Abstract
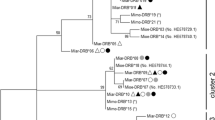
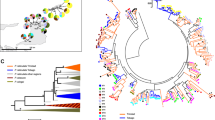
Major histocompatibility complex (MHC) genes play important role in host–parasite interactions and parasites are crucial factors influencing the population dynamics of hosts. We described the structure and diversity of exon 2 of the MHC class II DQA gene in three species of voles (Arvicolinae) exhibiting regular multi-annual fluctuations of population density and analysed the processes leading to the observed MHC polymorphism. By using cloning–sequencing methodology and capillary electrophoresis-single strand conformation polymorphism, we described seven sequences in the water, eight in the common, and seven in the bank voles coming from an area of 70 km2 around the Nozeroy canton in the Jura Mountains (Franche Comté, France). All exon 2 sequences translate to give unique amino acid sequences and positive selection was found to act very intensively on antigen binding sites. We documented the presence of recombination at vole DQA region but its importance in generating allelic polymorphism seems to be relatively limited. For the first time within rodents, we documented the duplication of the DQA gene in all three species with both copies being transcriptionally active. Phylogenetic analysis of allelic sequences revealed extensive trans-species polymorphism within the subfamily although no alleles were shared between species in our data set. We discuss possible role of parasites in forming the recent polymorphism pattern of the DQA locus in voles.



Similar content being viewed by others
References
Andersson L, Rask L (1988) Characterization of the MHC class II region in cattle. The number of DQ genes varies between haplotypes. Immunogenetics 27:110–120
Andersson G, Andersson L, Larhammar D, Rask L, Sigurdardottis S (1994) Simplifying genetic locus assignment of HLA-DRB genes. Immunol Today 15:58–62
Apanius V, Penn D, Slev PR, Ruff LR, Potts WK (1997) The nature of selection on the major histocompatibility complex. Crit Rev Immunol 17:179–224
Awadalla P, Charlesworth D (1999) Recombination and selection at Brassica self-incompatibility loci. Genetics 152:413–425
Ballingall KT, Luyai A, McKeever DJ (1997) Analysis of genetic diversity at the DQA loci in African cattle: evidence for a BoLA-DQA3 locus. Immunogenetics 46:237–244
Berthier K, Galan M, Foltete JC, Charbonnel N, Cosson JF (2005) Genetic structure of the cyclic fossorial water vole (Arvicola terrestris): landscape and demographic influences. Mol Ecol 14:2861–2871
Blake JA, Richardson JE, Bult CJ, Kadin JA, Eppig JT, Mouse Genome Database Group (2003) MGD: the mouse genome database. Nucleic Acids Res 31:193–195
Bowen L, Aldridge BM, Gulland F, Van Bonn W, DeLong R, Melin S, Lowenstine LJ, Stott JL, Johnson ML (2004) Class II multiformity generated by variable MHC-DRB region configurations in the California sea lion (Zalophus californianus). Immunogenetics 56:12–27
Brown JH, Jardetzky TS, Gorga JC, Stern LJ, Urban RG, Strominger JL, Wiley DC (1993) Three-dimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature 64:33–39
Bryja J, Galan M, Charbonnel N, Cosson JF (2005) Analysis of major histocompatibility complex class II gene in water voles using capillary electrophoresis-single stranded conformation polymorphism. Mol Ecol Notes 5:173–176
Butet A, Spitz F (2001) Cyclic fluctuations of microtine populations: half a century of research. Rev Ecol (Terre Vie) 56:353–372
Cavanagh RD, Lambin X, Ergon T, Bennett M, Graham IM, van Soolingen D, Begon M (2004) Disease dynamics in cyclic populations of field voles (Microtus agrestis): cowpox virus and vole tuberculosis (Mycobacterium microti). Proc R Soc Lond B 271:859–867
