Abstract
This study was aimed at developing the discriminant models for distinguishing the tomato seeds based on texture parameters of the outer surface of seeds calculated from the images (scans) converted to individual color channels R, G, B, L, a, b, X, Y, Z. The seeds of tomatoes ‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’, Sacher F1 and Sandoline F1 were discriminated in pairs. The highest results were observed for models built based on sets of textures selected individually from color channels R, L and X and sets of textures selected from all color channels. In all cases, the tomato seeds ‘Green Zebra’ and ‘Ożarowski’ were discriminated with the highest average accuracy equal to 97% for the Multilayer Perceptron classifier and 96.25% for Random Forest for color channel R, 95.25% (Multilayer Perceptron) and 95% (Random Forest) for color channel L, 93% (Multilayer Perceptron) and 95% (Random Forest) for color channel X, 99.75% (Multilayer Perceptron) and 99.5% (Random Forest) for a set of textures selected from all color channels (R, G, B, L, a, b, X, Y, X). The highest average accuracies for other pairs of cultivars reached 98.25% for ‘Ożarowski’ vs. Sacher F1, 95.75% for ‘Pineapple’ vs. Sandoline F1, 97.5% for ‘Green Zebra’ vs. Sandoline F1, 97.25% for Sacher F1 vs. Sandoline F1 for models built based on textures selected from all color channels. The obtained results may be used in practice for the identification of cultivar of tomato seeds. The developed models allow to distinguish the tomato seed cultivars in an objective and fast way using digital image processing. The results confirmed the usefulness of texture parameters of the outer surface of tomato seeds for classification purposes. The discriminative models allow to obtain a very high probability and may be applied to authenticate and detect seed adulteration.
Similar content being viewed by others
Explore related subjects
Find the latest articles, discoveries, and news in related topics.Avoid common mistakes on your manuscript.
Introduction
The origin of tomato (Solanum lycopersicum L.) is not confirmed by archaeological evidence, but on the basis of DNA sequence analyses of plants currently found in Latin America, Peru and Ecuador are most often indicated as the place of origin. It is estimated that about 7000 years ago the selection of wild plants led to the development of the domestic tomato. For centuries, the yield has been the main criterion for breeding the tomato, which has led to a significant enlargement of the fruit with a simultaneous reduction in sugar and aroma content [1]. In 2019, the global cultivated area was 6.1 million hectares and total production was around 243.6 million tons [2], making tomato one of the world's main food crops. The common tomato produces fruits in a large variety of shapes, colors, and sizes. Tomato quality factors for fresh consumption are overall appearance, firmness and taste, whereas the quality of tomatoes for processing is determined by total solids content, color, pH and firmness [3]. For the consumers, color is an indicator of maturity level and in many cases, this feature of the fruit has a decisive influence on the preference for selection. The color of tomatoes fruits depends mainly on lycopene content. The second-most important compound affecting the color of the fruit is β-Carotene [4]. The nutritional value of tomatoes is mainly due to their nutrient content (carotenoids, polyphenols, ascorbic acid), minerals (Ca, Mg, Cu, Zn, K, Fe) and fiber [5]. In comparison to other vegetables, tomato fruits have intermediate levels of carotenoids; however, high dietary intake makes it a very important source. It has recently been shown that the content of lycopene and β-carotene in the fresh weight of tomatoes fruits is in the range 0.02–422 mg/100 g and 0.01–4.44 mg/100 g, respectively [6]. A similar situation applies to ascorbic acid content. Comparing to other vegetables, its concentration remains at an average level, but the large quantities consumed make tomatoes great contributors of this nutrient in diet [3]. Large differences in ascorbic acid levels have been reported among tomato cultivars and growing conditions; however, concentration in tomatoes fruits was estimated between 1 and 64 mg/100 g in fresh weight [6] The level of phenolic compounds in tomato fruit is influenced by a huge number of factors (variety, cultivation method, weather conditions, degree of ripeness) and therefore average content may not be representative [6, 7]. However, the content of polyphenolic compounds is not high, as it was reported, the fruits contain flavonoids in concentration ranges 2.57–4.37 mg/100 g [8] and phenolic acids (5-caffeoylquinic acid and caffeoylquinic acids derivatives) 10.5 mg/100 g fresh weight [5]. Epidemiological evidence indicates an association between the consumption of tomatoes and reduced cardiovascular risk. Lycopene administered at 200 mg/day has a significant effect on normalizing the blood lipid profile [9]. Tomato intake was found to have a reduction effect on LDL, total cholesterol, TG, and an increase in HDL levels [10].
