Abstract
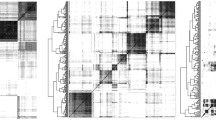
The grasses (Poaceae) represent a monophyletic lineage that arose about 70 million years ago. The lineage contains about 10,000 species that differ widely in morphology and physiology. Species show striking differences in genome size, a feature important in the context of conservation of gene content and order (synteny and colinearity) and in the extension of genomic information directly from one grass species to another using comparative approaches. Grass diversification has been a contentious issue, as the exact branching order of the various subfamilies has been difficult to establish with standard methods. This motivated an evolutionary study of deep phylogenetic relationships based on the structure of coding and non-coding RNA molecules and on chromosomal rearrangements. Phylogenetic relationships in the grass family were inferred directly from the structure of RNA using cladistic principles and considerations in statistical mechanics. Coded attributes describing topological and thermodynamic information embedded in RNA molecules were treated as linearly ordered multi-state characters and were polarized by fixing the direction of character transformation toward molecular order. Intrinsically rooted phylogenies derived from the structure of signal recognition particle (SRP) RNA, the mRNA encoded by the early nodulation gene enod40, the small subunit of ribosomal RNA (rRNA), and the internal transcribed spacer ITS1 of rRNA established an order for the diversification of major grass lineages, suggesting a sister relationship of the Pooideae and the PACCAD clade. This same conclusion was reached when large-scale chromosomal rearrangements derived from the comparative genetic mapping of cereal genomes were studied. Chromosomal complements aligned in the most parsimonious manner allowed identification and coding of characters depicting chromosomal translocations, insertions, and linkage block arrangements and the reconstruction of phylogenetic trees based on large-scale chromosomal structure. Congruent reconstruction of deep branching relationships using geometrical and statistical features of RNA structure and orthology and large scale chromosomal recombination events support assumptions of polarization in character argumentation, and fail to falsify the claim that extant grass chromosomes can be considered combinations of linkage blocks of an ancestor of the rice genome. Congruence also suggests that the universal tendency toward order in RNA and the search for the most parsimonious organization of be genome architecture appear to be mutually supported drivers of molecular evolution. The study clarifies the relationship of major clades in the grasses, shows that phylogenetic history can be reconstructed effectively from the combinatorial exchange of chromosomal linkage blocks, and reveals considerable phylogenetic signal embedded in the structure of signal polypeptide-coding mRNA molecules, describing an instance where mRNA structure is the subject of strong evolutionary constraint.







Similar content being viewed by others
References
LW Ancel W Fontana (2000) ArticleTitlePlasticity, evolvability, and modularity in RNA J Exp Zool 288 242–283 Occurrence Handle10.1002/1097-010X(20001015)288:3<242::AID-JEZ5>3.0.CO;2-O Occurrence Handle1:CAS:528:DC%2BD3cXnsFChs70%3D
MD Bennet IJ Leitch (2001) Plant DNA C-value database (release 1.0, September 2001) Royal Botanic Gardens Kew
JL Bennetzen EA Kellogg (1997) ArticleTitleDo plants have a one-way ticket to genomic obesity? Plant Cell 9 1509–1514 Occurrence Handle1:CAS:528:DyaK2sXmsFelt7k%3D Occurrence Handle12237393