Easterfield AJ, Bradley JA, Bolton EM (2003) Complementary DNA sequences encoding the rat MHC class II RT1-Bu and RT1-Du α and β chains. Immunogenetics 55:344–350
Edwards SV, Wakeland EK, Potts WK (1995) Contrasting histories of avian and mammalian MHC genes revealed by class II B sequences from songbirds. Proc Natl Acad Sci U S A 92:12200–12204
Edwards SV, Chesnut K, Satta Y, Wakeland EK (1997) Ancestral polymorphism of Mhc class II genes in mice: implications for balancing selection and the mammalian molecular clock. Genetics 146:655–668
Fabb SA, Maddox JF, Gogolin EKJ, Baker L, Wu MJ, Brandon MR (1993) Isolation, characterization and evolution of ovine major histocompatibility complex class II DRA and DQA genes. Anim Genet 24:249–255
Figueroa F, Gunther E, Klein J (1988) MHC polymorphism pre-dating speciation. Nature 355:265–267
Fremont DH, Monnaie D, Nelson CA, Hendrickson WA, Unanue ER (1998) Crystal structure of I-Ak in complex with a dominant epitope of lysozyme. Immunity 8:305–317
Gill TJ (1992) Influence of MHC and MHC-linked genes on reproduction. Am J Hum Genet 50:1–5
Godot V, Harraga S, Beurton I, Tiberghien P, Sarciron E, Gottstein B, Vuitton DA (2000) Resistance/susceptibility to Echinococcus multilocularis infection and cytokine profile in humans. II. Influence of the HLA B8, DR3, DQ2 haplotype. Clin Exp Immunol 121:491–498
de Bellocq JG, Delarbre C, Gachelin G, Morand S (2005) Allelic diversity at the Mhc-DQA locus of woodmouse populations (Apodemus sylvaticus) present in the islands and mainland of the northern Mediterranean. Glob Ecol Biogeogr 14:115–122
Günther E, Walter L (2001) The major histocompatibility complex of the rat (Rattus norvegicus). Immunogenetics 53:520–542
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Harf R, Sommer S (2005) Association between major histocompatibility complex class II DRB alleles and parasite load in the hairy-footed gerbil, Gerbillurus paeba, in the southern Kalahari. Mol Ecol 14:85–91
Haukisalmi V, Henttonen H (2003) What is Paranoplocephala macrocephala (Douthitt, 1915) (Cestoda: Anoplocephalidae)? Syst Parasitol 54:53–69
Haukisalmi V, Wixkström LM, Henttonen H, Hantula J, Gubanyi HA (2004) Molecular and morphological evidence for multiple species within Paranoplocephala omphalodes (Cestoda, Anoplocephalidae) in Microtus voles (Arvicolinae). Zool Scr 33:277–290
Hedrick PW, Lee RN, Parker KM (2000) Major histocompatibility complex (MHC) variation in the endangered Mexican wolf and related canids. Heredity 85:617–624
Hedrick PW, Parker KM, Gutierrez-Espeleta GA, Rattink A, Lievers K (2000) Major histocompatibility complex variation in the Arabian oryx. Evolution 54:2145–2151
Higasa K, Kukita Y, Baba S, Hayashi K (2002) Software for machine-independent quantitative interpretation of SSCP in capillary array electrophoresis (QUISCA). Biotechniques 33:1342–1348
Hohenhaus MA, Outteridge PM (1995) The immunogenetics of resistance to Trichostrongylus colubriformis and Haemonchus contortus parasites in sheep. Br Vet J 151:119–140
Hudson RR (2001) Two-locus sampling distributions and their application. Genetics 159:1805–1817
Hughes AL (1994) The evolution of functionally novel proteins after gene duplication. Proc R Soc Lond B 256:119–124
Hughes AL, Nei M (1990) Evolutionary relationships of class II MHC genes in mammals. Mol Biol Evol 7:491–514
Hughes AL, Yeager M (1998) Natural selection at major histocompatibility complex loci of vertebrates. Annu Rev Genet 32:415–435
Hughes AL, Yeager M (1998) Natural selection and the evolutionary history of MHC loci. Front Biosci 3:509–516