Spectral and image analysis acquires using various methods and provides valuable information for classifying the physiological condition of seeds, their defects invisible to the eye, and for variety discrimination. Nowadays, non-destructive, rapid classification methods based on imaging, tomography and infrared spectroscopy (NIR) are under development for such use. For example, excellent results were obtained when the possibility of using near-infrared spectroscopy (reflection spectrum) to classify damaged and correct tomato seeds was investigated. The study showed that these discrimination models can be used to differentiate thermally damaged seeds. Total classification accuracy for the validation sample was 96.7% when five factors were selected for partial least squares discriminant analysis [11]. The potential of NIR spectroscopy for discrimination of tomato seed quality (viable and non-viable) using spectral analysis was evidenced. The ability to correctly identify the positive samples and to reject the negative samples of the model for prediction of viable and non-viable seeds were in both cases: 0.94 [12]. Rapid non-destructive grading of tomato seeds was also developed based on the hyperspectral technique. The area, circularity and average gray of seeds were analyzed to correlate with standard germination test performance. Image acquisition system equipped with line scanning spectrometer, gives a good result when 713 nm of wavelength was selected for prediction analysis. The accuracy of the calibration and validation data set was above 90.00% [13]. In another case, the physiological maturity of tomato seeds determined by X-ray image analysis proved to be an effective method for selecting high-quality seeds. The internal features of the seeds: embryo morphology and presence of free areas, (which represent the physiological potential of the seeds) were analyzed on the radiographic images [14]. Own research proposes the application of image textures for cultivar discrimination of tomato seeds. In the available literature, there is a lack of information on the presence of models based on textures extracted from the color channels R, G, B, L, a, b, X, Y, Z from digital color images acquired using a flatbed scanner for distinguishing of tomato seed cultivars. The performed experiments were intended to supplement this scarcity.
The objective of this study was to develop the discriminant models for distinguishing the tomato seeds based on texture parameters of the outer surface of seeds calculated from the images (scans) converted to individual color channels R, G, B, L, a, b, X, Y, Z.
Materials and methods
Materials
The tomatoes belonging to cultivars ‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’, Sacher F1 and Sandoline F1 were used in the experiments. The tomatoes were purchased from a local manufacturer. The seeds were manually prepared for the image acquisition. The individual tomato fruits were cut into quarters. Then, the seed chambers were emptied. The extracted seeds were covered with a protective tissue (mucilaginous gel) which was removed to obtain clean seeds. During the process of seed extraction, the seeds were rinsed in a sieve under tap water. In the next step, the mucilaginous gel was removed mechanically by sponge on absorption paper. Before scanning, the seeds were dried with paper towels.
Image analysis
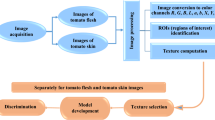
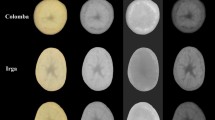
The tomato seeds images were obtained with the use of a flatbed scanner. The outer surfaces of seeds belonging to tomatoes ‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’, Sacher F1 and Sandoline F1 were scanned on the black background that facilitated the segmentation and ROI (region of interest) identification. For each cultivar, two scans were acquired. One scan included one-hundred seeds. Therefore, the images of two hundred seeds were obtained for each tomato cultivar. The images were characterized by the following parameters: 800 dpi resolution, TIFF format. After the image acquisition, the Mazda software (Łódź University of Technology, Institute of Electronics, Poland) [15] was applied for image processing. Before image analysis using the Mazda application, the images were converted to BMP format. Then, the conversion of tomato seed images to individual color channels R, G, B, L, a, b, X, Y, Z was carried out. The exemplary images of tomato seeds are presented in Fig. 1. The individual seeds were separated from the background and the region of interest (ROI) was overlaid on each seed. For one image from each color channel, almost 200 textures based on the run-length matrix, histogram, co-occurrence matrix, autoregressive model and gradient map [15] were computed for each ROI (one seed) and these features were used for the stage of attribute selection. Of the 200 features, the features with the highest discriminatory power were selected and used to build models to distinguish tomato seed cultivars. In this study, the texture parameters of tomato seeds were calculated from images based on spatial variation of pixel brightness intensities. Analysis of textures can provide numerical data about the structure of objects, which can determine the changes that are difficult to notice to the naked eye. The images of objects with the same color histograms and number of pixels can differ in textures if they have dissimilar color distributions [16,17,18]. The texture parameters were successfully applied to distinguish seed cultivars [19,20,21,22]. The proposed procedure of cultivar discrimination of tomato seeds is presented in Fig. 2.
