Abstract
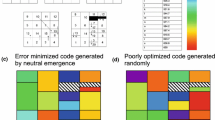
Studies on the origin of the genetic code compare measures of the degree of error minimization of the standard code with measures produced by random variant codes but do not take into account codon usage, which was probably highly biased during the origin of the code. Codon usage bias could play an important role in the minimization of the chemical distances between amino acids because the importance of errors depends also on the frequency of the different codons. Here I show that when codon usage is taken into account, the degree of error minimization of the standard code may be dramatically reduced, and shifting to alternative codes often increases the degree of error minimization. This is especially true with a high CG content, which was probably the case during the origin of the code. I also show that the frequency of codes that perform better than the standard code, in terms of relative efficiency, is much higher in the neighborhood of the standard code itself, even when not considering codon usage bias; therefore alternative codes that differ only slightly from the standard code are more likely to evolve than some previous analyses suggested. My conclusions are that the standard genetic code is far from being an optimum with respect to error minimization and must have arisen for reasons other than error minimization.


Similar content being viewed by others
References
L Achenbach-Ritcher R Gupta KO Stetter CR Woese (1987) ArticleTitleWere the original eubacteria thermophiles? Syst Appl Microbiol 9 34–39 Occurrence Handle11542087
SA Benner MA Cohen GH Gonnet (1994) ArticleTitleAmino acid substitution during functionally constrained divergent evolution of protein sequences Protein Eng 7 IssueID11 1323–1332 Occurrence Handle1:CAS:528:DyaK2MXit1Cku7k%3D Occurrence Handle7700864
MO Dayhoff RM Schwartz BC Orcutt (1978) A model of evolutionary change in proteins MO Dayhoff (Eds) Atlas of protein sequence and structure, Vol 5, Suppl 3 National Biomedical Research Foundation Washington 352
M Di Giulio (1989) ArticleTitleThe extension reached by the minimization of the polarity distances during the evolution of the genetic code J Mol Evol 29 IssueID4 288–293 Occurrence Handle1:CAS:528:DyaL1MXmt1Ogurc%3D Occurrence Handle2514270
M Di Giulio (1997a) ArticleTitleOn the origin of the genetic code J Theor Biol 187 IssueID4 573–581 Occurrence Handle1:CAS:528:DyaK2sXmtVGlsL0%3D
M Di Giulio (1997b) ArticleTitleThe origin of the genetic code Trends Biochem Sci 22 IssueID2 49–50 Occurrence Handle1:CAS:528:DyaK2sXht1Gnu78%3D
M Di Giulio (1999) ArticleTitleThe coevolution theory of the origin of the genetic code J Mol Evol 48 IssueID3 253–255 Occurrence Handle1:CAS:528:DyaK1MXhs1ymsr0%3D Occurrence Handle10627191
M Di Giulio (2000a) ArticleTitleGenetic code origin and the strength of natural selection J Theor Biol 205 659–661 Occurrence Handle10.1006/jtbi.2000.2115 Occurrence Handle1:CAS:528:DC%2BD3cXlsVSrsrw%3D
M Di Giulio (2000b) ArticleTitleThe origin of the genetic code Trends Biochem Sci 25 IssueID2 44 Occurrence Handle10.1016/S0968-0004(99)01522-4 Occurrence Handle1:CAS:528:DC%2BD3cXhsFejsbY%3D
M Di Giulio (2000c) ArticleTitleThe universal ancestor lived in a thermophilic or hyperthermophilic environment J Theor Biol 203 203–213 Occurrence Handle1:CAS:528:DC%2BD3cXhs12hsLk%3D
M Di Giulio (2001) ArticleTitleThe origin of the genetic code cannot be studied using measurements based on the PAM matrix because this matrix reflects the code itself, making any such analyses tautologous J Theor Biol 208 141–144 Occurrence Handle10.1006/jtbi.2000.2206 Occurrence Handle1:CAS:528:DC%2BD3MXjt1yitA%3D%3D Occurrence Handle11162059
CJ Epstein (1966) ArticleTitleRole of the amino-acid “code” and of selection for conformation in the evolution of proteins Nature 210 25–28 Occurrence Handle1:CAS:528:DyaF28XpsVOksg%3D%3D Occurrence Handle5956344
WM Fitch K Upper (1987) ArticleTitleThe phylogeny of tRNA sequences provides evidence for ambiguity reduction in the origin of the genetic code Cold Spring Harbor Symp Quant Biol 52 759–767 Occurrence Handle1:CAS:528:DyaL1cXlt1Cmtrc%3D Occurrence Handle3454288
