Abstract
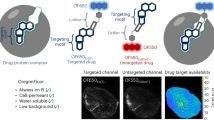
Here, we report a fluorescent probe based on a macrocyclic peptide scaffold that specifically stains EpCAM-expressing MCF7 cells. The 14-mer macrocyclic peptide binding to the extracellular domain of EpCAM with a dissociation constant in the low nM range (1.7 nM) was discovered using the random non-standard peptide-integrated discovery system. Notably, this probe containing a fluorescence tag is less than 3000 Da in total and able to visualize nearly every live cell under high cell-density conditions, which was not achieved by the conventional mAb staining method. This suggests that the molecular probe based on the compact macrocyclic scaffold has great potentials as an imaging tool for the EpCAM biomarker as well as a delivery vehicle for drug conjugates.




Similar content being viewed by others
References
Armstrong A, Eck SL (2003) EpCAM: a new therapeutic target for an old cancer antigen. Cancer Biol Ther 2:320–326
Baeuerle PA, Gires O (2007) EpCAM (CD326) finding its role in cancer. Br J Cancer 96:417–423. doi:10.1038/sj.bjc.6603494
Chaudry MA, Sales K, Ruf P, Lindhofer H, Winslet MC (2007) EpCAM an immunotherapeutic target for gastrointestinal malignancy: current experience and future challenges. Br J Cancer 96:1013–1019. doi:10.1038/sj.bjc.6603505
Cirulli V et al (1998) KSA antigen Ep-CAM mediates cell-cell adhesion of pancreatic epithelial cells: morphoregulatory roles in pancreatic islet development. J Cell Biol 140:1519–1534
Deonarain MP, Kousparou CA, Epenetos AA (2009) Antibodies targeting cancer stem cells: a new paradigm in immunotherapy? MAbs 1:12–25
Goto Y, Ohta A, Sako Y, Yamagishi Y, Murakami H, Suga H (2008) Reprogramming the translation initiation for the synthesis of physiologically stable cyclic peptides. ACS Chem Biol 3:120–129. doi:10.1021/cb700233t
Goto Y, Katoh T, Suga H (2011) Flexizymes for genetic code reprogramming. Nat Protoc 6:779–790. doi:10.1038/nprot.2011.331
Guillemot JC, Naspetti M, Malergue F, Montcourrier P, Galland F, Naquet P (2001) Ep-CAM transfection in thymic epithelial cell lines triggers the formation of dynamic actin-rich protrusions involved in the organization of epithelial cell layers. Histochem Cell Biol 116:371–378. doi:10.1007/s004180100329
Hayashi Y, Morimoto J, Suga H (2012) In vitro selection of anti-Akt2 thioether-macrocyclic peptides leading to isoform-selective inhibitors. ACS Chem Biol 7:607–613. doi:10.1021/cb200388k
Hristodorov D et al (2014) EpCAM-selective elimination of carcinoma cells by a novel MAP-based cytolytic fusion protein. Mol Cancer Ther 13:2194–2202. doi:10.1158/1535-7163.MCT-13-0781
Huls GA et al (1999) A recombinant, fully human monoclonal antibody with antitumor activity constructed from phage-displayed antibody fragments. Nat Biotechnol 17:276–281. doi:10.1038/7023
Hussain S, Pluckthun A, Allen TM, Zangemeister-Wittke U (2006) Chemosensitization of carcinoma cells using epithelial cell adhesion molecule-targeted liposomal antisense against bcl-2/bcl-xL. Mol Cancer Ther 5:3170–3180. doi:10.1158/1535-7163.MCT-06-0412
Ito K et al (2015) Artificial human Met agonists based on macrocycle scaffolds. Nat Commun 6:6373. doi:10.1038/ncomms7373
Iwasaki K, Goto Y, Katoh T, Suga H (2012) Selective thioether macrocyclization of peptides having the N-terminal 2-chloroacetyl group and competing two or three cysteine residues in translation. Org Biomol Chem 10:5783–5786. doi:10.1039/c2ob25306b