Discriminant analysis
The discrimination of tomato seeds belonging to different cultivars was carried out with the use of the WEKA 3.9 application (Machine Learning Group, University of Waikato) [23]. The cultivars were discriminated in pairs: ‘Green Zebra’ vs. ‘Ożarowski’, ‘Green Zebra’ vs. ‘Pineapple’, ‘Green Zebra’ vs. Sacher F1, ‘Green Zebra’ vs. Sandoline F1, ‘Ożarowski’ vs. ‘Pineapple’, ‘Ożarowski’ vs. Sacher F1, ‘Ożarowski’ vs. Sandoline F1, ‘Pineapple’ vs. Sacher F1, ‘Pineapple’ vs. Sandoline F1, Sacher F1 vs. Sandoline F1. Additionally, the discrimination of all five tomato seed cultivars was performed and the discrimination for one cultivar versus other cultivars. The discriminative models were developed individually for the sets of selected textures. The textures were selected using the Best First with the CFS (Correlation-based Feature Selection) subset evaluator. In the case of pair comparison, 10 textures were selected for individual color channels and 30 for all color channels for each pair of tomato seed cultivars. This was the optimal number of features that provided high correctness of discrimination and a short analysis time. For the classification of all five cultivars, there were more selected textures, about 15 for each color channel and 35 for model built for a set including textures selected from all color channels. It meant that there may be a need to use more features to distinguish more cultivars from each other. For example, in the case of color channel R, the following textures were selected: RHMean, RHVariance, RHPerc01, RHPerc50, RHPerc99, RSGSkewness, RS5SH1DifVarnc, RS5SV1SumVarnc, RS4RHLngREmph, RS4RVGLevNonU, RS4RVLngREmph, RS4RZRLNonUni, RAArea, RATeta2, RASigma. For color channel X, the selected textures were: XHMean, XHVariance, XHPerc01, XHPerc10, XHPerc50, XHPerc99, XSGArea, XS5SV1SumVarnc, XS5SN1DifEntrp, XS5SZ3AngScMom, XS4RVGLevNonU, XS4RZRLNonUni, XAArea, XATeta2, XASigma. The discrimination was performed using different classifiers from the groups of Functions, Decision Trees, Lazy and Rules which were available using the Weka application. The tenfold cross-validation mode was applied for the discrimination [24]. In the case of each pair, the discriminant models were built separately for individual color channels R, G, B, L, a, b, X, Y, Z from color spaces RGB, Lab, XYZ, respectively, using different classifiers. The main criterion for the evaluation of the analysis performance and selection of classifiers was the highest average accuracy (%). The accuracies of classification (%) for individual tomato seed cultivars were also evaluated. The highest discrimination accuracies were determined in the case of the Multilayer Perceptron and Random Forest classifiers, as well as the color channels R, L and X when the models were built for sets of textures selected individually for each color channel. Therefore, the results for these discriminative classifiers and individual color channels are presented in this paper.
Random Forest is one of the classifier algorithms from a group of Decision Trees. The function of this classifier is to build random forests by bagging ensembles of randomized decision trees. The Multilayer Perceptron classifier is a neural network belonging to a group of Functions that uses backpropagation for training [24].
