Abstract
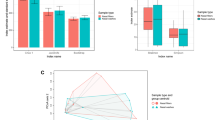
To date, there is a limited understanding of the role of the airway microbiome in the early life development of respiratory diseases such as asthma, partly due to a lack of simple and minimally invasive sample collection methods. In order to characterize the baseline microbiome of the upper respiratory tract (URT) in infants, a comparatively non-invasive method for sampling the URT microbiome suitable for use in infants was developed. Microbiome samples were collected by placing filter paper in the nostrils of 33 healthy, term infants enrolled as part of the Infant Susceptibility to Pulmonary Infections and Asthma Following RSV Exposure (INSPIRE) study. After bacterial genomic DNA was extracted from the filters, amplicons were generated with universal primers targeting the V1–V3 region of the 16S rRNA gene. This method was capable of capturing a wide variety of taxa expected to inhabit the nasal cavity. Analyses stratifying subjects by demographic and environmental factors previously observed or predicted to influence microbial communities were performed. Microbial community richness was found to be higher in infants who had been delivered via Cesarean section and in those who had been formula-fed; an association was observed between diet and delivery, which confounds this analysis. We have established a baseline URT microbiome using a non-invasive filter paper nasal sampling for this population, and future studies will be performed in this large observational cohort of infants to investigate the relationship between viral infections, the URT microbiota, and the development of childhood wheezing illnesses.


Similar content being viewed by others
References
The Human Microbiome Project Consortium (2012) Structure, function and diversity of the healthy human microbiome. Nature 486:207–214. doi:10.1038/nature11234
Eder W, Ege MJ, von Mutius E (2006) The asthma epidemic. N Engl J Med 355(21):2226–2235. doi:10.1056/NEJMra054308
Martinez FD, Vercelli D (2013) Asthma. Lancet 382(9901):1360–1372. doi:10.1016/S0140-6736(13)61536-6
Cho I, Blaser MJ (2012) The human microbiome: at the interface of health and disease. Nat Rev Genet 13(4):260–270. doi:10.1038/nrg3182
Madupu R, Szpakowski S, Nelson KE (2013) Microbiome in human health and disease. Sci Prog 96(Pt 2):153–170
Guinane CM, Cotter PD (2013) Role of the gut microbiota in health and chronic gastrointestinal disease: understanding a hidden metabolic organ. Ther Adv Gastroenterol 6(4):295–308. doi:10.1177/1756283X13482996
Legatzki A, Rosler B, von Mutius E (2014) Microbiome diversity and asthma and allergy risk. Curr Allergy Asthma Rep 14(10):466. doi:10.1007/s11882-014-0466-0
Bisgaard H, Hermansen MN, Buchvald F, Loland L, Halkjaer LB, Bonnelykke K, Brasholt M, Heltberg A, Vissing NH, Thorsen SV, Stage M, Pipper CB (2007) Childhood asthma after bacterial colonization of the airway in neonates. N Engl J Med 357(15):1487–1495. doi:10.1056/NEJMoa052632
von Linstow M-L, Schønning K, Hoegh AM, Sevelsted A, Vissing NH, Bisgaard H (2013) Neonatal airway colonization is associated with troublesome lung symptoms in infants. Am J Respir Crit Care Med 188(8):1041–1042. doi:10.1164/rccm.201302-0395LE
Hilty M, Burke C, Pedro H, Cardenas P, Bush A, Bossley C, Davies J, Ervine A, Poulter L, Pachter L, Moffatt MF, Cookson WOC (2010) Disordered microbial communities in asthmatic airways. PLoS ONE 5(1):e8578. doi:10.1371/journal.pone.0008578
Huang YJ, Nelson CE, Brodie EL, Desantis TZ, Baek MS, Liu J, Woyke T, Allgaier M, Bristow J, Wiener-Kronish JP, Sutherland ER, King TS, Icitovic N, Martin RJ, Calhoun WJ, Castro M, Denlinger LC, Dimango E, Kraft M, Peters SP, Wasserman SI, Wechsler ME, Boushey HA, Lynch SV, National Heart, Lung, and Blood Institute’s Asthma Clinical Research Network (2011) Airway microbiota and bronchial hyperresponsiveness in patients with suboptimally controlled asthma. J Allergy Clin Immunol 127(2):372–381 e371–373. doi:10.1016/j.jaci.2010.10.048
