Abstract
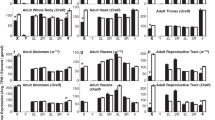
In Drosophila, there is a consistent deficit of male-biased genes on the X chromosome. It has been suggested that male-biased genes may evolve from initially unbiased genes as a result of increased expression levels in males. If transcription rates are limited, a large increase in expression in the testis may be harder to achieve for single-copy X-linked genes than for autosomal genes, because they are already hypertranscribed due to dosage compensation. This hypothesis predicts that the larger the increase in expression required to make a male-biased gene, the lower the chance of this being achievable if it is located on the X chromosome. Consequently, highly expressed male-biased genes should be located on the X chromosome less often than lowly expressed male-biased genes. This pattern is observed in our analysis of publicly available data, where microarray data or EST data are used to detect male-biased genes in D. melanogaster and to measure their expression levels. This is consistent with the idea that limitations in transcription rates may prevent male-biased genes from accumulating on the X chromosome.


Similar content being viewed by others
References
Betrán E, Thornton K, Long M (2002) Retroposed new genes out of the X in Drosophila. Genome Res 12:1854–1859
Charlesworth B (1978) Model for evolution of Y chromosomes and dosage compensation. Proc Natl Acad Sci USA 75:5618–5622
Charlesworth B, Coyne JA, Barton NH (1987) The relative rates of evolution of sex-chromosomes and autosomes. Am Nat 130:113–146
Connallon T, Knowles LL (2005) Intergenomic conflict revealed by patterns of sex-biased gene expression. Trends Genet 21:495–499
Drummond DA, Wilke CO (2008) Mistranslation-induced protein misfolding as a dominant constraint on coding-sequence evolution. Cell 134:341–352
Ellegren H, Parsch J (2007) The evolution of sex-biased genes and sex-biased gene expression. Nat Rev Genet 8:689–698
Emerson JJ, Cardoso-Moreira M, Borevitz JO, Long M (2008) Natural selection shapes genome-wide patterns of copy-number polymorphism in Drosophila melanogaster. Science 320:1629–1631
Gupta V, Parisi M, Sturgill D, Nuttall R, Doctolero M, Dudko OK, Malley JD, Eastman PS, Oliver B (2006) Global analysis of X-chromosome dosage compensation. J Biol 5:3.1–3.10
Hense W, Baines JF, Parsch J (2007) X chromosome inactivation during Drosophila spermatogenesis. PLoS Biol 5:e273
Kaiser VB, Ellegren H (2006) Nonrandom distribution of genes with sex-biased expression in the chicken genome. Evol Int J Org Evol 60:1945–1951
Khil PP, Smirnova NA, Romanienko PJ, Camerini-Otero RD (2004) The mouse X chromosome is enriched for sex-biased genes not subject to selection by meiotic sex chromosome inactivation. Nat Genet 36:642–646
Kondrashov FA, Rogozin IB, Wolf YI, Koonin EV (2002) Selection in the evolution of gene duplications. Genome Biol 3:RESEARCH0008
Lercher MJ, Urrutia AO, Hurst LD (2003) Evidence that the human X chromosome is enriched for male-specific but not female-specific genes. Mol Biol Evol 20:1113–1116
Lin H, Gupta V, Vermilyea MD, Falciani F, Lee JT, O’Neill LP, Turner BM (2007) Dosage compensation in the mouse balances up-regulation and silencing of X-linked genes. PLoS Biol 5:e326
Nguyen DK, Disteche CM (2006) Dosage compensation of the active X chromosome in mammals. Nat Genet 38:47–53
Ohno S (1967) Sex chromosomes and sex-linked genes. Springer, New York
Parisi M, Nuttall R, Naiman D, Bouffard G, Malley J, Andrews J, Eastman S, Oliver B (2003) Paucity of genes on the Drosophila X chromosome showing male-biased expression. Science 299:697–700
Reinke V, Gil IS, Ward S, Kazmer K (2004) Genome-wide germline-enriched and sex-biased expression profiles in Caenorhabditis elegans. Development 131:311–323
Rice WR (1984) Sex chromosomes and the evolution of sexual dimorphism. Evolution 38:735–742
Schwab M (1999) Oncogene amplification in solid tumors. Semin Cancer Biol 9:319–325
Seoighe C, Wolfe KH (1999) Yeast genome evolution in the post-genome era. Curr Opin Microbiol 2:548–554
Straub T, Becker PB (2007) Dosage compensation: the beginning and end of generalization. Nat Rev Genet 8:47–57
Sturgill D, Zhang Y, Parisi M, Oliver B (2007) Demasculinization of X chromosomes in the Drosophila genus. Nature 450:238–241
Vicoso B, Charlesworth B (2006) Evolution on the X chromosome: unusual patterns and processes. Nat Rev Genet 7:645–653
Wang PJ, McCarrey JR, Yang F, Page DC (2001) An abundance of X-linked genes expressed in spermatogonia. Nat Genet 27:422–426
Wu C-I, Xu EY (2003) Sexual antagonism and X inactivation—the SAXI hypothesis. Trends Genet 19:243–247
Zhang Y, Sturgill D, Parisi M, Kumar S, Oliver B (2007) Constraint and turnover in sex-biased gene expression in the genus Drosophila. Nature 450:233–237
Acknowledgments
We thank two referees for very helpful comments on the manuscript. This work was funded by a Portuguese Foundation for Science and Technology scholarship to B.V., and B.C. was supported by the Royal Society.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Vicoso, B., Charlesworth, B. The Deficit of Male-Biased Genes on the D. melanogaster X Chromosome Is Expression-Dependent: A Consequence of Dosage Compensation?. J Mol Evol 68, 576–583 (2009). https://doi.org/10.1007/s00239-009-9235-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-009-9235-4