Hughes AL, Hughes MK, Howell CY, Nei M (1994) Natural selection at the class II major histocompatibility complex loci of mammals. Philos Trans R Soc Lond B 346:359–366
Hurt P, Walter L, Sudbrak R, Klages S, Müller I, Shiina T, Inoko H, Lehrach H, Günther E, Reinhardt R, Himmelbauer H (2004) The genomic sequence and comparative analysis of the rat major histocompatibility complex. Genome Res 14:631–639
Jaarola M, Martinkova N, Gunduz I, Brunhoff C, Zima J, Nadachowski A, Amori G, Bulatova NS, Chondropoulos B, Fraguedakis-Tsolis S, Gonzalez-Esteban J, Lopez-Fuster MJ, Kandaurov AS, Kefelioglu H, Mathias MD, Villate I, Searle JB (2004) Molecular phylogeny of the speciose vole genus Microtus (Arvicolinae, Rodentia) inferred from mitochondrial DNA sequences. Mol Phylogenet Evol 33:647–663
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic, New York, pp 21–132
Karlin S, Altschul SF (1990) Methods for assessing the statistical significance of molecular sequences by using general scoring schemes. Proc Natl Acad Sci U S A 87:2264–2268
Kennedy LJ, Ryvar R, Gaskell RM, Addie DD, Willoughby K, Carter SD, Thomson W, Ollier WER, Radford AD (2002) Sequence analysis of MHC DRB alleles in domestic cats from the United Kingdom. Immunogenetics 54:348–352
Khazand M, Peiberg C, Nagy M, Sauermann U (1999) Mhc-DQ-DRB haplotype analysis in the rhesus macaque: evidence for a number of different haplotypes displaying a low allelic polymorphism. Tissue Antigens 54:615–624
Klein J (1986) Natural history of the major histocompatibility complex. Wiley, New York
Klein J (1987) Origin of major histocompatibility complex polymorphism: the trans-species hypothesis. Hum Immunol 19:155–162
Klein J, O’Huigin C (1994) MHC polymorphism and parasites. Philos Trans R Soc Lond B 346:351–358
Klein J, Bontrop RE, Dawkins RL, Erlich HA, Gyllensten UB, Heise ER, Jones PP, Parham P, Wakeland EK, Watkins DI (1990) Nomenclature for the major histocompatibility complexes of different species: a proposal. Immunogenetics 31:217–219
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Latek RR, Suri A, Petzold SJ, Nelson CA, Kanagawa O, Unanue ER, Fremont DH (2000) Structural basis of peptide binding and presentation by the type I diabetes-associated MHC class II molecule of NOD mice. Immunity 12:699–710
Martin Y, Gerlach G, Schlötterer C, Meyer A (2000) Molecular phylogeny of European muroid rodents based on complete cytochrome b sequences. Mol Phylogenet Evol 16(1):37–47
Martinsohn JT, Sousa AB, Guethlein LA, Howard JC (1999) The gene conversion hypothesis of MHC evolution: a review. Immunogenetics 50:168–200
McVean G, Awadalla P, Fearnhead P (2002) A coalescent-based method for detecting and estimating recombination from gene sequences. Genetics 160:1231–1241
Meyer-Lucht Y, Sommer S (2005) MHC diversity and the association to nematode parasitism in the yellow-necked mouse (Apodemus flavicollis). Mol Ecol 14:2233–2243
Michaux J, Catzeflis F (2000) The bushlike radiation of muroid rodents is exemplified by the molecular phylogeny of the LCAT nuclear gene. Mol Phylogenet Evol 17:280–293
Musolf K, Meyer-Lucht Y, Sommer S (2004) Evolution of MHC-DRB class II polymorphism in the genus Apodemus and a comparison of DRB sequences within the family Muridae (Mammalia: Rodentia). Immunogenetics 56:420–426
Musser GG, Carleton MD (1993) Family Muridae. In: Wilson DE, Reeder DM (eds) Mammal species of the world: a taxonomic and geographic reference, 2nd edn. Smithsonian Institution, Washington, pp 501–755
Nei M, Gojobori T (1986) Simple method for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Nei M, Gu X, Sitnikova T (1997) Evolution by the birth-and-death process in multigene families of the vertebrate immune system. Proc Natl Acad Sci U S A 94:7799–7806