Additionally, the models for sets of textures selected from all color channels were developed. The presented results include the confusion matrices for the pairs of cultivars and all five cultivars of tomato seeds, the average accuracies for each pair of cultivars and all five cultivars, and the TP (True Positive) Rate, Precision, F-Measure, ROC (Receiver Operating Characteristic) Area and PRC (Precision–Recall) Area. The values of these metrics were computed using the Weka. However, these parameters may be calculated manually using the following equations [22]:
where TP is the True Positive; FP is the False Positive; FN is the False Negative.
The interpretation of the results was based on the average accuracy (%) of classification of the tomato seeds belonging to different cultivars, the accuracies of classification (%) for individual cultivars and the values of other performance metrics, such as TP Rate, Precision, F-Measure, ROC Area and PRC Area. The higher the accuracies and the values of other metrics, the better the classification efficiency.
Additionally, the ROC (Receiver Operating Characteristic) and PRC (Precision–Recall) curves for selected model and classifier providing high accuracy were presented for discrimination of five cultivars of tomato seeds to show distinguishing one cultivar from the four others as follows:
-
‘Green Zebra’ versus tomato seeds other than 'Green Zebra' (‘Ożarowski’, ‘Pineapple’, Sacher F1 and Sandoline F1),
-
‘Ożarowski’ versus tomato seeds other than ‘Ożarowski’ (‘Green Zebra’, ‘Pineapple’, Sacher F1 and Sandoline F1),
-
‘Pineapple’ versus tomato seeds other than ‘Pineapple’ (‘Green Zebra’, ‘Ożarowski’, Sacher F1 and Sandoline F1),
-
Sacher F1 versus tomato seeds other than Sacher F1 (‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’ and Sandoline F1),
-
Sandoline F1 versus tomato seeds other than Sandoline F1 (‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’ and Sacher F1).
Results and discussion
The cultivar discrimination of tomato seeds was performed for pairs of cultivars. In the case of each pair, the discriminant models were built separately for color channels R, L, X based on selected textures, and two discriminative classifiers (Multilayer Perceptron and Random Forest) were applied for classification. In the case of the model built based on textures selected from color channel R of images of tomato seeds belonging to cultivars ‘Green Zebra’ and ‘Ożarowski’, very satisfactory discrimination accuracies were obtained for both classifiers (Table 1). The Multilayer Perceptron produced an average accuracy of 97% and the Random Forest allowed for the classification of seeds with an average accuracy equal to 96.25%. In the case of the Multilayer Perceptron, the confusion matrices indicated that the seeds ‘Green Zebra’ were correctly classified in 96.5% and 3.5% of the seeds belonging to tomato ‘Green Zebra’ were incorrectly included in class ‘Ożarowski’. Whereas the seeds ‘Ożarowski’ were discriminated with the correctness of 97.5% and only 2.5% of seeds ‘Ożarowski’ were incorrectly classified as ‘Green Zebra’. Also, the values of other metrics of discrimination were high. The TP Rate reached 0.975 for ‘Ożarowski’, Precision—0.975 for ‘Green Zebra’, F-Measure—0.970 and ROC Area—0.988 for both ‘Green Zebra’ and ‘Ożarowski’, and PRC Area was up to 0.992 for ‘Green Zebra’. Slightly worse but still satisfactory discrimination accuracies were obtained in the case of tomato seeds belonging to cultivars ‘Pineapple’ and Sandoline F1 for both analyzed classifiers. In the case of Multilayer Perceptron, only 9% of seeds were incorrectly classified (average accuracy