Abstract
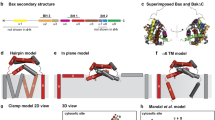
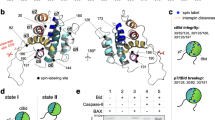
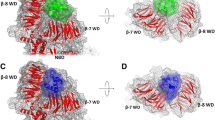
Bid, a BH3-only pro-apoptopic member of the BCL-2 protein family, regulates cell death at the level of mitochondrial cytochrome c efflux. Bid consists of 8 α-helices (H1–H8, respectively) and is soluble cytosolic protein in its native state. Proteolysis of the N-terminus (encompassing H1 and H2) of Bid by caspase 8 in apoptosis yields activated “tBid” (truncated Bid), which translocates to the mitochondria and induces the efflux of cytochrome c. The release of cytochrome c from mitochondria to the cytosol constitutes a critical control point in apoptosis that is regulated by interaction of tBid protein with mitochondrial membrane. tBid displays structural homology to channel-forming bacterial toxins, such as colicins or transmembrane domain of diphtheria toxin. By analogy, it has been hypothesized that tBid would unfold and insert into the lipid bilayer of the mitochondria outer membrane (MOM) upon membrane association. However, it has been shown recently that unlike colicins and the transmembrane domain of diphtheria toxin, tBid binds to the lipid bilayer maintaining α-helical conformation of its helices without adopting a transmembrane orientation by them. Here, the mechanism of the association of tBid with the model membrane mimicking the mitochondrial membrane is studied by Monte Carlo simulations, taking into account the underlying energetics. A novel two-stage hierarchical simulation protocol combining coarse-grained discretization of conformational space with subsequent refinements was applied which was able to generate the protein conformation and its location in the membrane using modest computational resources. The simulations show that starting from NMR-established conformation in the solution, the protein associates with the membrane without adopting the transmembrane orientation. The configuration (conformation and location) of tBid providing the lowest free energy for the system protein/membrane/solvent has been obtained. The simulations reveal that tBid upon association with the membrane undergoes significant conformational changes primarily due to rotations within the loops between helices H4 and H5, H6 and H7, H7 and H8. It is established that in the membrane-bound state of tBid-monomer helices H3 and H5 have the locations exposed to the solution, helices H6 and H8 are partly buried and helices H4 and H7 are buried into the membrane at shallow depth. The average orientation of tBid bound to the membrane in the most stable configuration reported here is in satisfactory agreement with the evaluations obtained by indirect experimental means.





Similar content being viewed by others
References
Adams JM (2003) Ways of dying: multiple pathways of apoptosis. Genes Dev 172:2481–2495
Allen MP, Tildesley DJ (1987) Computer simulations of liquids. Oxford, Clarendon
Annis MG, Soucie EL, Dlugosz PJ, Cruz-Aguado JC, Penn LZ, Leber B, Andrews DW (2005) Bax forms multispanning monomers that oligomerize to permeabilize membranes during apoptosis. EMBO J 24:2096–2103
Antonsson B, Montessuit S, Lauper S, Eskes R, Martinou JC (2000) Bax oligomerization is required for channel-forming activity in liposomes and to trigger cytochrome c release from mitochondria. Biochem J 345:271–278
Ardail D, Privat J-P, Egret-Charlier M, Levrat C, Lerme F, Louisot P (1990) Mitochondrial contact sites. Lipid composition and dynamics. J Biol Chem 265:18797–18802