Marri PR, Stern DA, Wright AL, Billheimer D, Martinez FD (2013) Asthma-associated differences in microbial composition of induced sputum. J Allergy Clin Immunol 131(2):346–352.e341–343. doi:10.1016/j.jaci.2012.11.013
Grad R, Morgan WJ (2012) Long-term outcomes of early-onset wheeze and asthma. J Allergy Clin Immunol 130(2):299–307. doi:10.1016/j.jaci.2012.05.022
Sigurs N, Bjarnason R, Sigurbergsson F, Kjellman B (2000) Respiratory syncytial virus bronchiolitis in infancy is an important risk factor for asthma and allergy at age 7. Am J Respir Crit Care Med 161(5):1501–1507. doi:10.1164/ajrccm.161.5.9906076
Sigurs N, Aljassim F, Kjellman B, Robinson PD, Sigurbergsson F, Bjarnason R, Gustafsson PM (2010) Asthma and allergy patterns over 18 years after severe RSV bronchiolitis in the first year of life. Thorax 65(12):1045–1052. doi:10.1136/thx.2009.121582
Sigurs N, Gustafsson PM, Bjarnason R, Lundberg F, Schmidt S, Sigurbergsson F, Kjellman B (2005) Severe respiratory syncytial virus bronchiolitis in infancy and asthma and allergy at age 13. Am J Respir Crit Care Med 171(2):137–141. doi:10.1164/rccm.200406-730OC
Lynch SV (2014) Viruses and microbiome alterations. Ann Am Thorac Soc 11(Suppl 1):S57–S60. doi:10.1513/AnnalsATS.201306-158MG
Larkin EK, Gebretsadik T, Moore ML, Anderson LJ, Dupont WD, Chappell JD, Minton PA, Peebles RS Jr, Moore PE, Valet RS, Arnold DH, Rosas-Salazar C, Das SR, Polack FP, Hartert TV (2015) Objectives, design and enrollment results from the Infant Susceptibility to Pulmonary Infections and Asthma following RSV exposure study (INSPIRE). BMC Pulm Med 15(1):45. doi:10.1186/s12890-015-0040-0
Folsgaard NV, Schjorring S, Chawes BL, Rasmussen MA, Krogfelt KA, Brix S, Bisgaard H (2013) Pathogenic bacteria colonizing the airways in asymptomatic neonates stimulates topical inflammatory mediator release. Am J Respir Crit Care Med 187(6):589–595. doi:10.1164/rccm.201207-1297OC
Fouts DE, Pieper R, Szpakowski S, Pohl H, Knoblach S, Suh M-J, Huang S-T, Ljungberg I, Sprague BM, Lucas SK, Torralba M, Nelson KE, Groah SL (2012) Integrated next-generation sequencing of 16S rDNA and metaproteomics differentiate the healthy urine microbiome from asymptomatic bacteriuria in neuropathic bladder associated with spinal cord injury. J Transl Med 10(1):174
Schloss P, Westcott S, Ryabin T, Hall J, Hartmann M, Hollister E, Lesniewski R, Oakley B, Parks D, Robinson C, Sahl JW, Stres B, Thallinger G, Van Horn D, Weber C (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541
Sebastian S YAP. https://github.com/shpakoo/YAP
Fouts DE, Szpakowski S, Purushe J, Torralba M, Waterman RC, MacNeil MD, Alexander LJ, Nelson KE (2012) Next generation sequencing to define prokaryotic and fungal diversity in the bovine rumen. PLoS ONE 7(11), e48289. doi:10.1371/journal.pone.0048289
Schloss PD, Gevers D, Westcott SL (2011) Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PLoS ONE 6(12), e27310. doi:10.1371/journal.pone.0027310
Hamady M, Knight R (2009) Microbial community profiling for human microbiome projects: tools, techniques, and challenges. Genome Res 19(7):1141–1152. doi:10.1101/gr.085464.108
Tovchigrechko A. MGSAT—statistical analysis of microbiome and proteome abundance matrices with automated report generation. https://bitbucket.org/andreyto/mgsat
Bassis CM, Tang AL, Young VB, Pynnonen MA (2014) The nasal cavity microbiota of healthy adults. Microbiome 2(1):27. doi:10.1186/2049-2618-2-27
Costello EK, Lauber CL, Hamady M, Fierer N, Gordon JI, Knight R (2009) Bacterial community variation in human body habitats across space and time. Science 326(5960):1694–1697. doi:10.1126/science.1177486