C Brunel R Marquet P Romby C Ehresmann (2002) ArticleTitleRNA loop-loop interactions as dynamic functional motifs Biochimie 84 925–944 Occurrence Handle1:CAS:528:DC%2BD38XovFersro%3D Occurrence Handle12458085
HN Bryant (1991) ArticleTitleThe polarization of character transformations in phylogenetic systematics: role of axiomatic and auxiliary assumptions Syst Zool 40 433–445
IV ES Buckler A Ippolito TP Holtsford (1997) ArticleTitleThe evolution of ribosomal DNA: divergent paralogues and phylogenetic implications Genetics 145 821–832 Occurrence Handle1:CAS:528:DyaK2sXmsVaktLc%3D Occurrence Handle9055091
G Caetano-Anollés (2001) ArticleTitleNovel strategies to study the role of mutation and nucleic acid structure in evolution Plant Cell Tissue Org Cult 67 115–132
G Caetano-Anollés (2002a) ArticleTitleEvolved RNA secondary structure and the rooting of the universal tree J Mol Evol 54 333–345
G Caetano-Anollés (2002b) ArticleTitleTracing the evolution of RNA structure in ribosomes Nucleic Acids Res 30 2575–2587
JH Cate MM Yusupov GZ Yusupova TN Earnest HF Noller (1999) ArticleTitleX-ray crystal structure of 70S ribosome functional complexes Science 285 2095–2104 Occurrence Handle1:CAS:528:DyaK1MXmt1yrt74%3D Occurrence Handle10497122
WD Clayton SA Renvoize (1986) Genera Graminium: Grasses of the world. Kew Bulletin Additional Series XIII. Royal Botanical Gardens, Kew Her Majesty’s Stationery Office London
KM Devos MD Gale (2000) ArticleTitleGenome relationships: the grass model in current research Plant Cell 12 637–646 Occurrence Handle1:CAS:528:DC%2BD3cXjvFyhurw%3D Occurrence Handle10810140
KM Devos TS Pittaway A Reynolds MD Gale (2000) ArticleTitleComparative mapping reveals a complex relationship between the pearl millet genome and those of foxtail millet and rice Theor Appl Genet 100 190–198 Occurrence Handle1:CAS:528:DC%2BD3cXhs1Gmsbk%3D
JS Farris M Kallersjö AG Kluge C Bull (1995) ArticleTitleTesting significance of incongruence Cladistics 10 315–319 Occurrence Handle10.1006/clad.1994.1021
J Felsenstein (1985) ArticleTitleConfidence limits on phylogenies: an approach using the bootstrap Evolution 39 783–791
W Fontana (2002) ArticleTitleModelling ‘evo-devo’ with RNA BioEssays 24 1164–1177 Occurrence Handle1:CAS:528:DC%2BD38Xpsleltb4%3D Occurrence Handle12447981
W Fontana DA Konings PF Stadler P Schuster (1993) ArticleTitleStatistics of RNA secondary structures Biopolymers 33 1389–1404 Occurrence Handle1:CAS:528:DyaK3sXmtF2ku7s%3D Occurrence Handle7691201
W Fontana P Schuster (1998) ArticleTitleContinuity in evolution: on the nature of transitions Science 280 1451–1455 Occurrence Handle1:CAS:528:DyaK1cXjtlars78%3D Occurrence Handle9603737
MD Gale KM Devos (1998a) ArticleTitlePlant comparative genetics after 10 years Science 282 656–659 Occurrence Handle1:CAS:528:DyaK1cXmvFOrtrg%3D
MD Gale KM Devos (1998b) ArticleTitleComparative genetics in the grasses Proc Natl Acad Sci USA 95 1971–1974 Occurrence Handle1:CAS:528:DyaK1cXhslejsLk%3D
G Girard A Roussis AP Gultyaev CWA Preij HP Paink (2003) ArticleTitleStructural motifs in the RNA encoded by the early nodulation gene enod40 of soybean Nucleic Acids Res 31 5003–5015 Occurrence Handle1:CAS:528:DC%2BD3sXmvVWrur8%3D Occurrence Handle12930950
GP Gladyshev YA Ershov (1982) ArticleTitlePrinciples of the thermodynamics of biological systems J Theor Biol 94 301–343 Occurrence Handle1:CAS:528:DyaL38Xhtlyns74%3D Occurrence Handle7078210
InstitutionalAuthorNameGPWG (Grass Phylogeny Working Group) (2001) ArticleTitlePhylogeny and subfamilial classification of the grasses (Poaceae) Ann Missouri Bot Gard 88 373–457
PA Gultyaev FHD van Batenburg CWA Pleij (2002) ArticleTitleSelective pressures on RNA hairpins in vivo and in vitro J Mol Evol 54 1–8 Occurrence Handle1:CAS:528:DC%2BD3MXovFKnsrg%3D Occurrence Handle11734892