of 91%) regardless of cultivar. The Random Forest classifier allowed for the classification of seeds with an average accuracy equal to 95.25%, with very high values of ROC Area and PRC Area—0.985 for both ‘Pineapple’ and Sandoline F1. In order of classification accuracy, the next pair was ‘Green Zebra’ vs. Sandoline F1. For both cultivars, all discrimination metric values had almost no difference and were relatively high. Average accuracy reached 93.25% (Multilayer Perceptron) and 92.75% (Random Forest). In the case of both cultivars, regardless of the applied discriminative classifiers, TP Rate, ROC Area, PRC Area reached values: 0.935 (‘Green Zebra’, Multilayer Perceptron and Sandoline F1, Random Forest), 0.973 (both cultivars and both classifiers), and 0.973 (Sandoline F1, Multilayer Perceptron and Random Forest), respectively. For some pairs of cultivars, the average accuracies were between 80 and 90% and in these cases, the metric scores of the TP Rate, Precision, F-Measure, ROC Area, PRC Area for Multilayer Perceptron reached the following values: 0.835–0.940, 0.835–0.935, 0.835–0.906, 0.908–0.964, 0.877–0.969, respectively. In this group of cultivar pairs (average accuracy: 90% or less) when the Random Forest classifier was applied, slightly better performers were obtained. In this case, the average metric scores of the TP Rate, Precision, F-Measure, ROC Area, PRC Area reached the values: 0.874; 0.875; 0.874; 0.945; 0.943, respectively. In the case of the tomato seed discrimination of just one pair of cultivars (‘Pineapple’ vs. Sacher F1), the lowest accuracies (less than 80%) were obtained for both classifiers. In the case of ‘Pineapple’ (Multilayer Perceptron), 25.5% of seeds were incorrectly classified as Sacher F1, which resulted in low average accuracy (77.25%). The Random Forest classifier gave slightly better results with an obtained average accuracy of 79.5%.
In the next step of the analysis, the discriminant models were built for the sets of textures selected from color channel L (Table 2). For the pairs of cultivars of tomato seeds, the highest accuracies reaching 95.25% (Multilayer Perceptron) and 95% (Random Forest) were determined for ‘Green Zebra’ vs. ‘Ożarowski’. The values of TP Rate, Precision, F-Measure, ROC Area and PRC Area were also very high and reached 0.965 (‘Ożarowski’, Random Forest), 0.964 (‘Green Zebra’, Random Forest), 0.953 (‘Ożarowski’, Multilayer Perceptron), 0.991 (‘Green Zebra’, ‘Ożarowski’, Random Forest), 0.993 ‘Green Zebra’, Random Forest), respectively. Therefore, very high correctness greater than 90% were found for the following pairs of cultivars: ‘Pineapple’ vs. Sandoline F1 (94.5% for Multilayer Perceptron and 93.5% for Random Forest), ‘Ożarowski’ vs. Sandoline F1 (91% for Multilayer Perceptron and 91.75% for Random Forest), ‘Green Zebra’ vs. Sandoline F1 (90.75% for Multilayer Perceptron and 91.75% for Random Forest). The accuracy of up to 90.5% was also obtained for Green Zebra vs. Sacher F1 for the Random Forest classifier. The discrimination of tomato seeds of ‘Pineapple’ and Sacher F1 was characterized by the lowest results equal to 74.5% for Multilayer Perceptron and 79% for Random Forest. Also, other metrics of discriminant analysis were low equal even 0.740 (TP Rate), 0.743 (Precision), 0.744 (F-Measure), 0.820 (ROC Area), 0.810 (PRC Area). The accuracies of cultivar discrimination of the other pairs of the tomato seeds reached 87% (Random Forest) for ‘Green Zebra’ vs. ‘Pineapple’, 86% (Multilayer Perceptron) for ‘Ożarowski’ vs. ‘Pineapple’, 83.5% (Random Forest) for ‘Ożarowski’ vs. Sacher F1, 85.75% (Random Forest) for Sacher F1 vs. Sandoline F1.