Ohta T (1999) Effect of gene conversion on polymorphic patterns at major histocompatibility complex loci. Immunol Rev 167:319–325
Otting N, de Groot NG, Doxiadis GGM, Bontrop RE (2002) Extensive Mhc-DQB variation in humans and non-human primate species. Immunogenetics 54:230–239
Ottova E, Simkova A, Martin JF, Gouy de Bellocq J, Gelnar M, Allienne JF, Morand S (2005) Evolution and trans-species polymorphism of MHC class IIβ genes in cyprinid fish. Fish Shellfish Immunol 18:199–222
Paterson S, Wilson K, Pemberton JM (1998) Major histocompatibility complex variation associated with juvenile survival and parasite resistance in a large unmanaged ungulate population. Proc Natl Acad Sci U S A 95:3714–3719
Pfau RS, Van Den Bussche RA, McBee K, Lochmiller RL (1999) Allelic diversity at the Mhc-DQA locus in cotton rats (Sigmodon hispidus) and a comparison of DQA sequences within the family Muridae (Mammalia: Rodentia). Immunogenetics 49:886–893
Pfau RS, Van Den Bussche RA, McBee K (2001) Population genetics of the hispid cotton rat (Sigmodon hispidus): patterns of genetic diversity at the major histocompatibility complex. Mol Ecol 10:1939–1945
Posada D (2002) Evaluation of methods for detecting recombination from DNA sequences: empirical data. Mol Biol Evol 19:708–717
Quinnell RJ (2003) Genetics of susceptibility to human helminth infection. Int J Parasitol 33:1219–1231
Reusch TBH, Schaschl H, Wegner KM (2004) Recent duplication and inter-locus gene conversion in major histocompatibility class II genes in a teleost, the three-spined stickleback. Immunogenetics 56:427–437
Richman AD, Herrera LG, Nash D (2002) Characterization of Peromyscus MHC class II beta sequences by ligation-anchored RT-PCR and denaturing gradient gel electrophoresis. Eur J Immunogenet 29:213–217
Richman AD, Herrera LG, Nash D (2003a) Evolution of MHC class II Eβ diversity within the genus Peromyscus. Genetics 164:289–297
Richman AD, Herrera LG, Nash D, Schierup MH (2003b) Relative roles of mutation and recombination in generating allelic polymorphism at an MHC class II locus in Peromyscus maniculatus. Genet Res 82:89–99
Robinson J, Waller MJ, Stoehr P, Marsh SGE (2005) IPD—the immuno polymorphism database. Nucleic Acids Res 331:523–526
Roitt IM (1997) Essential immunology, 9th edn. Blackwell Science, Oxford
Sauermann U, Nurnberg P, Bercovitch FB, Berard JD, Trefilov A, Widdig A, Kessler M, Schmidtke J, Krawczak M (2001) Increased reproductive success of MHC class II heterozygous males among free-ranging rhesus macaques. Hum Genet 108:249–254
Sawyer SA (1999) GENECONV: a computer package for statistical detection of gene conversion. Available at http://www.math.wustl.edu/∼sawyer/mbprogs/
Schad J, Ganzhorn JU, Sommer S (2005) Parasite burden and constitution of major histocompatibility complex in the Malagasy mouse lemur, Microcebus murinus. Evolution 59:439–450
Schaschl H, Suchentruk F, Hammer S, Goodman SJ (2005) Recombination and the origin of sequence diversity in the DRB MHC class II locus in chamois (Rupicapra spp.). Immunogenetics 57:108–115
Schierup MH, Hein J (2000) Consequences of recombination on traditional phylogenetic analysis. Genetics 156:879–891
Scott CA, Peterson PA, Teyton L, Wilson IA (1998) Crystal structures of two I-Ad–peptide complexes reveal that high affinity can be achieved without large anchor residues. Immunity 8:319–329
Seddon JM, Baverstock PR (1998) Eight rat RT1Ba sequences. Immunogenetics 48:161–162
Seddon JM, Baverstock PR (1999) Variation on islands: major histocompatibility complex (Mhc) polymorphism in populations of the Australian bush rat. Mol Ecol 8:2071–2079