RK Hamby EA Zimmer (1988) ArticleTitleRibosomal RNA sequences for inferring phylogeny within the grass family (Poaceae) Plant Syst Evol 160 29–37 Occurrence Handle1:CAS:528:DyaL1cXls1Ogs7w%3D
T Hermann DJ Patel (1999) ArticleTitleStitching together RNA tertiary architectures J Mol Biol 294 829–849 Occurrence Handle1:CAS:528:DyaK1MXnslCqt7o%3D Occurrence Handle10588890
PG Higgs (1993) ArticleTitleRNA secondary structure: a comparison of real and random sequences J Phys I France 3 43–59 Occurrence Handle1:CAS:528:DyaK3sXit1yns7o%3D
PG Higgs (1995) ArticleTitleThermodynamic properties of transfer RNA: a computational study J Chem Soc Faraday Trans 91 2531– 2540 Occurrence Handle1:CAS:528:DyaK2MXns1Siu78%3D
IL Hofacker M Fekete PF Stadler (2002) ArticleTitleSecondary structure prediction for aligned RNA sequences J Mol Biol 319 1059–1066 Occurrence Handle10.1016/S0022-2836(02)00308-X Occurrence Handle1:CAS:528:DC%2BD38XkslantLk%3D Occurrence Handle12079347
IL Hofacker W Fontana PF Stadler LS Bonhoeffer M Tacker P Schuster (1994) ArticleTitleFast folding and comparison of RNA secondary structures Monatshefte Chem 125 167–188 Occurrence Handle1:CAS:528:DyaK2cXivFaht7k%3D
C Hsiao SWL Jacobs NJ Chatterton KH Asay (1999) ArticleTitleA molecular phylogeny of the grass family (Poaceae) based on the sequences of nuclear ribosomal DNA (ITS) Aust Syst Bot 11 667–688
M Huynen R Gutell D Konings (1997) ArticleTitleAssessing the reliability of RNA folding using statistical mechanics J Mol Biol 267 1104–1112 Occurrence Handle1:CAS:528:DyaK2sXjtFSqsbk%3D Occurrence Handle9150399
MA Huynen PF Stadler W Fontana (1996) ArticleTitleSmoothness with ruggedness: the role of neutrality in adaptation Proc Natl Acad Sci USA 86 397–401
BF Jacobs JD Kingston LL Jacobs (1999) ArticleTitleThe origin of grass-dominated ecosystems Ann Missouri Bot Gard 86 590–643
BD James GJ Olsen NR Pace (1989) ArticleTitlePhylogenetic comparative analysis of RNA secondary structure Methods Enzymol 180 227–239 Occurrence Handle1:CAS:528:DyaK3cXitFent7c%3D Occurrence Handle2482415
SA Kauffmann (1993) The origins of order Oxford University Press New York
RJ Keenan DM Freymann RM Stroud P Walter (2001) ArticleTitleThe signal recognition particle Annu Rev Biochem 70 755–775 Occurrence Handle1:CAS:528:DC%2BD3MXlsVehtLY%3D Occurrence Handle11395422
EA Kellogg (1998) ArticleTitleRelationships of cereal crops and other grasses Proc Natl Acad Sci USA 95 2005–2010 Occurrence Handle1:CAS:528:DyaK1cXhslejsbg%3D Occurrence Handle9482825
EA Kellogg (2001) ArticleTitleEvolutionary history of the grasses Plant Physiol 125 1198–1205 Occurrence Handle1:CAS:528:DC%2BD3MXitFWrt70%3D Occurrence Handle11244101
W Kennard R Phillips R Porter A Grombacher (1999) ArticleTitleA comparative map of wild rice (Zizania palustris L 2n=2x=30). Theor Appl Genet 99 793–799 Occurrence Handle1:CAS:528:DyaK1MXnsVKjtL4%3D
E Kierzek E Biala R Kierzek (2001) ArticleTitleElements of thermodynamics in RNA evolution Acta Biochim Polonica 48 485–493 Occurrence Handle1:CAS:528:DC%2BD3MXlvVKgtrg%3D
DAM Konings RR Gutell (1995) ArticleTitleA comparison of thermodynamic foldings with comparatively derived structures of 16S and 16S-like rRNA RNA 1 559–574 Occurrence Handle1:CAS:528:DyaK2MXot1Wqsb8%3D Occurrence Handle7489516
K Le SY Zhang JV Maizel SuffixJr (2002) ArticleTitleRNA molecules with structure dependent functions are uniquely folded Nucleic Acids Res 30 3574–3582 Occurrence Handle12177299
WP Maddison MJ Donoghue DR Maddison (1984) ArticleTitleOutgroup analysis and parsimony Syst Zool 33 83–103
WP Maddison DR Maddison (1999) MacClade: analysis of phylogeny and character evolution, Version 3.08 Sinauer Associates Sunderland, MA
