Abstract
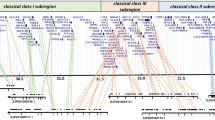
Genes of the major histocompatibility complex (MHC) are exceptionally polymorphic due to the combined effects of natural and sexual selection. Most research in wild populations has focused on the second exon of a single class II locus (DRB), but complete gene sequences can provide an illuminating backdrop for studies of intragenic selection, recombination, and organization. To this end, we characterized class II loci in the banner-tailed kangaroo rat (Dipodomys spectabilis). Seven DRB-like sequences (provisionally named MhcDisp-DRB*01 through *07) were isolated from spleen cDNA and most likely comprise ≥5 loci; this multiformity is quite unlike the situation in muroid rodents such as Mus, Rattus, and Peromyscus. In silico translation revealed the presence of important structural residues for glycosylation sites, salt bonds, and CD4+ T-cell recognition. Amino-acid distances varied widely among the seven sequences (2–34%). Nuclear DNA sequences from the Disp-DRB*07 locus (∼10 kb) revealed a conventional exon/intron structure as well as a number of microsatellites and short interspersed nuclear elements (B4, Alu, and IDL-Geo subfamilies). Rates of nucleotide substitution at Disp-DRB*07 are similar in both exons and introns (π = 0.015 and 0.012, respectively), which suggests relaxed selection and may indicate that this locus is an expressed pseudogene. Finally, we performed BLASTn searches against Dipodomys ordii genomic sequences (unassembled reads) and find 90–97% nucleotide similarity between the two kangaroo rat species. Collectively, these data suggest that class II diversity in heteromyid rodents is based on polylocism and departs from the muroid architecture.



Similar content being viewed by others
References
Acevedo-Whitehouse K, Cunningham AA (2006) Is MHC enough for understanding wildlife immunogenetics? Trends Ecol Evol 21:433–438 doi:10.1016/j.tree.2006.05.010
Andersson L, Gustafsson K, Jonsson A-K, Rask L (1991) Concerted evolution in a segment of the first domain exon of polymorphic MHC class II b loci. Immunogenetics 33:235–242
Apanius V, Penn D, Slev PR, Ruff LR, Potts WK (1997) The nature of selection on the major histocompatibility complex. Crit Rev Immunol 17:179–224
Axtner J, Sommer S (2007) Gene duplication, allelic diversity, selection processes and adaptive value of MHC class II DRB genes of the bank vole, Clethrionomys glareolus. Immunogenetics 59:417–426 doi:10.1007/s00251-007-0205-y
Babik W, Durka W, Radwan J (2005) Sequence diversity of the MHC DRB gene in the Eurasian beaver (Castor fiber). Mol Ecol 14:4249–4257 doi:10.1111/j.1365-294X.2005.02605.x
Babik W, Radwan J (2007) Sequence diversity of MHC class II DRB genes in the bank vole Myodes glareolus. Acta Theriol (Warsz) 52:227–235
Bernatchez L, Landry C (2003) MHC studies in nonmodel vertebrates: what have we learned about natural selection in 15 years? J Evol Biol 16:363–377 doi:10.1046/j.1420-9101.2003.00531.x
Bondinas GP, Moustakas AK, Papadopoulos GK (2007) The spectrum of HLA-DQ and HLA-DR alleles, 2006: a listing correlating sequence and structure with function. Immunogenetics 59:539–553 doi:10.1007/s00251-007-0224-8
Bos DH, Waldman B (2006) Evolution by recombination and transspecies polymorphism in the MHC class I gene of Xenopus laevis. Mol Biol Evol 23:137–143 doi:10.1093/molbev/msj016
Bos DH, Gopurenko D, Williams RN, DeWoody JA (2008) Inferring population history and demography using microsatellites, mitochondrial DNA, and major histocompatibility complex (MHC) genes. Evolution Int J Org Evolution 62:1458–1468 doi:10.1111/j.1558-5646.2008.00364.x