Jung YK, Woo MA, Soh HT, Park HG (2014) Aptamer-based cell imaging reagents capable of fluorescence switching. Chem Commun (Camb) 50:12329–12332. doi:10.1039/c4cc03888f
Kodan A et al (2014) Structural basis for gating mechanisms of a eukaryotic P-glycoprotein homolog. Proc Natl Acad Sci USA 111:4049–4054. doi:10.1073/pnas.1321562111
Kratz H, Haeckel A, Michel R, Schonzart L, Hanisch U, Hamm B, Schellenberger E (2012) Straightforward thiol-mediated protein labelling with DTPA: Synthesis of a highly active 111In-annexin A5-DTPA tracer. EJNMMI Res 2:17. doi:10.1186/2191-219X-2-17
Kwiatkowska-Borowczyk EP, Gabka-Buszek A, Jankowski J, Mackiewicz A (2015) Immunotargeting of cancer stem cells. Contemp Oncol (Pozn) 19:A52–A59. doi:10.5114/wo.2014.47129
Ladwein M et al (2005) The cell-cell adhesion molecule EpCAM interacts directly with the tight junction protein claudin-7. Exp Cell Res 309:345–357. doi:10.1016/j.yexcr.2005.06.013
Linke R, Klein A, Seimetz D (2010) Catumaxomab: clinical development and future directions. MAbs 2:129–136
Litvinov SV et al (1997) Epithelial cell adhesion molecule (Ep-CAM) modulates cell-cell interactions mediated by classic cadherins. J Cell Biol 139:1337–1348
Maetzel D et al (2009) Nuclear signalling by tumour-associated antigen EpCAM. Nat Cell Biol 11:162–171. doi:10.1038/ncb1824
Martin-Killias P, Stefan N, Rothschild S, Pluckthun A, Zangemeister-Wittke U (2011) A novel fusion toxin derived from an EpCAM-specific designed ankyrin repeat protein has potent antitumor activity. Clin Cancer Res 17:100–110. doi:10.1158/1078-0432.CCR-10-1303
Morimoto J, Hayashi Y, Suga H (2012) Discovery of macrocyclic peptides armed with a mechanism-based warhead: isoform-selective inhibition of human deacetylase SIRT2. Angew Chem Int Ed Engl 51:3423–3427. doi:10.1002/anie.201108118
Morioka T, Loik ND, Hipolito CJ, Goto Y, Suga H (2015) Selection-based discovery of macrocyclic peptides for the next generation therapeutics. Curr Opin Chem Biol 26:34–41. doi:10.1016/j.cbpa.2015.01.023
Munz M, Kieu C, Mack B, Schmitt B, Zeidler R, Gires O (2004) The carcinoma-associated antigen EpCAM upregulates c-myc and induces cell proliferation. Oncogene 23:5748–5758. doi:10.1038/sj.onc.1207610
Nemoto N, Miyamoto-Sato E, Husimi Y, Yanagawa H (1997) In vitro virus: bonding of mRNA bearing puromycin at the 3′-terminal end to the C-terminal end of its encoded protein on the ribosome in vitro. FEBS Lett 414:405–408
Nochi T et al (2004) Biological role of Ep-CAM in the physical interaction between epithelial cells and lymphocytes in intestinal epithelium. Clin Immunol 113:326–339. doi:10.1016/j.clim.2004.08.013
Osta WA et al (2004) EpCAM is overexpressed in breast cancer and is a potential target for breast cancer gene therapy. Cancer Res 64:5818–5824. doi:10.1158/0008-5472.CAN-04-0754
Passioura T, Katoh T, Goto Y, Suga H (2014) Selection-based discovery of druglike macrocyclic peptides. Annu Rev Biochem 83:727–752. doi:10.1146/annurev-biochem-060713-035456
Patriarca C, Macchi RM, Marschner AK, Mellstedt H (2012) Epithelial cell adhesion molecule expression (CD326) in cancer: a short review. Cancer Treat Rev 38:68–75. doi:10.1016/j.ctrv.2011.04.002
Roberts RW, Szostak JW (1997) RNA-peptide fusions for the in vitro selection of peptides and proteins. Proc Natl Acad Sci USA 94:12297–12302