Seddon JM, Baverstock PR (2000) Evolutionary lineages of RT1.Ba in the Australian Rattus. Mol Biol Evol 17:768–772
Seddon JM, Ellegren H (2002) MHC class II genes in European wolves: a comparison with dogs. Immunogenetics 54:490–500
Smulders MJM, Snoek LB, Booy G, Vosman B (2003) Complete loss of MHC genetic diversity in the common hamster (Cricetus cricetus) populations in The Netherlands. Consequences for conservation strategies. Cons Gen 4:441–451
Snibson KJ, Maddox JF, Fabb SA, Brandon MR (1998) Allelic variation of ovine MHC class II DQA1 and DQA2 genes. Anim Genet 29:356–362
Sommer S, Tichy H (1999) Major histocompatibility complex (MHC) class II polymorphism and paternity in the monogamous Hypogeomys antimena, the endangered, largest endemic Malagasy rodent. Mol Ecol 8:1259–1272
Sommer S, Schwab D, Ganzhorn JU (2002) MHC diversity of endemic Malagasy rodents in relation to geographic range and social system. Behav Ecol Sociobiol 51:214–221
Steppan SJ, Adkins RM, Anderson J (2004) Phylogeny and divergence-date estimates of rapid radiations in muroid rodents based on multiple nuclear genes. Syst Biol 53:533–553
Takahata N, Nei M (1990) Allelic genealogy under overdominant and frequency-dependent selection and polymorphism of major histocompatibility complex loci. Genetics 124:967–978
The MHC sequencing consortium (1999) Complete sequence and gene map of a human major histocompatibility complex. Nature 401:921–923
Vestberg M, Brunsberg U, Bergsteinsdottir K, Karlsson M, Gustafsson K, Wedekind D, Hedrich H, Holmdahl R (1998) Limited polymorphism in the first domain of the rat MHC class II RT1-D molecule. Immunogenetics 48:344–349
Waine GJ, Ross AG, Williams GM, Sleigh AC, McManus DP (1998) HLA class II antigens are associated with resistance or susceptibility to hepatosplenic disease in a Chinese population infected with Schistosoma japonicum. Int J Parasitol 28:537–542
Wittzell H, Madsen T, Westerdahl H, Shine R, von Schantz T (1999) MHC variation in birds and reptiles. Genetica 104:301–309
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13:555–556
Yang ZH, Swanson WJ, Vacquier VD (2000) Maximum-likelihood analysis of molecular adaptation in abalone sperm lysin reveals variable selective pressures among lineages and sites. Mol Biol Evol 17:1446–1455
Yang ZH, Wong WSW, Nielsen R (2005) Bayes empirical Bayes inference of amino acid sites under positive selection. Mol Biol Evol 22:1107–1118
Acknowledgements
The authors acknowledge the assistance of Julie Deter and Yannick Chaval in the provision of samples, Serge Morand for access to his database on rodent parasites and three anonymous referees for useful comments on the manuscript. This work received financial support from the Institut National de la Recherche Agronomique, the Region of Franche-Comté, the DIREN Franche-Comté, and the Ministry of the Environment of France. This publication has been also partially funded under the EU 6th Framework Program (GOCE-CT-2003-010284 Emerging Diseases in a Changing European Environment (EDEN) and is officially catalogued by the EDEN Steering Committee as EDEN0006). Its content does not represent the official position of the European Commission and is entirely under the responsibility of the authors.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bryja, J., Galan, M., Charbonnel, N. et al. Duplication, balancing selection and trans-species evolution explain the high levels of polymorphism of the DQA MHC class II gene in voles (Arvicolinae). Immunogenetics 58, 191–202 (2006). https://doi.org/10.1007/s00251-006-0085-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-006-0085-6