Bowen L, Aldridge BM, Gulland F, Van Bonn W, Delong R, Melin S et al (2004) Class II multiformity generated by variable MHC-DRB region configurations in the California sea lion (Zalophus californianus). Immunogenetics 56:12–27 doi:10.1007/s00251-004-0655-4
Brown JH, Davidson DW, Munger JC, Inouye RS (1986) Experimental community ecology: the desert granivore system. Harper & Row, New York
Brown JH, Jardetzky TS, Gorga JC, Stern LJ, Urban RG, Strominger JL et al (1993) Three-dimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature 364:33–39 doi:10.1038/364033a0
Busch JD, Waser PM, DeWoody JA (2007) Recent demographic bottlenecks are not accompanied by a genetic signature in banner-tailed kangaroo rats. Mol Ecol 16:2450–2462 doi:10.1111/j.1365-294X.2007.03283.x
Carrington M (1999) Recombination within the human MHC. Immunol Rev 167:245–256 doi:10.1111/j.1600-065X.1999.tb01397.x
Cocquet J, Chong A, Zhang GL, Veitia RA (2006) Reverse transcriptase template switching and false alternative transcripts. Genomics 88:127–131 doi:10.1016/j.ygeno.2005.12.013
Cutrera AP, Lacey EA (2007) Trans-species polymorphism and evidence of selection on class II MHC loci in tuco-tucos (Rodentia: Ctenomyidae). Immunogenetics 59:937–948 doi:10.1007/s00251-007-0261-3
de Groot N, Doxiadis GGM, de Groot NG, Otting N, Heijmans C, Rouweler AJ et al (2004) Genetic makeup of the DR region in rhesus macaques: gene content, transcripts, and pseudogenes. J Immunol 172:6152–6157
Debenham SL, Hart EA, Ashurst JL, Howe KL, Quail MA, Ollier WER et al (2005) Genomic sequence of the class II region of the canine MHC: comparison with the MHC of other mammalian species. Genomics 85:48–59 doi:10.1016/j.ygeno.2004.09.009
Doxiadis GGM, Otting N, De Groot NG, Bontrop RE (2001) Differential evolutionary MHC class II strategies in humans and rhesus macaques: relevance for biomedical studies. Immunol Rev 183:76–85 doi:10.1034/j.1600-065x.2001.1830106.x
Doxiadis GGM, Van Der Wiel MKH, Brok HPM, Groot NG, Otting N, 't Hart BA et al (2006) Reactivation by exon shuffling of a conserved HLA-DR3-like pseudogene segment in a new world primate species. Proc Natl Acad Sci U S A 103:5864–5868 doi:10.1073/pnas.0600643103
Figueroa F, Gutknecht J, Tichy H, Klein J (1990) Class II MHC genes in rodent evolution. Immunol Rev 113:27–46 doi:10.1111/j.1600-065X.1990.tb00035.x
Froeschke G, Sommer S (2005) MHC class II DRB variability and parasite load in the striped mouse (Rhabdomys pumilio) in the southern Kalahari. Mol Biol Evol 22:1254–1259 doi:10.1093/molbev/msi112
Gogolevsky KP, Kramerov DA (2006) Short interspersed elements (SINEs) of the Geomyoidea superfamily rodents. Gene 373:67–74 doi:10.1016/j.gene.2006.01.007
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nuc Acid Sym Ser 41:95–98
Harf R, Sommer S (2005) Association between major histocompatibility complex class II DRB alleles and parasite load in the hairy-footed gerbil, Gerbillurus paeba, in the southern Kalahari. Mol Ecol 14:85–91 doi:10.1111/j.1365-294X.2004.02402.x
Huchon D, Madsen O, Sibbald M, Ament K, Stanhope MJ, Catzeflis F et al (2002) Rodent phylogeny and a timescale for the evolution of Glires: evidence from an extensive taxon sampling using three nuclear genes. Mol Biol Evol 19:1053–1065
Huchon D, Chevret P, Jordan U, Kilpatrick CW, Ranwez V, Jenkins PD et al (2007) Multiple molecular evidences for a living mammalian fossil. Proc Natl Acad Sci U S A 104:7495–7499 doi:10.1073/pnas.0701289104
Hughes AL, Yeager M (1998) Natural selection at major histocompatibility complex loci of vertebrates. Annu Rev Genet 32:415–435 doi:10.1146/annurev.genet.32.1.415
IMGT/HLA Database. release 2.19.0 (October 2007). April 15, 2008