Frank DN, Feazel LM, Bessesen MT, Price CS, Janoff EN, Pace NR (2010) The human nasal microbiota and Staphylococcus aureus carriage. PLoS ONE 5(5), e10598. doi:10.1371/journal.pone.0010598
Lemon KP, Klepac-Ceraj V, Schiffer HK, Brodie EL, Lynch SV, Kolter R (2010) Comparative analyses of the bacterial microbiota of the human nostril and oropharynx. mBio 1(3). doi:10.1128/mBio.00129-10
Biesbroek G, Bosch AATM, Wang X, Keijser BJF, Veenhoven RH, Sanders EAM, Bogaert D (2014) The impact of breastfeeding on nasopharyngeal microbial communities in infants. Am J Respir Crit Care Med 190(3):298–308. doi:10.1164/rccm.201401-0073OC
Biesbroek G, Tsivtsivadze E, Sanders EAM, Montijn R, Veenhoven RH, Keijser BJF, Bogaert D (2014) Early respiratory microbiota composition determines bacterial succession patterns and respiratory health in children. Am J Respir Crit Care Med 190(11):1283–1292. doi:10.1164/rccm.201407-1240OC
Verduin CM, Hol C, Fleer A, van Dijk H, van Belkum A (2002) Moraxella catarrhalis: from emerging to established pathogen. Clin Microbiol Rev 15(1):125–144. doi:10.1128/CMR.15.1.125-144.2002
Verhaegh SJC, Lebon A, Saarloos JA, Verbrugh HA, Jaddoe VWV, Hofman A, Hays JP, Moll HA, van Belkum A (2010) Determinants of Moraxella catarrhalis colonization in healthy Dutch children during the first 14 months of life. Clin Microbiol Infect 16(7):992–997. doi:10.1111/j.1469-0691.2009.03008.x
Dominguez-Bello MG, Costello EK, Contreras M, Magris M, Hidalgo G, Fierer N, Knight R (2010) Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci U S A 107(26):11971–11975. doi:10.1073/pnas.1002601107
Biasucci G, Benenati B, Morelli L, Bessi E, Boehm G (2008) Cesarean delivery may affect the early biodiversity of intestinal bacteria. J Nutr 138(9):1796S–1800S
Jakobsson HE, Abrahamsson TR, Jenmalm MC, Harris K, Quince C, Jernberg C, Björkstén B, Engstrand L, Andersson AF (2014) Decreased gut microbiota diversity, delayed Bacteroidetes colonisation and reduced Th1 responses in infants delivered by caesarean section. Gut 63(4):559–566. doi:10.1136/gutjnl-2012-303249
Azad MB, Konya T, Maughan H, Guttman DS, Field CJ, Chari RS, Sears MR, Becker AB, Scott JA, Kozyrskyj AL (2013) Gut microbiota of healthy Canadian infants: profiles by mode of delivery and infant diet at 4 months. Can Med Assoc J 185(5):385–394. doi:10.1503/cmaj.121189
Tlaskalová-Hogenová H, Štěpánková R, Kozáková H, Hudcovic T, Vannucci L, Tučková L, Rossmann P, Hrnčíř T, Kverka M, Zákostelská Z, Klimešová K, Přibylová J, Bártová J, Sanchez D, Fundová P, Borovská D, Šrůtková D, Zídek Z, Schwarzer M, Drastich P, Funda DP (2011) The role of gut microbiota (commensal bacteria) and the mucosal barrier in the pathogenesis of inflammatory and autoimmune diseases and cancer: contribution of germ-free and gnotobiotic animal models of human diseases. Cell Mol Immunol 8(2):110–120. doi:10.1038/cmi.2010.67
Gill SR, Pop M, DeBoy RT, Eckburg PB, Turnbaugh PJ, Samuel BS, Gordon JI, Relman DA, Fraser-Liggett CM, Nelson KE (2006) Metagenomic analysis of the human distal gut microbiome. Science. doi:10.1126/science.1124234
Yan M, Pamp SJ, Fukuyama J, Hwang PH, Cho D-Y, Holmes S, Relman DA (2013) Nasal microenvironments and interspecific interactions influence nasal microbiota complexity and S. aureus carriage. Cell Host Microbe 14(6):631–640. doi:10.1016/j.chom.2013.11.005
Lina G, Boutite F, Tristan A, Bes M, Etienne J, Vandenesch F (2003) Bacterial competition for human nasal cavity colonization: role of staphylococcal agr alleles. Appl Environ Microbiol 69(1):18–23. doi:10.1128/aem.69.1.18-23.2003
Rutebemberwa A, Stevens MJ, Perez MJ, Smith LP, Sanders L, Cosgrove G, Robertson CE, Tuder RM, Harris JK (2014) Novosphingobium and its potential role in chronic obstructive pulmonary diseases: insights from microbiome studies. PLoS ONE 9(10), e111150. doi:10.1371/journal.pone.0111150
