Abstract
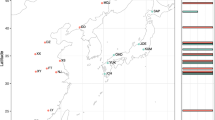
Most insects are associated with bacterial symbionts. The bacterial diversity and community composition within hosts may play an important role in shaping insect population biology, ecology and evolution. We focussed on the bacterial microbiome of the Australian fig homotomid Mycopsylla fici (Hemiptera: Psylloidea), which can cause defoliation of its only host tree, Ficus macrophylla. This sap-feeding insect is native to mainland Australia and Lord Howe Island (LHI) but also occurs where its host has been planted, notably in New Zealand. By using a high-throughput 16S rDNA amplicon sequencing approach, we compared the bacterial diversity and community composition in individual adult males of four host populations, Sydney, Brisbane, LHI and Auckland. We also compared males, females and nymphs of the Sydney population. The microbiome of M. fici was simple and consisted mostly of the following three maternally inherited endosymbiont species: the primary endosymbiont Carsonella, a secondary (S-) endosymbiont and Wolbachia. However, the relative abundance of their sequence reads varied between host populations, except for similarities between Sydney and Auckland. In addition, insects from Sydney and Auckland had identical bacterial strains supporting the hypothesis that Sydney is the source population for Auckland. In contrast, mainland and LHI populations harboured the same S-endosymbiont, co-diverged Carsonella but different Wolbachia strains. Besides detecting endosymbiont-specific patterns of either co-evolution or horizontal acquisition, our study highlights that relative abundance of maternally inherited endosymbionts should also be taken into account when studying bacterial communities across host populations, as variations in bacterial density may impact host biology and ecology.




Similar content being viewed by others
References
Buchner P (1953) Endosymbiose der Tiere mit pflanzlichen Mikroorganismen. Verlag Birkhauser, Basel
Moran NA, McCutcheon JP, Nakabachi A (2008) Genomics and evolution of heritable bacterial symbionts. Annu. Rev. Genet. 42:165–190. doi:10.1146/annurev.genet.41.110306.130119
Morrow JL, Frommer M, Shearman DCA, Riegler M (2014) Tropical tephritid fruit fly community with high incidence of shared Wolbachia strains as platform for horizontal transmission of endosymbionts. Environ. Microbiol. 16:3622–3637. doi:10.1111/1462-2920.12382/abstract
Morrow JL, Frommer M, Shearman DCA, Riegler M (2015) The microbiome of field-caught and laboratory-adapted Australian tephritid fruit fly species with different host plant use and specialisation. Microb. Ecol. 70:498–508. doi:10.1007/s00248-015-0571-1
Douglas AE (2009) The microbial dimension in insect nutritional ecology. Funct. Ecol. 23:38–47. doi:10.1111/j.1365-2435.2008.01442.x
Sloan DB, Moran NA (2012) Genome reduction and co-evolution between the primary and secondary bacterial symbionts of psyllids. Mol. Biol. Evol. 29:3781–3792. doi:10.1093/molbev/mss180
Douglas AE (2006) Phloem-sap feeding by animals: problems and solutions. J. Exp. Bot. 57:747–754. doi:10.1093/jxb/erj067
Shigenobu S, Watanabe H, Hattori M, et al (2000) Genome sequence of the endocellular bacterial symbiont of aphids Buchnera sp. APS. Nature 407:81–86. doi:10.1038/35024074
Shigenobu S, Wilson ACC (2011) Genomic revelations of a mutualism: the pea aphid and its obligate bacterial symbiont. Cell. Mol. Life Sci. 68:1297–1309. doi:10.1007/s00018-011-0645-2