Jukes TH, Cantor CR (1969) Evolution of protein molecules. Academic, New York
Kaufman J, Salomonsen J, Flajnik MF (1994) Evolutionary conservation of MHC class I and class II molecules—different yet the same. Semin Immunol 6:411–424 doi:10.1006/smim.1994.1050
Kelley J, Walter L, Trowsdale J (2005) Comparative genomics of major histocompatibility complexes. Immunogenetics 56:683–695 doi:10.1007/s00251-004-0717-7
Kindt TJ, Singer DS (1987) Class I major histocombatibility complex genes in vertebrate species: what is the common denominator? Immunol Res 6:57–66 doi:10.1007/BF02918104
Klein J (1986) Natural history of the major histocompatibility complex. Wiley, New York
Klein J (1987) Origin of major histocompatibility complex polymorphism—the transspecies hypothesis. Hum Immunol 19:155–162 doi:10.1016/0198-8859(87)90066-8
Klein J, Bontrop RE, Dawkins RL, Erlich HA, Gyllensten UB, Heise ER et al (1990) Nomenclature for the major histocompatibility complexes of different species—a proposal. Immunogenetics 31:217–219
Li WH, Ellsworth DL, Krushkal J, Chang BHJ, Hewett-Emmett D (1996) Rates of nucleotide substitution in primates and rodents and the generation-time effect hypothesis. Mol Phyl Evol 5:182–187 doi:10.1006/mpev.1996.0012
Martin DP, Williamson C, Posada D (2005) RDP2: recombination detection and analysis from sequence alignments. Bioinformatics 21:260–262 doi:10.1093/bioinformatics/bth490
Maynard Smith J (1992) Analyzing the mosaic structure of genes. J Mol Evol 34:126–129
Meyer-Lucht Y, Sommer S (2005) MHC diversity and the association to nematode parasitism in the yellow-necked mouse (Apodemus flavicollis). Mol Ecol 14:2233–2243 doi:10.1111/j.1365-294X.2005.02557.x
Milinski M (2006) The major histocompatibility complex, sexual selection, and mate choice. Annu Rev Ecol Evol Syst 37:159–186 doi:10.1146/annurev.ecolsys.37.091305.110242
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Nigenda-Morales S, Flores-Ramirez S, Urban-R J, Vazquez-Juarez R (2008) MHC DQB1 polymorphism in the Gulf of California fin whale (Balaenoptera physalus) population. J Hered 99:14–21 doi:10.1093/jhered/esm087
Nizetic D, Figueroa F, Dembic Z, Nevo E, Klein J (1987) Major histocompatibility complex gene organization in the mole rat Spalax ehrenbergi—evidence for transfer of function between class II genes. Proc Natl Acad Sci U S A 84:5828–5832 doi:10.1073/pnas.84.16.5828
Oliver MK, Piertney SB (2006) Isolation and characterization of a MHC class II DRB locus in the European water vole (Arvicola terrestris). Immunogenetics 58:390–395 doi:10.1007/s00251-006-0121-6
Piertney SB, Oliver MK (2006) The evolutionary ecology of the major histocompatibility complex. Heredity 96:7–21
Posada D, Crandall KA (2001) Selecting the best-fit model of nucleotide substitution. Syst Biol 50:580–601 doi:10.1080/106351501750435121
Reusch TBG, Langefors A (2005) Inter- and intralocus recombination drive MHC class IIB gene diversification in a teleost, the threespined stickleback Gasterosteus aculeatus. J Mol Evol 61:531–541 doi:10.1007/s00239-004-0340-0
Richman AD, Herrera LG, Nash D (2002) Characterization of Peromyscus MHC class II beta sequences by ligation-anchored RT-PCR and denaturing gradient gel electrophoresis. Eur J Immunogenet 29:213–217 doi:10.1046/j.1365-2370.2002.00290.x
Richman AD, Herrera LG, Nash D, Schierup MH (2003) Relative roles of mutation and recombination in generating allelic polymorphism at an MHC class II locus in Peromyscus maniculatus. Genet Res 82:89–99 doi:10.1017/S0016672303006347
