Abstract
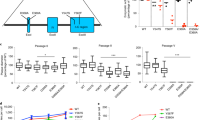
Two lines of the bacteriophage T7 were grown to fix mutations indiscriminately, using a combination of population bottlenecks and mutagenesis. Complete genome sequences revealed 404 and 299 base substitutions in the two lines, the largest number characterized in functional microbial genomes so far. Missense substitutions outnumbered silent substitutions. Silent substitutions occurred at similar rates between essential and nonessential genes, but missense substitutions occurred at a higher rate in nonessential genes than in essential genes, as expected if they were less deleterious in the nonessential genes. Viral fitness declined during this protocol, and subsequent passaging of each mutated line in large population sizes restored some of the lost fitness. Substitution levels during these recoveries were less than 6% of those during the bottleneck phase, and only two changes during recovery were reversions of the original mutations. Exchanges of genomic fragments between the two recovered lines revealed that fitness effects of some substitutions were not additive—that interactions were accumulating which could lead to incompatibility between the diverged genomes. Based on these results, unprecedented high rates of nucleotide and functional divergence in viral genomes should be attainable experimentally by using repeated population bottlenecks at a high mutation rate interspersed with recovery.


Similar content being viewed by others
References
MR Badgett A Auer LE Carmichael CR Parrish JJ Bull (2002) ArticleTitleEvolutionary dynamics of viral attenuation. J Virol 76 10524–10529 Occurrence Handle10.1128/JVI.76.20.10524-10529.2002 Occurrence Handle12239331
MJ Brauer (2000) Geometry and genetics of microbial adaptation. Integrative Biology. University of Texas Austin
JJ Bull CW Cunningham IJ Molineux M Badgett DM Hillis (1993) ArticleTitleExperimental molecular evolution of bacteriophage T7. Evolution 47 993–1007 Occurrence Handle1:CAS:528:DC%2BD3cXkslyksbc%3D Occurrence Handle10853957
JJ Bull MR Badgett HA Wichman JP Huelsenbeck DM Hillis A Gulati C Ho IJ Molineux (1997) ArticleTitleExceptional convergent evolution in a virus. Genetics 147 1497–1507 Occurrence Handle9409816
JJ Bull MR Badgett HA Wichman (2000) ArticleTitleBig-benefit mutations in a bacteriophage inhibited with heat. Mol Biol Evol 17 942–950 Occurrence Handle10833201
JJ Bull BR Levin T DeRouin N Walker CA Bloch (2002) ArticleTitleDynamics of success and failure in phage and antibiotic therapy in experimental infections. BMC Microbiol 2 35 Occurrence Handle10.1186/1471-2180-2-35 Occurrence Handle12453306
CL Burch L Chao (1999) ArticleTitleEvolution by small steps and rugged landscapes in the RNA virus phi6. Genetics 151 921–927 Occurrence Handle10049911
L Chao (1990) ArticleTitleFitness of RNA virus decreased by Muller’s ratchet. Nature 348 454–455 Occurrence Handle1:STN:280:By6D287ovV0%3D
WD Crill HA Wichman JJ Bull (2000) ArticleTitleEvolutionary reversals during viral adaptation to alternating hosts. Genetics 154 27–37 Occurrence Handle10628966
JA de Visser (2002) ArticleTitleThe fate of microbial mutators. Microbiology 148 1247–1252 Occurrence Handle11988499
JJ Dunn FW Studier (1983) ArticleTitleComplete nucleotide sequence of bacteriophage T7 DNA and the locations of T7 genetic elements. J Mol Biol 166 477–535 Occurrence Handle6864790
SF Elena RE Lenski (1997) ArticleTitleTest of synergistic interactions among deleterious mutations in bacteria. Nature 390 395–398 Occurrence Handle10.1038/37108 Occurrence Handle9389477