DH Mathews J Sabina M Zuker DH Turner (1999) ArticleTitleExpanded sequence dependence of thermodynamic parameters provides robust prediction of RNA secondary structure J Mol Biol 288 911–940 Occurrence Handle1:CAS:528:DyaK1MXjtVers7Y%3D Occurrence Handle10329189
Y Matsuoka Y Yamazaki Y Ogihara K Tsunewaki (2002) ArticleTitleWhole chloroplast genome comparison of rice, maize, and wheat: implications for chloroplast gene diversification and phylogeny of cereals Mol Biol Evol 19 2084–2091 Occurrence Handle1:CAS:528:DC%2BD38Xps12hsrY%3D Occurrence Handle12446800
JS McCaskill (1990) ArticleTitleThe equilibrium partition function and base pair binding probabilities fo RNA secondary structures Biopolymers 29 1105–1119 Occurrence Handle1:CAS:528:DyaK3cXkt1Snurk%3D Occurrence Handle1695107
RDM Page EC Holmes (1998) Molecular evolution: A phylogenetic approach Blackwell Science Oxford
DA Petrov (2001) ArticleTitleEvolution of genome size: new approaches to an old problem Trends Genet 17 23–28 Occurrence Handle1:CAS:528:DC%2BD3MXktl2ksw%3D%3D Occurrence Handle11163918
DD Pollock (2003) ArticleTitleThe Zuckerkandl Prize: structure and evolution J Mol Evol 56 375–376 Occurrence Handle1:CAS:528:DC%2BD3sXisV2qsb4%3D
E Rivas SR Eddy (2000) ArticleTitleSecondary structure alone is generally not statistically significant for the detection of noncoding RNAs Bioinformatics 16 583–605 Occurrence Handle1:CAS:528:DC%2BD3cXosFSqs7k%3D Occurrence Handle11038329
H Rohrig J Schmidt E Miklashevichs J Schell M John (2002) ArticleTitleSoybean enod40 encodes two peptides that bind to sucrose synthase Proc Natl Acad Sci USA 99 1915–1920 Occurrence Handle1:CAS:528:DC%2BD38XitVSrs7w%3D Occurrence Handle11842184
MA Rosenblad J Gorodkin B Knudsen C Zwieb T Samuelsson (2003) ArticleTitleSRPDB: Signal recognition particle database Nucleic Acids Res 31 363–364 Occurrence Handle1:CAS:528:DC%2BD3sXhvFSmtL8%3D Occurrence Handle12520023
G Savva J Dicks IN Roberts (2003) ArticleTitleCurrent approaches to whole genome phylogenetic analysis Brief Bioinform 4 63–74 Occurrence Handle1:CAS:528:DC%2BD3sXktVWgtLc%3D Occurrence Handle12715835
EA Schultes DP Bartel (2000) ArticleTitleOne sequence, two ribozymes: implications for the emergence of new ribozyme folds Science 289 448–452 Occurrence Handle1:CAS:528:DC%2BD3cXlsV2gtLg%3D Occurrence Handle10903205
EA Schultes PT Hraber TH LaBean (1999) ArticleTitleEstimating the contributions of selection and self-organization in RNA secondary structure J Mol Evol 49 76–83 Occurrence Handle1:CAS:528:DyaK1MXkt1Oktrs%3D Occurrence Handle10368436
P Schuster W Fontana PF Stadler IL Hofacker (1994) ArticleTitleFrom sequences to shapes and back: a case study in RNA secondary structure Proc R Soc London Ser B 255 279–284 Occurrence Handle1:CAS:528:DyaK2cXmslCmsL0%3D
P Schuster PF Stadler A Renner (1997) ArticleTitleRNA structures and folding: from conventional to new issues in structure predictions Curr Opin Struct Biol 7 229–235 Occurrence Handle1:CAS:528:DyaK2sXisFymsbw%3D Occurrence Handle9094330
C Sousa C Johansson H CharonManyani C Sautter A Kondorosi M Crespi (2001) ArticleTitleTranslation and structural requirements of the early nodulin gene enod40, a short-open reading frame-containing RNA, for elicitation of a cell-specific growth response in the alfalfa root cortex Mol Cell Biol 21 354–366 Occurrence Handle1:CAS:528:DC%2BD3MXptVSk Occurrence Handle11113209
W Steffens D Digby (1999) ArticleTitlemRNA have greater negative folding free energies than shuffled or codon choice randomized sequences Nucleic Acids Res 27 1578–1584 Occurrence Handle10075987
G Stegger H Hofman J Fortsch HJ Gross JW Randles HL Sanger D Riesner (1984) ArticleTitleConformational transitions in viroids and virusoids: comparison of results from energy minimization algorithm and from experimental data J Biomol Struct Dynam 2 543–571