Shaunak S et al (2006) Site-specific PEGylation of native disulfide bonds in therapeutic proteins. Nat Chem Biol 2:312–313. doi:10.1038/nchembio786
Song Y et al (2013) Selection of DNA aptamers against epithelial cell adhesion molecule for cancer cell imaging and circulating tumor cell capture. Anal Chem 85:4141–4149. doi:10.1021/ac400366b
Spizzo G et al (2004) High Ep-CAM expression is associated with poor prognosis in node-positive breast cancer. Breast Cancer Res Treat 86:207–213. doi:10.1023/B:BREA.0000036787.59816.01
Spizzo G et al (2006) Overexpression of epithelial cell adhesion molecule (Ep-CAM) is an independent prognostic marker for reduced survival of patients with epithelial ovarian cancer. Gynecol Oncol 103:483–488. doi:10.1016/j.ygyno.2006.03.035
Stefan N, Martin-Killias P, Wyss-Stoeckle S, Honegger A, Zangemeister-Wittke U, Pluckthun A (2011) DARPins recognizing the tumor-associated antigen EpCAM selected by phage and ribosome display and engineered for multivalency. J Mol Biol 413:826–843. doi:10.1016/j.jmb.2011.09.016
Subramanian N, Sreemanthula JB, Balaji B, Kanwar JR, Biswas J, Krishnakumar S (2014) A strain-promoted alkyne-azide cycloaddition (SPAAC) reaction of a novel EpCAM aptamer-fluorescent conjugate for imaging of cancer cells. Chem Commun (Camb) 50:11810–11813. doi:10.1039/c4cc02996h
Tanaka Y et al (2013) Structural basis for the drug extrusion mechanism by a MATE multidrug transporter. Nature 496:247–251. doi:10.1038/nature12014
Trzpis M, McLaughlin PM, de Leij LM, Harmsen MC (2007) Epithelial cell adhesion molecule: more than a carcinoma marker and adhesion molecule. Am J Pathol 171:386–395. doi:10.2353/ajpath.2007.070152
Went P, Dirnhofer S, Salvisberg T, Amin MB, Lim SD, Diener PA, Moch H (2005) Expression of epithelial cell adhesion molecule (EpCam) in renal epithelial tumors. Am J Surg Pathol 29:83–88
Went P et al (2006) Frequent high-level expression of the immunotherapeutic target Ep-CAM in colon, stomach, prostate and lung cancers. Br J Cancer 94:128–135. doi:10.1038/sj.bjc.6602924
Yamagishi Y, Shoji I, Miyagawa S, Kawakami T, Katoh T, Goto Y, Suga H (2011) Natural product-like macrocyclic N-methyl-peptide inhibitors against a ubiquitin ligase uncovered from a ribosome-expressed de novo library. Chem Biol 18:1562–1570. doi:10.1016/j.chembiol.2011.09.013
Zielonka S et al (2014) Shark Attack: high affinity binding proteins derived from shark vNAR domains by stepwise in vitro affinity maturation. J Biotechnol 191:236–245. doi:10.1016/j.jbiotec.2014.04.023
Acknowledgments
We thank Drs. M. Ohuchi and T. Yoshitomi for supporting in the preparation of the manuscript. This work was supported by Ministry of Health, Labor and Welfare Grants-in-aid for Scientific Research programs from 2010 to 2012 to H. S., Y. T., and S. K., Grants-in-Aid for JSPS Fellows (7734) to K. I., and partly grants of Japan Society for the Promotion of Science Grants-in-Aid for Young Scientists (A) (24681047) to T. K., and Challenging Exploratory Research (15K12739) to Y. G.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Iwasaki, K., Goto, Y., Katoh, T. et al. A Fluorescent Imaging Probe Based on a Macrocyclic Scaffold That Binds to Cellular EpCAM. J Mol Evol 81, 210–217 (2015). https://doi.org/10.1007/s00239-015-9710-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-015-9710-z