In the case of discrimination of the pairs of tomato cultivars based on seed textures from the images converted to color channel X, the average accuracies were very high and reached 95% for ‘Green Zebra’ vs. ‘Ożarowski’ for the Random Forest classifier (Table 3). The seeds belonging to tomato ‘Ożarowski’ were correctly classified in 97% and for the seeds ‘Green Zebra’, the correctness was 93%. The TP Rate reached 0.970 (‘Ożarowski’), Precision was up to 0.969 (‘Green Zebra’), F-Measure—up to 0.951 (‘Ożarowski’), ROC Area—up to 0.987 (‘Ożarowski’, ‘Green Zebra’), PRC Area—up to 0.989 (‘Green Zebra’). Also, in the case of the Multilayer Perceptron, the average accuracy of 93% (93% for ‘Green Zebra’ and 93% for ‘Ożarowski’) was very satisfactory. The average accuracy of 92.75% was obtained in the case of the discrimination of seeds ‘Pineapple’ and Sandoline F1, for both Multilayer Perceptron and Random Forest. Additionally, the seeds belonging to tomatoes ‘Green Zebra’ vs. Sandoline F1 were distinguished with very high accuracies equal to 92% (Multilayer Perceptron) and 91.25% (Random Forest). The seeds of tomatoes ‘Green Zebra’ vs. Sacher F1 were correctly discriminated in 90.25% (Multilayer Perceptron) to 91.5% (Random Forest). Also, a pretty high accuracy of 91.75% was observed in the case of seeds ‘Ożarowski’ vs. Sandoline F1 for the Random Forest. Whereas the tomato seeds ‘Pineapple’ vs. Sacher F1 were correctly distinguished with the lowest accuracy of 76.75% (Multilayer Perceptron)—81.25% (Random Forest). The seeds belonging to other pairs of cultivars were correctly discriminated in 83.75–85.75% (‘Green Zebra’ vs. ‘Pineapple’), 82.75–85% (‘Ożarowski’ vs. ‘Pineapple’), 80.75–81.75% (‘Ożarowski’ vs. Sacher F1), 84.75–87.5% (Sacher F1 vs. Sandoline F1) for the Multilayer Perceptron and Random Forest classifiers (Table 3).
The increase in the correctness was obtained by combining the textures from all color channels R, G, B, L, a, b, X, Y, X in the discriminative models (Table 4). The tomato seeds belonging to ‘Green Zebra’ and ‘Ożarowski’ were correctly discriminated with an average accuracy of 99.75% to 99.5% for the Multilayer Perceptron and Random Forest, respectively. For this pair of cultivars, the accuracy of 100% was observed for seeds ‘Ożarowski’ and the seeds ‘Green Zebra’ were correctly discriminated in 99.5% of the cases. In the case of other pairs, very high correctness was also achieved. The average accuracies of seed discrimination reached the following values: ‘Green Zebra’ vs. ‘Pineapple’—92% (Multilayer Perceptron), ‘Green Zebra’ vs. Sacher F1—91.25% (Random Forest), ‘Green Zebra’ vs. Sandoline F1—97.5% (Multilayer Perceptron, Random Forest), ‘Ożarowski’ vs. ‘Pineapple’—95% (Random Forest), ‘Ożarowski’ vs. Sacher F1—98.25% (Multilayer Perceptron, Random Forest), ‘Ożarowski’ vs. Sandoline F1—95.25% (Random Forest), ‘Pineapple’ vs. Sacher F1—84.5% (Random Forest), ‘Pineapple’ vs. c Sandoline F1—95.75% (Random Forest), Sacher F1 vs. Sandoline F1 97.25% (Random Forest).
The average accuracies of discrimination of all five cultivars of tomato seeds were slightly lower than for pair comparisons. The tomato seeds ‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’, Sacher F1 and Sandoline F1 were correctly discriminated with average accuracies reaching 83.6% (Random Forest) for model developed using a set of textures selected from all color channels R, G, B, L, a, b, X, Y, Z and 73.7% (Random Forest) for color channel R for analysis performed for individual color channels. In the case of individual cultivars, the tomato seeds ‘Ożarowski’ were discriminated with the highest accuracy of up to 93.5% (Random Forest classifier, textures selected from a set of all color channels R, G, B, L, a, b, X, Y, Z) (Table 5). The other performance metrics for the discrimination of five cultivars of tomato seeds reached 0.935 for TP Rate, 0.912 for F-Measure, 0.990 for ROC Area, 0.960 for PRC Area for ‘Ożarowski’ and 0.894 for Precision for Sandoline F1 in the case of a set of textures selected from all color channels R, G, B, L, a, b, X, Y, Z and the Random Forest classifier (Table 6).