Ash WL, Zlomislic MR, Oloo EQ, Tieleman DP (2004) Computer simulations of membrane proteins. Biochim Biophys Acta 1666:158–189
Basanez G, Sharpe JC, Galanis J, Brandt TB, Hardwick JM, Zimmerberg J (2002) Bax-type apoptotic proteins porate pure lipid bilayers through a mechanism sensitive to intrinsic monolayer curvature. J Biol Chem 277:49360–49365
Baumgartner A (1996) Insertion and hairpin formation of membrane proteins: a Monte Carlo study. Biophys J 71:1248–1255
Belzacq AS, Vieira HL, Verrier F, Vandescasteele G, Cohen I, Prevost MC, Larquet E, Pariselli F, Petit PX, Kahn A, Rizzutto R, Brenner C, Kroemer C (2003) Bcl-2 and Bax modulate adenine nucleotides translocase activity. Cancer Res 63:541–546
Benz R, Kottke M, Brdiczka D (1990) The cationically selective state of the mitochondrial outer membrane pore: a study with intact mitochondria and reconstituted mitochondrial porin. Biochim Biophys Acta 1022:311–318
Bond PJ, Sansom MS (2004) The simulation approach to bacterial outer membrane proteins. Mol Membr Biol 21:151–161
Bond PJ, Sansom MS (2006) Insertion and assembly of membrane protein via simulation. J Am Chem Soc 128:2697–2704
Bychkova VE, Dujsekina AE, Klenin SI, Tiktopulo EI, Uversky VN, Ptitsyn OB (1996) Molten globule like state of cytochrome c under conditions simulating those near the membrane surface. Biochemistry 35:6058–6063
Chenal A, Savarin P, Nizard P, Guillain F, Gillet D, Forge V (2002) Membrane protein insertion regulated by bringing electrostatic and hydrophobic interactions into play. A case study with the translocation domain of diphtheria toxin. J Biol Chem 277:43425–43432
Choe S, Bennett MJ, Fujii G, Curmi PM, Kantardjieff KA, Collier RJ, Eisenberg D (1992) The crystal structure of diphtheria toxin. Nature 357:216–222
Chou J, Li H, Salvesen G, Yuan J, Wagner G (1999) Solution structure of Bid, an intracellular amplifier of apoptopic signaling. Cell 96:615–624
Chang G, Guida WC, Still WC (1989) An internal coordinate Monte-Carlo method for searching conformational space. J Am Chem Soc 111:4379–4386
Cory S, Huang DC, Adams JM (2003) The Bcl-2 family: roles in cell survival and oncogenesis. Oncogene 22:8590–8607
Cramer WA, Heimann JB, Shendel SL, Deriy BN, Cohen FS, Etkins PA, Stauffacher CV (1995) Structure-function of the channel-forming colicins. Annu Rev Biophys Biomol Struct 24:611–641
Daniel NN, Korsmeyer SJ (2004) Cell death: critical control points. Cell 116:205–219
Desagher S, Osen-Sand A, Nichols A, Eskes R, Montessuit S, Lauper S, Maundrell K, Antonsson B, Martinou JC (1999) Bid-induced conformational change of Bax is responsible for mitochondrial cytochrome c release during apoptosis. J Cell Biol 144:891–901
Ducarme P, Rahman M, Brasseur R (1998) IMPALA: a simple restraint field to simulate the biological membranes in molecular structure studies. Proteins 30:357–371
Dunfield LG, Burgess AW, Scheraga HA (1978) Energy parameters in polypeptides. 8. Empirical potential energy algorithm for the conformational analysis of large molecules. J Phys Chem 82:2609–2616
Efremov RG, Nolde DE, Vergoten G, Arseniev AS (1999a) A solvent model for simulations of peptides in bilayers. I. Membrane-promoting α-helix formation. Biophys J 76:2448–2459
Efremov RG, Nolde DE, Vergoten G, Arseniev AS (1999b) A solvent model for simulations of peptides in bilayers. II. Membrane-spanning α-helices. Biophys J 76:2460–2471
Efremov RG, Volynsky PE, Nolde DE, Dubovskii PV, Arseniev AS (2002) Interaction of cardiotoxins with membranes: a molecular modeling study. Biophys J 83:144–153
