Abstract
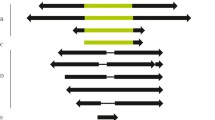
We compared deleted copies of the seven mauritiana subfamilies of mariner transposable elements in species of the Drosophilidae. All elements were detected by PCR using the inverted terminal repeats of the Mos1 element of Drosophila mauritiana as primers. A higher frequency of breakpoints in the 5′ part of the element compared to the 3′ part was observed. Of the 27 deletions, 9 (33%) occurred between short direct repeats (SDR) of 5 to 8 bp. The SDRs can be at or close to the breakpoints of the deletion. A deleted copy of D. simulans (St. Martin population) had three repeats of a motif present only once in the complete consensus sequence. The high frequency of SDRs at or near the breakpoints of the deletions strongly suggests that some of them do not occur at random. Mechanisms that might explain these deletions, such as unequal crossing-over, ectopic recombination, and abortive gap repair, are discussed.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 22 December 2000 / Accepted: 12 July 2001
Rights and permissions
About this article
Cite this article
Brunet, F., Giraud, T., Godin, F. et al. Do Deletions of Mos1-Like Elements Occur Randomly in the Drosophilidae Family?. J Mol Evol 54, 227–234 (2002). https://doi.org/10.1007/s0023901-0004-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s0023901-0004-2