Tamames J, Gil R, Latorre A, et al (2007) The frontier between cell and organelle: genome analysis of Candidatus Carsonella ruddii. BMC Evol. Biol. 7:181. doi:10.1186/1471-2148-7-181
Baumann P (2005) Biology of bacteriocyte-associated endosymbionts of plant sap-sucking insects. Annu. Rev. Microbiol. 59:155–189. doi:10.1146/annurev.micro.59.030804.121041
Oliver KM, Noge K, Huang EM, et al (2012) Parasitic wasp responses to symbiont-based defense in aphids. BMC Biol. 10:11. doi:10.1186/1741-7007-10-11
Dunbar HE, Wilson ACC, Ferguson NR, Moran NA (2007) Aphid thermal tolerance is governed by a point mutation in bacterial symbionts. PLoS Biol. 5:e96. doi:10.1371/journal.pbio.0050096
Lukasik P, van Asch M, Guo H, et al (2012) Unrelated facultative endosymbionts protect aphids against a fungal pathogen. Ecol. Lett. 16:214–218. doi:10.1111/ele.12031
Gueguen G, Vavre F, Gnankine O, et al (2010) Endosymbiont metacommunities, mtDNA diversity and the evolution of the Bemisia tabaci (Hemiptera: Aleyrodidae) species complex. Mol. Ecol. 19:4365–4378. doi:10.1111/j.1365-294X.2010.04775.x
Hansen AK, Jeong G, Paine TD, Stouthamer R (2007) Frequency of secondary symbiont infection in an invasive psyllid relates to parasitism pressure on a geographic scale in California. Appl. Environ. Microbiol. 73:7531–7535. doi:10.1128/AEM.01672-07
Gauthier J-P, Outreman Y, Mieuzet L, Simon J-C (2015) Bacterial communities associated with host-adapted populations of pea aphids revealed by deep sequencing of 16S ribosomal DNA. PLoS One 10:e0120664. doi:10.1371/journal.pone.0120664
Tsuchida T, Koga R, Shibao H, et al (2002) Diversity and geographic distribution of secondary endosymbiotic bacteria in natural populations of the pea aphid, Acyrthosiphon pisum. Mol. Ecol. 11:2123–2135
Smith CCR, Snowberg LK, Caporaso GJ, et al (2015) Dietary input of microbes and host genetic variation shape among-population differences in stickleback gut microbiota. ISME J 9: 2515-2526. doi:10.1038/ismej.2015.64
Hoffmann M, Coy MR, Kingdom Gibbard HN, Pelz-Stelinzki KS (2014) Wolbachia infection density in populations of the Asian citrus psyllid (Hemiptera: Liviidae). Environ. Entomol. 43:1215–1222
Bansal R, Mian MAR, Michel AP (2014) Microbiome diversity of Aphis glycines with extensive superinfection in native and invasive populations. Environ. Microbiol. Rep. 6:57–69. doi:10.1111/1758-2229.12108
Hall AAG, Morrow JL, Fromont C, et al (2016) Codivergence of the primary bacterial endosymbiont of psyllids versus host switches and replacement of their secondary bacterial endosymbionts. Environ. Microbiol. 18:2591–2603
Dossi FCA, da Silva EP, Cônsoli FL (2014) Population dynamics and growth rates of endosymbionts during Diaphorina citri (Hemiptera, Liviidae) ontogeny. Microb. Ecol. 68:881–889. doi:10.1007/s00248-014-0463-9
Fromont C, Riegler M, Cook JM (2016) Phylogeographic analyses of bacterial endosymbionts in fig homotomids (Hemiptera: Psylloidea) reveal codiversification of both primary and secondary endosymbionts. FEMS Microbiol. Ecol. 92:fiw205. doi:10.1093/femsec/fiw205
Aksoy E, Telleria EL, Echodu R, et al (2014) Analysis of multiple tsetse fly populations in Uganda reveals limited diversity and species-specific gut microbiota. Appl. Environ. Microbiol. 80:4301–4312. doi:10.1128/AEM.00079-14
Jing X, Wong AC-N, Chaston JM, et al (2014) The bacterial communities in plant phloem-sap-feeding insects. Mol. Ecol. 23:1433–1444. doi:10.1111/mec.12637
Capinha C, Essl F, Seebens H, et al (2015) The dispersal of alien species redefines biogeography in the Anthropocene. Science 348:1248–1251