Epand RM, Vogel HJ (1999) Diversity of antimicrobial peptides and their mechanisms of ac tion. Biochim Biophys Acta 1462:11–28
Epand RF, Martinou J-C, Fornallaz-Mulhauser M, Hughes DW, Epand RM (2002a) The apoptotic protein tBid promotes leakage by altering membrane curvature. J Biol Chem 277:32632–32639
Epand RF, Martinou J-C, Montessuit S, Epand RM, Yip CM (2002b) Direct evidence for membrane pore formation by the apoptotic protein Bax. Biochem Biophys Res Commun 298:744–749
Eskes R, Desagher S, Antonsson B, Martinou J-C (2000) Bid induces the oligomerization and insertion of BAX into the outer mitochondrial membrane. Mol Cell Biol 20:929–935
Forsten KE, Kosack RE, Lauffenburger DA, Subramanian (1994) Numerical solution of nonlinear Poisson–Boltzmann equation for a membrane electrolyte system. J Phys Chem 98:5580–5586
Fraczkiewicz R, Braun W (1998) Exact and efficient analytical calculation of the accessible surface areas and their gradients for macromolecules. J Comp Chem 19:319–333
Franzin CM, Choi J, Zhai J, Reed D, Marassi FM (2004) Structural studies of apoptosis and ion transport regulatory proteins in membranes. Magn Reson Chem 42:172–179
Garcia-Saez AJ, Mingarro I, Perez-Paya E, Salgado J (2004) Membrane-insertion fragments of Bcl-XL, Bax, and Bid. Biochemistry 43:10930–10943
Garcia-Saez A, Coraiola M, Dalla Serra M, Mingarro I, Muller P, Salgado J (2006) Peptides corresponding to helices 5 and 6 of BAX can independently form lipid pores. FEBS J 273:971–981
Gong X-M, Choi J, Franzin CM, Zhai D, Reed JC, Marassi FM (2004) Conformation of membrane-associated proapoptopic tBid. J Biol Chem 279:28954–28960
Grinberg M, Sarig R, Zaltsman Y, Frumkin D, Grammatikakis N, Reuveny E, Gross A (2002) tBID homooligomerizes in the mitochondrial membrane to induce apoptosis. J Biol Chem 277:12237–12245
Gross A, McDonnel JM, Korsemeyer SJ (1999a) BCL-2 family members and the mitochondria in apoptosis. Genes Dev 13:1899–1911
Gross A, Yin XM, Wang K, Wei MC, Jockel J, Milliman C, Erdjument-Bromage H, Tempst P, Korsemeyer SJ (1999b) Caspase cleaved BID targets mitochondria and is required for cytochrome c release, while BCL-XL prevents this release but not tumor necrosis factor R1/FAS death. J Biol Chem 274:1156–1163
Gumbart J, Wang Y, Aksimentiev A, Tajkhotshid E, Schulten K (2005) Molecular dynamics simulations of proteins in lipid bilayers. Curr Opin Struct Biol 15:423–431
Holm L, Park J (2000) DaliLite workbench for protein structure comparison. Bioinformatics 16:566–567
Huang HW, Chen FY, Lee MT (2004) Molecular mechanism of peptide-induced pores in membranes. Phys Rev Lett 92:198304
Huang HW (2006) Molecular mechanism of antimicrobial peptides. The origin of cooperativity. Biochim Biophys Acta 1758:1292–1302
Kessel A, Shental-Bechor D, Haliloglu T, Ben-Tal N (2003) Interaction of hydrophobic peptides with lipid bilayers: Monte Carlo simulations with M2δ. Biophys J 85:3431–3444
Kim TH, Zhao Y, Ding WX, Shin JN, He X, Seo YW, Chen J, Rabinowich H, Amoscato AA, Yin XM (2004) Bid-cardiolipin interaction at Mitochondrial contact site contributes to mitochondrial cristae reorganization and cytochrome c release. Mol Biol Cell 15:3061–3072
Kirkpatrick S, Gelati CD, Vecchi MP (1983) Optimization by simulated annealing. Science 220:671–680
Korsmeyer SJ, Wei MC, Saito M, Weiler S, Oh KJ, Schlesinger PH (2000) Pro-apoptopic cascade activates BID, which oligomerizes BAK or BAX into pores that result in the release of cytochrome c. Cell Death Differ 7:1166–1173
