Abstract
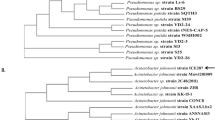
A strain of Pseudomonasputida ZWL73 was isolated from soil contaminated with chloronitrobenzenes and identified by 16S rDNA sequencing. This bacterium released chloride and ammonia into the medium when grown on 4-chloronitrobenzene (4CNB) as the sole source of carbon, nitrogen and energy. A plasmid designated pZWL73 of approximately 100 kb in this strain was found to be responsible for 4CNB degradation. This was based on the fact that the plasmid-cured strains showed 4CNB− phenotype and the 4CNB+ phenotype could be conjugally transferred. The cell-free extracts of strain ZWL73 exhibited chloronitrobenzene nitroreductase and 2-amino-5-chlorophenol 1, 6-dioxygenase (2A5CPDO) activities, but neither activity was found from that of the plasmid-cured strain. We have also cloned a 4.9-kb EcoRI fragment exhibiting 2A5CPDO activity. Sequencing results revealed β-subunit (cnbCa) and α subunit (cnbCb) of a meta-cleavage dioxygenase, which were subsequently expressed in E. coli with 2A5CPDO activity. The phylogenetic analysis suggested that 2A5CPDO may form a new subgroup in class III meta-cleavage dioxygenase with its close homologs.





Similar content being viewed by others
References
Bradford MM (1976) A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Corbett MD, Corbett BR (1981) Metabolism of 4-chloronitrobenzene by the yeast Rhodosporidium sp. Appl Environ Microbiol 41:942–949
Dennis JJ, Zylstra GJ (2004) Complete sequence and genetic organization of pDTG1, the 83 kilobase naphthalene degradation plasmid from Pseudomonas putida strain NCIB 9816-4. J Mol Biol 341:753–768
Ditta G, Stanfield S, Corbin D, Helinski DR (1980) Broad host range DNA cloning system for gram-negative bacteria: construction of a gene bank of Rhizobium meliloti. Proc Natl Acad Sci USA 77:7347–7351
Eaton RW (2001) Plasmid-encoded phthalate catabolic pathway in Arthrobacter keyseri 12B. J Bacteriol 183:3689–3703
Eltis LD, Bolin JT (1996) Evolutionary relationships among extradiol dioxygenases. J Bacteriol 178:5930–5937
Gerhardt P, Murray RGE, Wood WA, Krieg NR (1994) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, District of Columbia
Greated A, Lambertsen L, Williams PA, Thomas CM (2002) Complete sequence of the IncP-9 TOL plasmid pWW0 from Pseudomonas putida. Environ Microbiol 4:856–871
Johnson GR, Jain RK, Spain JC (2002) Origins of the 2, 4-dinitrotoluene pathway. J Bacteriol 184:4219–4232
Katsivela E, Wray V, Pieper DH, Wittich RM (1999) Initial reactions in the biodegradation of 1-chloro-4-nitrobenzene by a newly isolated bacterium, Strain LW1. Appl Environ Microbiol 65:1405–1412
Keen NT, Tamaki S, Kobayashi D, Trollinger D (1988) Improved broad-host-range plasmids for DNA cloning in Gram-negative bacteria. Gene 70:191–197
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Liu H, Wang SJ, Zhou NY (2005) A new isolate of Pseudomonas stutzeri that degrades 2-chloronitrobenzene. Biotechnol Lett 27:275–278
Nishino SF, Spain JC (1993) Degradation of nitrobenzene by a Pseudomonas pseudoalcaligenes. 59:2520–2525
Park HS, Kim HS (2000) Identification and characterization of the nitrobenzene catabolic plasmids pNB1 and pNB2 in Pseudomonasputida HS12. J Bacteriol 182:573–580
Park HS, Lim SJ, Chang YK, Livingston AG, Kim HS (1999) Degradation of chloronitrobenzenes by a coculture of Pseudomonas putida and a Rhodococcus sp. Appl Environ Microbiol 65:1083–1091
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Schackmann A, Muller R (1991) Reduction of nitroaromatic compounds by different Pseudomonas species under aerobic conditions. Appl Microbiol Biotechnol 34:809–813
Schuman JD, Zottola EA, Harlander SK (1989) Preliminary characterization of a food-borne multiple-antibiotic-resistant Salmonella typhimurium Strain. Appl Environ Microbiol 55:2344–2348
Spence EL, Kawamukai M, Sanvoisin J, Braven H, Bugg TD (1996) Catechol dioxygenases from Escherichia coli (MhpB) and Alcaligenes eutrophus (MpcI): sequence analysis and biochemical properties of a third family of extradiol dioxygenases. J Bacteriol 178:5249–5256
Sugimoto K, Senda T, Aoshima H, Masai E, Fukuda M, Mitsui Y (1999) Crystal structure of an aromatic ring opening dioxygenase LigAB, a protocatechuate 4,5-dioxygenase, under aerobic conditions. Structure 7:953–965
Top EM, Springael D (2003) The role of mobile genetic elements in bacterial adaptation to xenobiotic organic compounds. Curr Opin Biotechnol 14:262–269
Wheatcroft R, Williams PA (1981) Rapid methods for the study of both stable and unstable plasmids in Pseudomonas. J Gen Microbiol 124:433–437
Williams PA, Murray K (1974) Metabolism of benzoate and the methylbenzoates by Pseudomonas putida (arvilla) mt-2: evidence for the existence of a TOL plasmid. J Bacteriol 120:416–423
Wu JF, Sun CW, Jiang CY, Liu ZP, Liu SJ (2005) A novel 2-aminophenol 1,6-dioxygenase involved in the degradation of p-chloronitrobenzene by Comamonasstrain CNB-1: purification, properties, genetic cloning and expression in Escherichia coli. Arch Microbiol 183:1–8
Acknowledgements
This work was supported by the National Natural Science Foundation of China (NSFC30230010). The authors would like to thank Peter Williams for Pseudomonas putida PaW340 and PaW1.
Author information
Authors and Affiliations
Corresponding author
Additional information
# D.Z. and H.L. equally contributed to this work.
Rights and permissions
About this article
Cite this article
Zhen, D., Liu, H., Wang, SJ. et al. Plasmid-mediated degradation of 4-chloronitrobenzene by newly isolated Pseudomonas putida strain ZWL73. Appl Microbiol Biotechnol 72, 797–803 (2006). https://doi.org/10.1007/s00253-006-0345-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-006-0345-2