Robinson J, Waller MJ, Parham P, De Groot N, Bontrop R, Kennedy LJ et al (2003) IMGT/HLA and IMGT/MHC: sequence databases for the study of the major histocompatibility complex. Nucleic Acids Res 31:311–314 doi:10.1093/nar/gkg070
Rozas J, Sanchez-DelBarrio JC, Messeguer X, Rozas R (2003) DNA Sequence Polymorphism v.4.20.2. Free software distributed by author (http://www.ub.es/dnasp)
Schaschl H, Wandeler P, Suchentrunk F, Obexer-Ruff G, Goodman SJ (2006) Selection and recombination drive the evolution of MHC class II DRB diversity in ungulates. Heredity 97:427–437 doi:10.1038/sj.hdy.6800892
Schaschl H, Wegner KM (2007) Contrasting mode of evolution between the MHC class I genomic region and class II region in the three-spined stickleback (Gasterosteus aculeatus L.; Gasterosteidae: Teleostei). Immunogenetics 59:295–304 doi:10.1007/s00251-007-0192-z
Smulders MJM, Snoek LB, Booy G, Vosman B (2003) Complete loss of MHC genetic diversity in the common hamster (Cricetus cricetus) population in The Netherlands: consequences for conservation strategies. Con Gen 4:441–451 doi:10.1023/A:1024767114707
Strand T, Westerdahl H, Hoeglund J, Alatalo RV, Siitari H (2007) The MHC class II of the black grouse (Tetrao tetrix) consists of low numbers of B and Y genes with variable diversity and expression. Immunogenetics 59:725–734 doi:10.1007/s00251-007-0234-6
Swofford DL (1998) PAUP* Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sinauer, Sunderland
Triant DA, DeWoody JA (2006) Accelerated molecular evolution in Microtus (Rodentia) as assessed via complete mitochondrial genome sequences. Genetica 128:95–108 doi:10.1007/s10709-005-5538-6
von Salome J, Gyllensten U, Bergström TF (2007) Full-length analysis of the HLA-DRB1 locus suggests a recent origin of alleles. Immunogenetics 59:261–271 doi:10.1007/s00251-007-0196-8
Waser PM, Busch JD, McCormick CR, DeWoody JA (2006) Parentage analysis detects cryptic pre-capture dispersal in a philopatric rodent. Mol Ecol 15:1929–1937 doi:10.1111/j.1365-294X.2006.02893.x
Wilson and Reeder (2005) (eds) Mammal species of the world: a taxonomic and geographic reference, 3rd edn. The Johns Hopkins University Press, Baltimore
Worley K, Carey J, Veitch A, Coltman DW (2006) Detecting the signature of selection on immune genes in highly structured populations of wild sheep (Ovis dalli). Mol Ecol 15:623–637 doi:10.1111/j.1365-294X.2006.02829.x
Yauk CL, Bois PR, Jeffreys AJ (2003) High-resolution sperm typing of meiotic recombination in the mouse MHC E-beta gene. EMBO J 22:1389–1397 doi:10.1093/emboj/cdg136
Yuhki N, Beck T, Stephens RM, Nishigaki Y, Newmann K, O'Brien SJ (2003) Comparative genome organization of human, murine, and feline MHC class II region. Genet Res 13:1169–1179 doi:10.1101/gr.976103
Acknowledgements
We thank the members of the DeWoody lab, D. Bos, and J. Patton for their insight. J. McCreight provided assistance in DNA sequencing. Funding to J.D.B. was provided by the AMNH Theodore Roosevelt Memorial Fund, the American Society of Mammalogists, and Purdue University. J.A.D.’s lab is funded primarily by the National Science Foundation. This manuscript is ARP#2008-18322 from Purdue University.
Author information
Authors and Affiliations
Corresponding author
Additional information
Nucleotide sequence data reported are available in the DDBJ/EMBL/GenBank databases under the accession numbers EU817477–EU817485.
Rights and permissions
About this article
Cite this article
Busch, J.D., Waser, P.M. & DeWoody, J.A. Characterization of expressed class II MHC sequences in the banner-tailed kangaroo rat (Dipodomys spectabilis) reveals multiple DRB loci. Immunogenetics 60, 677–688 (2008). https://doi.org/10.1007/s00251-008-0323-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-008-0323-1