Kudla G, Montessuit S, Eskes R, Berrier C, Martinou JC, Gazi A, Antonsson B (2000) The destabilization of lipid membranes induced by the C-terminal fragment of caspase 8-cleaved bid is inhibited by the N-terminal fragment. J Biol Chem 275:22713–22718
Kuwana T, Mackey MR, Perkins G, Ellisman MH, Latterich M, Schneiter R, Green DR, Newmeyer DD (2002) Bid, Bax, and lipids cooperate to form supramolecular openings in the outer mitochondrial membrane. Cell 111:331–342
Lazaridis T (2005) Implicit solvent simulations of peptide interactions with anionic lipid membranes. Proteins 58:518–527
Lesieur C, Vecsey-Semjen B, Abrami L, Fivaz M, van der Goot FG (1997) Membrane insertion: the strategies of toxins. Mol Membr Biol 14:45–64
Lee J, Scheraga HA, Rackovsky S (1997) New optimization method for conformational energy calculations on polypeptides: conformational space annealing. J Comput Chem 18:1222–1232
Letai A, Bassik MC, Walensky LD, Sorcinelli MD, Weiler S, Korsmeyer SJ (2002) Distinct BH3 domains either sensitize or activate mitochondrial apoptosis, serving as prototype cancer therapeutics. Cancer Cell 2:183–192
Li Z, Scheraga HA (1987) Monte Carlo-minimization approach to the multiple-minima problem in protein folding. Proc Natl Acad Sci USA 84:6611–6615
Li H, Zhu H, Xu CJ, Yuan J (1998) Cleavage of Bid by caspase 8 mediates the mitochondrial damage in the Fas pathway of apoptosis. Cell 94:491–501
London E (1992) Diphtheria toxins: membrane interaction and membrane translocation. Biochim Biophys Acta 1113:25–51
Lu JX, Damodaran K, Blazyk J, Lorigan GA (2005) Solid-state nuclear magnetic resonance relaxation studies of the interaction mechanism of antimicrobial peptides with phospholipid bilayer membranes. Biochemistry 44:10208–10217
Luo X, Budihardjo I, Zou H, Slaughter C, Wang X (1998) Bid, a Bcl-2 interacting protein, mediates cytochrome c release from mitochondria in response to activation of cell surface death receptors. Cell 94:481–490
Lutter M, Fang M, Lun X, Nishijima M, Xie M, Wang X (2000) Cardiolipin provides specificity for targeting of tBid to mitochondria. Nat Cell Biol 2:754–761
Maddox MW, Longo ML (2002) A Monte Carlo study of peptide insertion into lipid bilayers: equilibrium conformations and insertion mechanisms. Biophys J 82:244–263
Matsuzaki K (1999) Why and how are peptide-lipid interactions utilized for self-defense? Magainins and tacheplysins as archetypes. Biochim Biophys Acta 1452:1–10
Mau NV, Kajava AV, Bonfils C, Martinou J-C, Harricane M-C (2006) Interactions of Bax and tBid with lipid monolayers. J Membr Biol 207:1–9
McDonnel J, Fushman D, Milliman C, Korsmeyer S, Cowburn D (1999) Solution structure of the proapoptopic molecule BID: a structural basis for apoptopic agonists and antagonists. Cell 96:625–634
McLaughlin S (1989) The electrostatic properties of membranes. Annu Rev Biophys Biophys Chem 18:113–136
Metropolis N, Rosenbluth AW, Rosenbluth MN, Teller AH, Teller EJ (1953) Equation of state calculations by fast computing machines. J Chem Phys 21:1087–1092
Milik M, Skolnick J (1993) Insertion of peptide chains into lipid membranes: an off-lattice Monte Carlo dynamics model. Proteins 15:10–25
Milik M, Skolnick J (1995) Monte Carlo model of FD and PF1 coat proteins in lipid membranes. Biophys J 69:1382–1386
Muchmore SW, Sattler M, Liang H, Meadows RP, Harian JE, Yoon JE, Nettesheim D, Chang BS, Thompson CB, Wong SL, Ng SL, Fesic SW (1996) X-ray and NMR structure of human Bcl-xL, an inhibitor of programmed cell death. Nature 381:335–341