Zouache K, Voronin D, Tran-Van V, Mavingui P (2009) Composition of bacterial communities associated with natural and laboratory populations of Asobara tabida infected with Wolbachia. Appl. Environ. Microbiol. 75:3755–3764. doi:10.1128/AEM.02964-08
Nguyen DT, Spooner-Hart RN, Riegler M (2016) Loss of Wolbachia but not Cardinium in the invasive range of the Australian thrips species, Pezothrips kellyanus. Biol. Invasions 18:197–214. doi:10.1007/s10530-015-1002-4
Newman AKL (2004) The biology of Mycopsylla fici Tryon on its sole host, Ficus macrophylla Desf. Ex Pers. Macquarie University, Sydney
Bain J (2004) New records. For Heal News 142:1–2
Herlemann DPR, Labrenz M, Jürgens K, et al (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571–1579. doi:10.1038/ismej.2011.41
Schloss PD, Westcott SL, Ryabin T, et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75:7537–7541. doi:10.1128/AEM.01541-09
Kozich JJ, Westcott SL, Baxter NT, et al (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq illumina sequencing platform. Appl. Environ. Microbiol. 79:5112–5120. doi:10.1128/AEM.01043-13
Pruesse E, Quast C, Knittel K, et al (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35:7188–7196. doi:10.1093/nar/gkm864
Edgar RC, Haas BJ, Clemente JC, et al (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. doi:10.1093/bioinformatics/btr381
Tamura K, Stecher G, Peterson D, et al (2013) MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30:2725–2729
Ronquist F, Teslenko M, van der Mark P, et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61:539–542. doi:10.1093/sysbio/sys029
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29:1969–1973. doi:10.1093/molbev/mss075
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57:289–300
Team RDC (2014) A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
De Mendiburu F (2014) Agricolae: statistical procedures for agricultural research. R package version 1.2-4. http://CRAN.R-project.org/package=agricolae
Oksanen J, Blanchet G, Kindt R, et al (2015) Vegan: community ecology package. R package version 2:3–1 http://CRAN.R-project.org/package=vegan
Warnes GR, Bolker B, Bonebakker L, et al (2015) gplots: various R programming tools for plotting data. R package version 2.17.0. http://CRAN.R-project.org/package=gplots.
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405. doi:10.1093/bioinformatics/btn129
Fromont C, DeGabriel JL, Riegler M, Cook JM (2017) Diversity and specificity of sap-feeding herbivores and their parasitoids on Australian fig trees. Insect Conserv Divers 70:107–119
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. doi:10.1093/bioinformatics/btg180
Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20:289–290
Overholt WA, Diaz R, Rosskopf E, et al (2015) Deep characterization of the microbiomes of Calophya spp. (Hemiptera: Calophyidae) gall-inducing psyllids reveals the absence of plant pathogenic bacteria and three dominant endosymbionts. PLoS One 10:e0132248. doi:10.1371/journal.pone.0132248
Nakabachi A, Yamashita A, Toh H, et al (2006) The 160-kilobase genome of the bacterial endosymbiont Carsonella. Science 314:267
McCutcheon JP, Von Dohlen CD (2011) An interdependent metabolic patchwork in the nested symbiosis of mealybugs. Curr. Biol. 21:1366–1372. doi:10.1016/j.cub.2011.06.051
Zug R, Hammerstein P (2012) Still a host of hosts for Wolbachia: analysis of recent data suggests that 40% of terrestrial arthropod species are infected. PLoS One 7:e38544. doi:10.1371/journal.pone.0038544
Weinert LA, Araujo-Jnr EV, Ahmed MZ, Welch JJ (2015) The incidence of bacterial endosymbionts in terrestrial arthropods. Proc. R. Soc. B Biol. Sci. 282:20150249. doi:10.1098/rspb.2015.0249