DL Swofford (1998) Phylogenetic analysis using parsimony and other programs (PAUP*), version 4.0 Sinauer Associates Sunderland, MA
M Tacker PF Stadler EG Bornberg-Bauer IL Hofacker P Schuster (1996) ArticleTitleAlgorithm independent properties of RNA secondary structure predictions Eur Biophys J 25 115–130 Occurrence Handle1:CAS:528:DyaK2sXhtFGksro%3D
F Takaiwa K Oono Y lida M Sugiura (1995) ArticleTitleThe complete nucleotide sequence of a rice 25S rRNA gene Gene 37 255–259
K Thiele (1993) ArticleTitleThe holy grail of the perfect character: the cladistic treatment of morphometric data Cladistics 9 275–304 Occurrence Handle10.1006/clad.1993.1020
JL Thorley RDM Page (2000) ArticleTitleRadCon: phylogenetic tree comparison and consensus Bioinformatics 16 486–487 Occurrence Handle10.1093/bioinformatics/16.5.486 Occurrence Handle1:CAS:528:DC%2BD3cXlvVKqt7c%3D Occurrence Handle10871273
S Washietl IL Hofacker (2004) ArticleTitleConsensus folding of aligned sequences as a new measure for the detection of functional RNAs by comparative genomics J Mol Biol 342 19–30 Occurrence Handle1:CAS:528:DC%2BD2cXmslegsLw%3D Occurrence Handle15313604
L Watson MJ Dallwitz (1992) The grass genera of the world CAB International Wallingford, Oxon, UK
WC Wheeler (1990) ArticleTitleCombinatorial weights in phylogenetic analysis A statistical parsimony procedure. Cladistics 6 269–275
M Wilkinson JL Thorley P Upchurch (2000) ArticleTitleA chain is no longer than its weakest link: double decay analysis of phylogenetic hypothesis Syst Biol 49 754–776 Occurrence Handle10.1080/106351500750049815 Occurrence Handle1:STN:280:DC%2BD38zntVOisw%3D%3D Occurrence Handle12116438
S Wright (1932) ArticleTitleThe roles of mutation, inbreeding, crossbreeding and selection in evolution Proc Sixth Intl Congr Genet 1 356–366
J Wuyts Y Van de Peers R Wachter ParticleDe (2001) ArticleTitleDistribution of substitution rates and location of insertion sites in the tertiary structure of ribosomal RNA Nucleic Acids Res 29 5017–5028 Occurrence Handle1:CAS:528:DC%2BD38XmslWgsQ%3D%3D Occurrence Handle11812832
MM Yusupov GZ Yusupova A Baucom K Lieberman TN Earnest JHD Cate HF Noller (2001) ArticleTitleCrystal structure of the ribosome at 5.5 Å resolution Science 292 883–896 Occurrence Handle10.1126/science.1060089 Occurrence Handle1:CAS:528:DC%2BD3MXjsVSrsb4%3D Occurrence Handle11283358
H Zhang J Jia MD Gale KM Devos (1998) ArticleTitleRelationships between the chromosomes of Aegilops umbellulata and wheat Theor Appl Genet 96 69–75 Occurrence Handle1:CAS:528:DyaK1cXhtlSksbo%3D
M Zuker (1989) ArticleTitleOn finding all suboptimal foldings of an RNA molecule Science 244 48–52 Occurrence Handle1:CAS:528:DyaL1MXkt1SnsbY%3D Occurrence Handle2468181 Occurrence HandleMR989985
Acknowledgments
I gratefully acknowledge G.E. Caetano-Anollés for support and help in character coding, V. Knudsen, USIT, University of Oslo, for computer programs, and D. Pollock and an anonymous reviewer for valuable suggestions. Research was supported by CSREES-USDA-ILLU Hatch Project 802-316 and NSF Project MCB-0343126. Any opinious, findings, and conclusions or recommendations expressed in this material are those of the author and do not necessarily reflect the views of the fundings agencies.
Author information
Authors and Affiliations
Corresponding author
Additional information
Reviewing Editor: Dr. David Pollock
Rights and permissions
About this article
Cite this article
Caetano-Anollés, G. Grass Evolution Inferred from Chromosomal Rearrangements and Geometrical and Statistical Features in RNA Structure. J Mol Evol 60, 635–652 (2005). https://doi.org/10.1007/s00239-004-0244-z
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-004-0244-z