martínez MJ, Gutiérrez A, del Río JC (2005) Biodegradation of lignocellulosics: microbial, chemical, and enzymatic aspects of the fungal attack of lignin. Int Microbiol 8:195–204
Miller MP (1997) Tools for Population Genetic Analyses (TFPGA) version 1.3 A Windows® program for the analysis of allozyme and molecular population genetic data. Computer software distributed by author
Miyazaki Y, Nakamura M, Babasaki K (2005) Molecular cloning of developmentally specific genes by representational difference analysis during the fruiting body formation in the basidiomycete Lentinula edodes. Fungal Genet Biol 42:493–498
Mueller UG, Wolfenbarger L (1999) AFLP genotyping and fingerprinting. Trends Ecol Evol 14:389–394
Nei M (1978) Estimation of average heterozygosity and genetics distance from a small number of individuals. Genetics 89:583–590
Noël T, Labarere J (1987) Isolation of DNA from Agrocybe aegerita for the construction of a genomic library in E. coli. Mushroom Sci 12:187–201
Palmieri G, Giardina P, Bianco C, Fontanella B, Sannia G (2000) Copper induction of laccase isoenzymes in the ligninolytic fungus Pleurotus ostreatus. Appl Envion Microbiol 66(3):920–924
Ramanata Rao V, Hodgkin T (2002) Genetic diversity and conservation and utilisation of plant genetic resources. Plant Cell Tissue Organ Cult 68:1–19
Ramírez L, Muez V, Alfonso M, Garcia Barrenechea A, Alfonso L, Pisabarro AG (2001) Use of molecular markers to differentiate between commercial strains of the button mushrooms Agaricus bisporus. FEMS Microbiol Lett 198(1):45–48
Soden DM, Dobson ADW (2001) Differential regulation of laccase gene expression in Pleurotus sajor-caju. Microbiology 147:1755–1763
Sokal RR, Michener CD (1958) A statistical method for evaluating systematic relationships. Univ Kans Sci Bull 38:1409–1438
Terashima K, Matsumoto T (2004) Strain typing of shiitake (Lentinula edodes) cultivars by AFLP analysis, focusing on a heat-dried fruiting body. Mycoscience 45:79–82
Terefework Z, Kaijalainen S, Lindström K (2001) AFLP fingerprinting as a tool to study the genetic diversity of Rhizobium galegae from Galega orientalis and Galega officinalis. J Biotechnol 91:169–180
Urbanelli S, Fanelli C, Fabbri AA, Della Rosa V, Maddau L, Marras F, Reverberi M (2002) Molecular genetic analysis of two taxa of the Pleurotus eryngii complex: P. eryngii (DC.Fr.) Quel. var. eryngii and P. eryngii (DC.Fr.) Quel. var. ferulae. Biol J Linn Soc 75(1):125–136
Urbanelli S, Fanelli C, Fabbri AA, Della Rosa V, Reverberi M (2003) Genetic diversity and population structure of the Italian fungi belonging to the taxa Pleurotus eryngii and Pleurotus ferulae (DC.:Fr.). Heredity 90(3):253–259
Valsangiacomo C, Baggi F, Gaia V, Balmelli T, Peduzzi R, Piffaretti JC (1995) Use of amplified fragment length polymorphism in molecular typing of Legionella pneumophila and application to epidemiological studies. J Clin Microbiol 33:1716–1719
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Homes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Xu J, Kerrigan RW, Callac P, Horgen PA, Anderson JB (1997) Genetic structure of natural populations of Agaricus bisporus, the commercial button mushroom. J Hered 88:482–488
Xu J, Kerrigan RW, Sonnenberg A, Callac P, Horgen PA, Anderson JB (1998) Mitochondrial DNA variation in natural populations of the mushroom Agaricus bisporus. Mol Ecol 7:19–33
Zervakis GI, Venturella G, Papadopoulou K (2001) Genetic polymorphism and taxonomic infrastructure of the Pleurotus eryngii species-complex as determined by RAPD analysis, isozyme profiles and ecomorphological characters. Microbiology 147:3183–3194
Zhang JX, Huang CY, Ng TB, Wang HX (2006) Genetic polymorphism of ferula mushroom growing on Ferula sinkiangensis. Appl Microbiol Biotechnol 71(3):304–309
Acknowledgements
We thank the anonymous reviewer for his helpful comments, Alessandra Spanò and Slaven Zjalic for their invaluable cooperation, and Monica Brocco for the linguistic revision. The research was funded with grants from the Ministero delle Politiche Agricole e Forestali and the Life Science Project of the European Community.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Urbanelli, S., Della Rosa, V., Punelli, F. et al. DNA-fingerprinting (AFLP and RFLP) for genotypic identification in species of the Pleurotus eryngii complex. Appl Microbiol Biotechnol 74, 592–600 (2007). https://doi.org/10.1007/s00253-006-0684-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-006-0684-z