Hosokawa T, Koga R, Kikuchi Y, et al (2010) Wolbachia as a bacteriocyte-associated nutritional mutualist. Proc. Natl. Acad. Sci. 107:769–774. doi:10.1073/pnas.0911476107
Andersen SB, Boye M, Nash DR, Boomsma JJ (2012) Dynamic Wolbachia prevalence in Acromyrmex leaf-cutting ants: potential for a nutritional symbiosis. J. Evol. Biol. 25:1340–1350. doi:10.1111/j.1420-9101.2012.02521.x
Riegler M, Stauffer C (2002) Infections and superinfections in cytoplasmically incompatible populations of the European cherry fruit fly. Mol. Ecol. 11:2425–2434
Schuler H, Köppler K, Daxböck-Horvath S, et al (2016) The hitchhiker’s guide to Europe: the infection dynamics of an ongoing Wolbachia invasion and mitochondrial selective sweep in Rhagoletis cerasi. Mol. Ecol. 25:1595–1609. doi:10.1111/mec.13571
Clark EL, Daniell TJ, Wishart J, et al (2012) How conserved are the bacterial communities associated with aphids? A detailed assessment of the Brevicoryne brassicae (Hemiptera: Aphididae) using 16S rDNA. Environ. Entomol. 41:1386–1397. doi:10.1603/EN12152
XiaoFang M, XueChao Z, HaiJun X (2012) The endosymbiotic microbiota of two geographical populations of the Asian citrus psyllid, Diaphorina citri, in China and its ability of pathogen transmission. Acta Entomol. Sin. 55:1149–1153
Vicente CSL, Nascimento FX, Espada M, et al (2013) Characterization of bacterial communities associated with the pine sawyer beetle Monochamus galloprovincialis, the insect vector of the pinewood nematode Bursaphelenchus xylophilus. FEMS Microbiol. Lett. 347:130–139. doi:10.1111/1574-6968.12232
Gendrin M, Christophides GK (2013) The anopheles mosquito microbiota and their impact on pathogen transmission. In: Anopheles mosquitoes - New insights into Malar. vectors. pp 525–548
Davidson EW, Rosell RC, Hendrix DL (2000) Culturable bacteria associated with the whitefly, Bemisia argentifolii (Homoptera: Aleyrodidae). Florida Entomol 82:159–171
Grenier A-M, Nardon C, Rahbé Y (1994) Observations on the micro-organisms occurring in the gut of the pea aphid Acyrthosiphon pisum. Entomol Exp Appl 70:91–96. doi:10.1007/BF02380635
Dillon RJ, Dillon VM (2004) The gut bacteria of insects: nonpathogenic interactions. Annu. Rev. Entomol. 49:71–92. doi:10.1146/annurev.ento.49.061802.123416
Yun J, Roh W, Whon W, et al (2014) Insect gut bacterial diversity determined by environmental habitat, diet, developmental stage, and phylogeny of host. Appl. Environ. Microbiol. 80:5254–5264. doi:10.1128/AEM.01226-14
Minard G, Tran F-H, Dubost A, et al (2014) Pyrosequencing 16S rRNA genes of bacteria associated with wild tiger mosquito Aedes albopictus: a pilot study. Front. Cell. Infect. Microbiol. doi:10.3389/fcimb.2014.00059
Zheng L, Crippen TL, Singh B, et al (2013) A survey of bacterial diversity from successive life stages of black soldier fly (Diptera: Stratiomyidae) by using 16S rDNA pyrosequencing. J. Med. Entomol. 50:647–658
Thao ML, Clark MA, Baumann L, et al (2000) Secondary endosymbionts of psyllids have been acquired multiple times. Curr. Microbiol. 41:300–304. doi:10.1007/s002840010138
Thao ML, Moran NA, Abbot P, et al (2000) Cospeciation of psyllids and their primary prokaryotic endosymbionts. Appl. Environ. Microbiol. 66:2898–2905
Wernegreen JJ (2002) Genome evolution in bacterial endosymbionts of insects. Nat Rev Genet 3:850–861. doi:10.1038/nrg931
Fromont C, Rymer PD, Riegler M, Cook JM (submitted) Hop, step and jump—natural and anthropogenic dispersal of a host-specific insect to habitat and oceanic islands.