SJ Freeland LD Hurst (1998) ArticleTitleThe genetic code is one in a million J Mol Evol 47 238–248 Occurrence Handle1:CAS:528:DyaK1cXmt1ansrs%3D Occurrence Handle9732450
SJ Freeland RD Knight LF Landweber (2000a) ArticleTitleMeasuring adaptation within the genetic code Trends Biochem Sci 25 IssueID2 44–45 Occurrence Handle1:CAS:528:DC%2BD3cXhsFejtr4%3D
SJ Freeland RD Knight LF Landweber LD Hurst (2000b) ArticleTitleEarly fixation of an optimal genetic code Mol Biol Evol 17 IssueID4 511–518 Occurrence Handle1:CAS:528:DC%2BD3cXisVSgt70%3D
DG George WC Barker LT Hunt (1990) ArticleTitleMutation data matrix and its uses Meth Enzym 183 333–351 Occurrence Handle1:CAS:528:DyaK3MXivFGg Occurrence Handle2314281
D Haig LD Hurst (1991) ArticleTitleA quantitative measure of error minimization in the genetic code J Mol Evol 33 IssueID5 412–417 Occurrence Handle1:CAS:528:DyaK3MXmsleqtrs%3D Occurrence Handle1960738
OP Judson D Haydon (1999) ArticleTitleThe genetic code: What is it good for? An analysis of the effects of selection pressures on genetic codes J Mol Evol 49 IssueID5 539–550 Occurrence Handle1:CAS:528:DyaK1MXnsVajsbw%3D Occurrence Handle10552035
RD Knight SJ Freeland LF Landweber (1999) ArticleTitleSelection, history and chemistry: The three faces of the genetic code Trends Biochem Sci 24 IssueID6 241–247 Occurrence Handle10.1016/S0968-0004(99)01392-4 Occurrence Handle1:CAS:528:DyaK1MXks1ansLg%3D Occurrence Handle10366854
AD McLachlan (1971) ArticleTitleTests for comparing related amino-acid sequences. Cytochrome c and cytochrome c 551 J Mol Biol 61 409–424 Occurrence Handle1:CAS:528:DyaE38XhsF2k Occurrence Handle5167087
J Overington D Donnelly MS Johnson et al. (1992) ArticleTitleEnvironment-specific amino-acid substitution tables - tertiary templates and prediction of protein folds Protein Sci 1 IssueID2 216–226 Occurrence Handle1:CAS:528:DyaK38XksVeqtb4%3D Occurrence Handle1304904
RP Riek MD Handschumacher SS Sung M Tan MJ Glynias MD Schluchter J Novotny RM Graham (1995) ArticleTitleEvolutionary conservation of both the hydrophilic and hydrophobic nature of transmembrane residues J Theor Biol 172 IssueID3 245–258 Occurrence Handle10.1006/jtbi.1995.0021 Occurrence Handle1:CAS:528:DyaK2MXkslGmsrk%3D Occurrence Handle7715195
JL Risler MO Delorme H Delacroix A Henaut (1988) ArticleTitleAmino acid substitutions in structurally related proteins. A pattern recognition approach. Determination of a new and efficient scoring matrix J Mol Biol 204 IssueID4 1019–1029 Occurrence Handle1:CAS:528:DyaL1MXptV2jtg%3D%3D
TM Sonneborn (1965) Degeneracy of the genetic code: extent, nature and genetic implications. Academic Press, New York Academic Press New York
CR Woese (1965) ArticleTitleOn the evolution of the genetic code Proc Natl Acad Sci USA 54 1546–1552 Occurrence Handle1:CAS:528:DyaF28Xltlagsg%3D%3D Occurrence Handle5218910
CR Woese (1973) ArticleTitleEvolution of genetic code J Naturwiss 60 447–459 Occurrence Handle1:CAS:528:DyaE2cXksVOmsA%3D%3D
CR Woese (1987) ArticleTitleBacterial evolution Microbiol Rev 51 221–271
CR Woese (1998) ArticleTitleThe universal ancestor Proc Natl Acad Sci USA 95 6854–6859 Occurrence Handle1:CAS:528:DyaK1cXjslynu7w%3D Occurrence Handle9618502
JT Wong (1975) ArticleTitleA co-evolution theory of the genetic code Proc Natl Acad Sci USA 72 IssueID5 1909–1912 Occurrence Handle1:CAS:528:DyaE2MXkt1ertbg%3D Occurrence Handle1057181
F Wright (1990) ArticleTitleThe ‘effective number of codons’ used in a gene Gene 87 23–29 Occurrence Handle10.1016/0378-1119(90)90491-9 Occurrence Handle1:CAS:528:DyaK3cXktVWmsbo%3D Occurrence Handle2110097
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Martin Kreitman]
Rights and permissions
About this article
Cite this article
Archetti, M. Codon Usage Bias and Mutation Constraints Reduce the Level of ErrorMinimization of the Genetic Code. J Mol Evol 59, 258–266 (2004). https://doi.org/10.1007/s00239-004-2620-0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-004-2620-0