Renz-Polster H, David MR, Buist AS, Vollmer WM, O’Connor EA, Frazier EA, Wall MA (2005) Caesarean section delivery and the risk of allergic disorders in childhood. Clin Exp Allergy 35(11):1466–1472. doi:10.1111/j.1365-2222.2005.02356.x
Hauck YL, Fenwick J, Dhaliwal SS, Butt J (2011) A Western Australian survey of breastfeeding initiation, prevalence and early cessation patterns. Matern Child Health J 15(2):260–268. doi:10.1007/s10995-009-0554-2
Prior E, Santhakumaran S, Gale C, Philipps LH, Modi N, Hyde MJ (2012) Breastfeeding after cesarean delivery: a systematic review and meta-analysis of world literature. Am J Clin Nutr 95(5):1113–1135. doi:10.3945/ajcn.111.030254
Zanardo V, Svegliado G, Cavallin F, Giustardi A, Cosmi E, Litta P, Trevisanuto D (2010) Elective cesarean delivery: does it have a negative effect on breastfeeding? Birth 37(4):275–279. doi:10.1111/j.1523-536X.2010.00421.x
Penders J, Thijs C, Vink C, Stelma FF, Snijders B, Kummeling I, van den Brandt PA, Stobberingh EE (2006) Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 118(2):511–521. doi:10.1542/peds.2005-2824
Negele K, Heinrich J, Borte M, von Berg A, Schaaf B, Lehmann I, Wichmann HE, Bolte G (2004) Mode of delivery and development of atopic disease during the first 2 years of life. Pediatr Allergy Immunol 15(1):48–54
Collado MC, Rautava S, Isolauri E, Salminen S (2015) Gut microbiota: a source of novel tools to reduce the risk of human disease? Pediatr Res 77(1-2):182–188. doi:10.1038/pr.2014.173
Fernández L, Langa S, Martín V, Maldonado A, Jiménez E, Martín R, Rodríguez JM (2013) The human milk microbiota: origin and potential roles in health and disease. Pharmacol Res 69(1):1–10. doi:10.1016/j.phrs.2012.09.001
Johnson CL, Versalovic J (2012) The human microbiome and its potential importance to pediatrics. Pediatrics 129(5):950–960. doi:10.1542/peds.2011-2736
Zhou Y, Gao H, Mihindukulasuriya KA, La Rosa PS, Wylie KM, Vishnivetskaya T, Podar M, Warner B, Tarr PI, Nelson DE, Fortenberry JD, Holland MJ, Burr SE, Shannon WD, Sodergren E, Weinstock GM (2013) Biogeography of the ecosystems of the healthy human body. Genome Biol 14(1):R1. doi:10.1186/gb-2013-14-1-r1
Acknowledgments
We thank Theresa Rodger for providing outstanding technical assistance and Dr. Karla M. Stucker for her critical review and editing of the manuscript. The clinical sample and data collection for this study were supported by a National Institute of Allergy and Infectious Diseases grant (AI U19-AI-095277) and a Vanderbilt Institute for Clinical and Translational Research Grant (UL1 TR000445) from NCATS/NIH. The sequencing work was generously supported by the NIAID/NIH Genomic Centers for Infectious Diseases (GCID) program (U19-AI-110819). MHS and SRD are supported by U19AI095227 supplement. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Authors’ Contributions
MHS, RSP, MLM, LJA, KEN, TVH, and SRD conceived and designed the study. CRS, RSP, and TH collected clinical samples, and MHS, MT, RH, and SRD performed sample processing and 16S rRNA gene sequencing. MHS, AT, KEN, and SRD performed the 16S rRNA gene sequence data analysis. MHS, CRS, AT, MLM, LJA, KEN, TVH, and SRD wrote the manuscript, and all authors reviewed and approved the final version.
Conflict of Interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 113 KB)
Rights and permissions
About this article
Cite this article
Shilts, M.H., Rosas-Salazar, C., Tovchigrechko, A. et al. Minimally Invasive Sampling Method Identifies Differences in Taxonomic Richness of Nasal Microbiomes in Young Infants Associated with Mode of Delivery. Microb Ecol 71, 233–242 (2016). https://doi.org/10.1007/s00248-015-0663-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-015-0663-y