The ROC (Receiver Operating Characteristic) and PRC (Precision–Recall) curves for each tomato seed cultivar versus other cultivars for the model developed based on a set of textures selected from all color channels (R, G, B, L, a, b, X, Y, X) and the Multilayer Perceptron classifier that provided high accuracies are presented in Figs. 3, 4, 5, 6 and 7. The ROC and PRC curves for the discrimination of the tomato seeds ‘Green Zebra’ versus all other tomato seed cultivars (‘Ożarowski’, ‘Pineapple’, Sacher F1 and Sandoline F1) are presented in Fig. 3, the seeds ‘Ożarowski’ versus other cultivars (‘Green Zebra’, ‘Pineapple’, Sacher F1 and Sandoline F1)—in Fig. 4, the tomato seeds ‘Pineapple’ and tomato seeds other than ‘Pineapple’ (‘Green Zebra’, ‘Ożarowski’, Sacher F1 and Sandoline F1)—in Fig. 5, the seeds Sacher F1 versus other tomato seeds (‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’ and Sandoline F1) in—Fig. 6, the tomato seeds Sandoline F1 versus other cultivars (‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’ and Sacher F1)—in Fig. 7. The area under ROC was the highest for tomato seeds ‘Ożarowski’ and tomato seeds other than ‘Ożarowski’ (‘Green Zebra’, ‘Pineapple’, Sacher F1 and Sandoline F1) and was equal to 0.976 for both curves (Fig. 4a, b). The values of PRC Area reached 0.927 for tomato seeds ‘Ożarowski’ (Fig. 4c) and 0.993 for tomato seeds other than ‘Ożarowski’ (Fig. 4d).
The ROC (Receiver Operating Characteristic) and PRC (Precision–Recall) curves for the tomato seeds for the model developed based on a set of textures selected from all color channels (R, G, B, L, a, b, X, Y, X) and the Multilayer Perceptron classifier, a ROC curve, tomato seeds ‘Green Zebra’, area under ROC = 0.972, b ROC curve, tomato seeds other than ‘Green Zebra’ (‘Ożarowski’, ‘Pineapple’, Sacher F1 and Sandoline F1), area under ROC = 0.972, c PRC curve, tomato seeds ‘Green Zebra’, PRC Area = 0.877, d PRC curve, tomato seeds other than ‘Green Zebra’, PRC Area = 0.993
The ROC (Receiver Operating Characteristic) and PRC (Precision–Recall) curves for the tomato seeds for the model developed based on a set of textures selected from all color channels (R, G, B, L, a, b, X, Y, X) and the Multilayer Perceptron classifier, a ROC curve, tomato seeds ‘Ożarowski’, area under ROC = 0.976, b ROC curve, tomato seeds other than ‘Ożarowski’ (‘Green Zebra’, ‘Pineapple’, Sacher F1 and Sandoline F1), area under ROC = 0.976, c PRC curve, tomato seeds ‘Ożarowski’, PRC Area = 0.927, d PRC curve, tomato seeds other than ‘Ożarowski’, PRC Area = 0.993
The ROC (Receiver Operating Characteristic) and PRC (Precision–Recall) curves for the tomato seeds for the model developed based on a set of textures selected from all color channels (R, G, B, L, a, b, X, Y, X) and the Multilayer Perceptron classifier, a ROC curve, tomato seeds ‘Pineapple’, area under ROC = 0.874, b ROC curve, tomato seeds other than ‘Pineapple’ (‘Green Zebra’, ‘Ożarowski’, Sacher F1 and Sandoline F1), area under ROC = 0.874, c PRC curve, tomato seeds ‘Pineapple’, PRC Area = 0.696, d PRC curve, tomato seeds other than ‘Pineapple’, PRC Area = 0.954
The ROC (Receiver Operating Characteristic) and PRC (Precision–Recall) curves for the tomato seeds for the model developed based on a set of textures selected from all color channels (R, G, B, L, a, b, X, Y, X) and the Multilayer Perceptron classifier, a ROC curve, tomato seeds Sacher F1, area under ROC = 0.941, b ROC curve, tomato seeds other than Sacher F1 (‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’ and Sandoline F1), area under ROC = 0.941, c PRC curve, tomato seeds Sacher F1, PRC Area = 0.794, d PRC curve, tomato seeds other than Sacher F1, PRC Area = 0.983
The ROC (Receiver Operating Characteristic) and PRC (Precision–Recall) curves for the tomato seeds for the model developed based on a set of textures selected from all color channels (R, G, B, L, a, b, X, Y, X) and the Multilayer Perceptron classifier, a ROC curve, tomato seeds Sandoline F1, area under ROC = 0.965, b ROC curve, tomato seeds other than Sandoline F1 (‘Green Zebra’, ‘Ożarowski’, ‘Pineapple’ and Sacher F1), area under ROC = 0.965, c PRC curve, tomato seeds Sandoline F1, PRC Area = 0.892, d PRC curve, tomato seeds other than Sandoline F1, PRC Area = 0.987