Mungikar AA, Forciniti D (2004) Conformational changes of peptides at solid/liquid interfaces: a Monte Carlo study. Biomacromolecules 5:2147–2159
Nam GH, Choi KY (2002) Association of human tumor necrosis factor-related apoptosis inducing ligand with membrane upon acidification. Eur J Biochem 269:5280–5287
Nelson AP, Colonomos P, McQuarrie DA (1975) Electrostatic coupling across a membrane with titratable surface groups. J Theor Biol 50:317–325
Némethy G, Pottle MS, Scheraga HA (1983) Energy parameters in polypeptides. 9. Updating of geometrical parameters, nonbonded interactions, and hydrogen bond interactions for the naturally occurring amino acids. J Phys Chem 87:1883–1887
Némethy G, Gibson KD, Palmer KA, Yoon CN, Paterlini G, Zagari A, Rumsey S, Scheraga HA (1992) Energy parameters in polypeptides. 10. Improved geometrical parameters and nonbonded interactions for use in ECEPP/3 algorithm with application to proline-containing peptides J Phys Chem 96:6472–6484
Oh KJ, Barbuto S, Meyer N, Kim R-S, Collier RJ, Korsmeyer SJ (2005) Conformational changes in BID, a pro-apoptopic BCL-2 family member, upon membrane binding. J Biol Chem 280:753–767
Opferman JT, Korsmeyer SJ (2003) Apoptosis in the development and maintenance of the immune system. Nat Immunol 4:410–415
Oshima H, Kondo T (1988) Membrane potential and Donnan potential. Biophys Chem 29:277–281
Ozkan SB, Meirovitch H (2004) Conformational search of peptides and proteins: Monte Carlo minimization with an adaptive bias method applied to the heptapeptide deltorphin. J Comput Chem 25:565–572
Peitzsch RM, Eisenberg M, Sharp KA, McLaughlin S (1995) Calculations of the electrostatic potential adjacent to model phospholipid bilayers. Biophys J 68:729–738
Petros AM, Oleiniczak ET, Fesik SW (2004) Structural biology of the Bcl-2 family of proteins. Biochim Biophys Acta 1644:83–94
Pillardy J, Czaplewski C, Wedemeyer WJ, Scheraga HA (2000) Conformation-family Monte Carlo (CFMC): an efficient computational method for identifying the low-energy states of a macromolecule. Helv Chim Acta 83:2214–2230
Pillardy J, Arnautova YA, Czaplewski C, Gibson KD, Scheraga HA (2001) Conformation-family Monte Carlo: a new method for crystal structure prediction. Proc Natl Acad Sci USA 98:12351–12356
Rosconi MP, Zhao G, London E (2004) Analyzing topography of membrane-inserted diphtheria toxin T domain using BODIPY-streptavidin: at low pH, helices 8 and 9 form a transmembrane hairpin but helices 5–7 form stable nonclassical inserted segments on the cis side of the bilayer. Biochemistry 43:9127–9139
Roseman MA (1988) Hydrophobicity of polar amino acid side chains is markedly reduced by flanking peptide bonds. J Mol Biol 200:513–522
Schendel SL, Azimov R, Pawlowski K, Godzik A, Kagan BL, Reed JC (1999) Ion channel activity of the BH3 only BCL-2 family member. BID J Biol Chem 274:21932–21936
Schlame M, Rua D, Greenberg ML (2000) The biosynthesis and functional role of cardiolipin. Prog Lipid Res 39:257–288
Scorrano L, Ashiya M, Buttle K, Weiler S, Oakes SA, Mannela CA, Korsmeyer SJ (2002) A distinct pathway remodels mitochondrial cristae and mobilizes cytochrome c during apoptosis. Dev Cell 2:55–67
Sintes T, Baumgartner A (1998) Membrane-mediated protein attraction. A Monte Carlo study. Physica A 249:571–575
Sperotto MM, May S, Baumgaertner A (2006) Modelling of proteins in membranes. Chem Phys Lipids 141:2–29
Stroud RM, Reiling K, Wiener M, Freymann D (1998) Ion-channel forming colicins. Curr Opin Struct Biol 8:525–533