McDougall I, Embleton BJJ, Stone DB (1981) Origin and evolution of Lord Howe Island, Southwest Pacific Ocean. J. Geol. Soc. Aust. 28:155–176. doi:10.1080/00167618108729154
Stireman III JO, Nason JD, Heard SB (2005) Host-associated genetic differentiation in phytophagous insects: general phenomenon or isolated exceptions? Evidence from a goldenrod-insect community. Evolution 59:2573–2587. doi:10.1111/j.0014-3820.2005.tb00970.x
Cook JM, Butcher RDJ (1999) The transmission and effects of Wolbachia bacteria in parasitoids. Res Popul Ecol (Kyoto) 41:15–28. doi:10.1007/PL00011978
Heath BD, Butcher RD, Whitfield WG, Hubbard SF (1999) Horizontal transfer of Wolbachia between phylogenetically distant insect species by a naturally occurring mechanism. Curr. Biol. 9:313–326
Amend AS, Seifert KA, Bruns TD (2010) Quantifying microbial communities with 454 pyrosequencing: does read abundance count? Mol. Ecol. 19:5555–5565. doi:10.1111/j.1365-294X.2010.04898.x
Fagen JR, Giongo A, Brown CT, et al (2012) Characterization of the relative abundance of the citrus pathogen Ca. Liberibacter asiaticus in the microbiome of its insect vector, Diaphorina citri, using high throughput 16S rRNA sequencing. Open Microbiol J 6:29–33
Pinto AJ, Raskin L (2012) PCR biases distort bacterial and archaeal community structure in pyrosequencing datasets. PLoS One 7:e43093. doi:10.1371/journal.pone.0043093
Chandler JA, Lang J, Bhatnagar S, et al (2011) Bacterial communities of diverse Drosophila species: ecological context of a host-microbe model system. PLoS Genet. 7:e1002272. doi:10.1371/journal.pgen.1002272
Ikeda T, Ishikawa H, Sasaki T (2003) Regulation of Wolbachia density in the Mediterranean flour moth, Ephestia kuehniella, and the almond moth, Cadra cautella. Zool. Sci. 20:153–157. doi:10.2108/zsj.20.153
Mouton L, Henri H, Bouletreau M, Vavre F (2006) Effect of temperature on Wolbachia density and impact on cytoplasmic incompatibility. Parasitology 132:49–56. doi:10.1017/S0031182005008723
Mouton L, Henri H, Bouletreau M, Vavre F (2003) Strain-specific regulation of intracellular Wolbachia density in multiply infected insects. Mol. Ecol. 12:3459–3465. doi:10.1046/j.1365-294X.2003.02015.x
Kondo N, Shimada M, Fukatsu T (2005) Infection density of Wolbachia endosymbiont affected by co-infection and host genotype. Biol. Lett. 1:488–491. doi:10.1098/rsbl.2005.0340
Mouton L, Henri H, Charif D, et al (2007) Interaction between host genotype and environmental conditions affects bacterial density in Wolbachia symbiosis. Biol. Lett. 3:210–213. doi:10.1098/rsbl.2006.0590
Anbutsu H, Goto S, Fukatsu T (2008) High and low temperatures differently affect infection density and vertical transmission of male-killing Spiroplasma symbionts in Drosophila hosts. Appl. Environ. Microbiol. 74:6053–6059. doi:10.1128/AEM.01503-08
Su Q, Xie W, Wang S, et al (2014) Location of symbionts in the whitefly Bemisia tabaci affects their densities during host development and environmental stress. PLoS One 9:e91802. doi:10.1371/journal.pone.0091802
Thomas MB, Blanford S (2003) Thermal biology in insect-parasite interactions. Trends Ecol. Evol. 18:344–350. doi:10.1016/S0169-5347(03)00069-7
Acknowledgements
CF was supported by a Hawkesbury Institute for the Environment Postgraduate Research Award. This research was also supported by a research grant of the Hermon Slade Foundation (HSF 12/10) to MR and JMC. We want to thank Tim Sutton, Desi Quintans, Aidan Hall, Jane DeGabriel, Courtney Campany and John Early for helping with the field collections as well as the Lord Howe Island Board and NSW Office of Environment and Heritage for the permission to collect specimens. We also want to thank Caroline Janitz, Emma Hackett and Jocelyn King from the Next-Generation Sequencing Facility at Western Sydney University for the 16S metagenomic sequencing services and advices.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fromont, C., Riegler, M. & Cook, J.M. Relative Abundance and Strain Diversity in the Bacterial Endosymbiont Community of a Sap-Feeding Insect Across Its Native and Introduced Geographic Range. Microb Ecol 74, 722–734 (2017). https://doi.org/10.1007/s00248-017-0971-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-0971-5