C Escarmis M Davila N Charpentier A Bracho A Moya E Domingo (1996) ArticleTitleGenetic lesions associated with Muller’s ratchet in an RNA virus. J Mol Biol 264 255–267 Occurrence Handle10.1006/jmbi.1996.0639 Occurrence Handle8951375
C Escarmis M Davila E Domingo (1999) ArticleTitleMultiple molecular pathways for fitness recovery of an RNA virus debilitated by operation of Muller’s ratchet. J Mol Biol 285 495–505 Occurrence Handle10.1006/jmbi.1998.2366 Occurrence Handle9878424
F Fenner J Cairns (1959) Variation in virulence in relation to adaptation to new hosts. FM Burnet WM Stanley (Eds) Animal viruses. Academic Press New York 225–249
SJ Flint (2000) Principles of virology: Molecular biology, pathogenesis, and control. ASM Press Washington, DC
DM Hillis JJ Bull ME White MR Badgett IJ Molineux (1992) ArticleTitleExperimental phylogenetics: generation of a known phylogeny. Science 255 589–592 Occurrence Handle1:CAS:528:DyaK38Xht1als7Y%3D Occurrence Handle1736360
DM Hillis JP Huelsenbeck CW Cunningham (1994) ArticleTitleApplication and accuracy of molecular phylogenies. Science 264 671–677 Occurrence Handle8171318
MJ Horsfall AJ Gordon PA Burns M Zielenska GM van der Vliet BW Glickman (1990) ArticleTitleMutational specificity of alkylating agents and the influence of DNA repair. Environ Mol Mutagen 15 102–22
RE Lenski CL Winkworth MA Riley (2003) ArticleTitleRates of DNA sequence evolution in experimental populations of Escherichia coli during 20,000 generations. J Mol Evol 56 498–508 Occurrence Handle10.1007/s00239-002-2423-0 Occurrence Handle12664169
W-H Li (1997) Molecular evolution. Sinauer Associates Sunderland, MA
JH Miller (1998) ArticleTitleMutators in Escherichia coli. Mutat Res 409 99–106 Occurrence Handle10.1016/S0921-8777(98)00049-4 Occurrence Handle1:CAS:528:DyaK1cXns1Ojsbs%3D Occurrence Handle9875286
FB Moore DE Rozen RE Lenski (2000) ArticleTitlePervasive compensatory adaptation in Escherichia coli. Proc R Soc Lond B Biol Sci 267 515–522 Occurrence Handle10.1098/rspb.2000.1030 Occurrence Handle10737410
D Rokyta MR Badgett IJ Molineux JJ Bull (2002) ArticleTitleExperimental genomic evolution: Extensive compensation for loss of DNA ligase activity in a virus. Mol Biol Evol 19 230–238 Occurrence Handle11861882
FW Studier (1979) ArticleTitleRelationships among different strains of T7 and among T7-related bacteriophages. Virology 95 70–84 Occurrence Handle375582
S Tabor CC Richardson (1989) ArticleTitleSelective inactivation of the exonuclease activity of bacteriophage T7 DNA polymerase by in vitro mutagenesis. J Biol Chem 264 6447–6458 Occurrence Handle2703498
JP Vartanian M Henry S Wain-Hobson (2001) ArticleTitleSimulating pseudogene evolution in vitro: Determining the true number of mutations in a lineage. Proc Natl Acad Sci USA 98 13172–13176 Occurrence Handle10.1073/pnas.221334898 Occurrence Handle11687609
HA Wichman LA Scott CD Yarber JJ Bull (2000) ArticleTitleExperimental evolution recapitulates natural evolution. Philos Trans R Soc Lond B Biol Sci 355 1677–1684 Occurrence Handle10.1098/rstb.2000.0731 Occurrence Handle11127915
Acknowledgements
We thank R. Springman for helping resolve some of the sequence ambiguities. This work was supported by Grants: NSF 9726902, NIH GM 57756 to J.J.B. and GM 32095 to I.J.M. J.J.B. is supported as the Friedrich Miescher Regents Professor. We thank H. Wichman and the reviewers for many useful comments.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bull, J., Badgett, M., Rokyta, D. et al. Experimental Evolution Yields Hundreds of Mutations in a Functional Viral Genome . J Mol Evol 57, 241–248 (2003). https://doi.org/10.1007/s00239-003-2470-1
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-003-2470-1