Sung S-S (1994) Helix folding simulations with various initial conformations. Biophys J 66:1796–1803
Sung S-S (1995) Folding simulations of alanine-based peptides with lysine residues. Biophys J 68:1796–1803
Terrones O, Antonsson B, Yamaguchi H, Wang HG, Liu J, Lee RM, Herrmann A, Basanez G (2004) Lipidic pore formation by the concerted action of proapoptopic BAX and tBid. J Biol Chem 279:30081–30091
Tzlil S, Ben-Schaul A (2005) Flexible charged molecules on mixed fluid lipid membranes: theory and Monte Carlo simulations. Biophys J 88:2391–2402
van der Goot FG, Gonzalez-Manas JM, Lakey JH, Pattus F (1991) A “molten-globule” membrane-insertion intermediate of the pore-forming domain of colicin A. Nature 354:408–410
Varfolomeev EE, Ashkenazi A (2004) Tumor necrosis factor: an apoptosis JuNKie? Cell 116:491–497
Vogt B, Ducarme P, Schinzel S, Brasseur R, Bechinger B (2000) The topology of lysine-containing amphipathic peptides in bilayers by circular dichroism, solid-state NMR, and molecular modeling. Biophys J 79:2644–2656
Wang X (2001) The expanding role of mitochondria in apoptosis. Genes Dev 15:2922–2933
Wang K, Yin XM, Chao DT, Milliman CL, Korsmeyer SJ (1996) Bid: a novel BH3 domain-only death agonist. Genes Dev 10:2859–2869
Wei MC, Lindsen T, Mootha VK, Weiler S, Gross A, Ashiya M, Thompson CB, Korsemeyer SJ (2000) tBid, a membrane-targeted death ligand, oligomerizes BAK to release cytochrome c. Genes Dev 14:2060–2071
Wei MC, Zong WX, Cheng EH, Lindsten T, Panautsakoupolo V, Ross AJ, Roth KA, McGregor GR, Thompson CB, Korsmayer SJ (2001) Proapoptotic Bax and Bak: a requisite gateway to mitochondrial dysfunction and death. Science 292:727–730
Willis SN, Adams JM (2005) Life in the balance: how BH3-only proteins induce apoptosis. Curr Opin Cell Biol 17:617–625
Winterhalter M, Helfrich W (1992) Bending elacticity of electrically charged bilayers: coupled monolayers, neutral surfaces, and balancing stresses. J Phys Chem 96:327–330
Yamaguchi S, Huster D, Waring A, Lehrer RI, Kearney W, Tack BF, Hong M (2001) Orientation and dynamics of an antimicrobial peptide in the lipid bilayer by solid-state NMR spectroscopy. Biophys J 81:2203–2214
Yin X-M (2006) Bid, a BH3-only multi-functional molecule, is at the cross road of life and death. Gene 369:7–19
Zakharov SD, Cramer WA (2002) Colicin crystal structures: pathways and mechanisms for colicin insertion into membranes. Biochim Biophys Acta 1565:333–346
Zemel A, Ben-Shaul A, May S (2004) Membrane perturbation induced by interfacially adsorbed peptides. Biophys J 86:3607–3619
Zha J, Weiler S, Oh KJ, Wei MC, Korsemeyer SJ (2000) Posttranslational N-Myristoylation of BID as a molecular switch for targeting mitochondria and apoptosis. Science 290:1761–1765
Zhan L, Chen JZY, Liu W-K (2006) Monte Carlo basin paving: an improved global optimization method. Phys Rev E 73:015701 (1–4)
Acknowledgments
The authors thank Prof. Gregory Nikiforovich (Washington University School of Medicine, St Louis, Missouri) for reading the early version of the manuscript and his insightful comments. This work was supported by the Program “Bioengineering and Biosecurity” of Republic of Belarus (Grant P-16).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Veresov, V.G., Davidovskii, A.I. Monte Carlo simulations of tBid association with the mitochondrial outer membrane. Eur Biophys J 37, 19–33 (2007). https://doi.org/10.1007/s00249-007-0149-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-007-0149-z