Computer vision systems can be of great practical importance for the cultivar classification. Correct cultivar identification is needed to authenticate and avoid adulteration and mixing cultivars with different properties and applications [25]. Computer vision systems can ensure objective, accurate and reproducible quality evaluation [26, 27]. The application of image processing can provide distributors, producers and consumers important information about both cultivar and quality of seeds as well as identification of aberrant seeds [28]. The seed classification based on images can be important for crops, both fruit and vegetables, disease recognition, or for archaeobotanical reasons related to obtaining specific feature information [29]. Image analysis is non-destructive and easier than other techniques used for distinguishing tomato cultivars reported by the available literature, e.g., based on genetics [30]. Besides, image analysis of seeds may be more advantageous compared to manual analysis due to the speeding up of the process, automaticity of classification using image pixel values, reduction of distortions caused by natural light [31]. Image analysis and machine learning may replace labor-intensive and time-consuming human visual procedures and can be used by seed laboratories or in the nursery industry for inspections of tomato seeds and evaluation of their germination rate [32]. The non-destructive cultivar discrimination of tomato seeds can be also useful for registration programs, protection of plant cultivars and management of plant genetic resources [33]. Own research proved the usefulness of images obtained with the use of a flatbed scanner for cultivar discrimination of tomato seeds based on selected texture parameters extracted from color channels R, G, B, L, a, b, X, Y, Z. Selection of textures allowed to build innovative discriminative models that provided high correctness. The developed non-destructive, objective, fast and inexpensive procedure can be of great practical importance for distinguishing tomato seeds.
Conclusion
The obtained results indicated that the tomato seeds belonging to different cultivars can be discriminated with a very high probability using the selected features calculated from the images. Therefore, the usefulness of textures of the outer surface for seed discrimination was confirmed. The models built based on sets of combined textures selected from all color channels proved to be more useful for tomato seeds discrimination than the models built separately for each color channel. The average accuracy reached 99.75% for distinguishing seeds ‘Green Zebra’ and ‘Ożarowski’ and in this case, the seeds ‘Ożarowski’ were correctly discriminated in 100% and the seeds ‘Green Zebra’—in 99.5%. These results are very satisfactory. Due to this, the image analysis can be applied to confirm the authenticity of the seed cultivar and avoid the adulteration that may be useful in various industries, e.g., for tomato seed processing and tomato cultivation.
References
Klee HJ, Resende MFR (2020) Plant domestication: reconstructing the route to modern tomatoes. Curr Biol 30:359–361
FAOSTAT (2021) Tomato production in 2019. UN Food and Agriculture Organization, Corporate Statistical Database. FAOSTAT, Rome
Stevens MA (1986) Inheritance of tomato fruit quality components. Plant Breed Rev 4:273–311
Arias R, Lee TC, Logendra L, Janes H (2000) Correlation of lycopene measured by HPLC with the L*, a*, b* color readings of a hydroponic tomato and the relationship of maturity with color and lycopene content. J Agric Food Chem 48:1697–1702
Erba D, Casiraghi MC, Ribas-Agustí A, Cáceres R, Marfà O, Castellari M (2013) Nutritional value of tomatoes (Solanum lycopersicum L.) grown in greenhouse by different agronomic techniques. J Food Comp Anal 31:245–251
Vélez-Terreros PY, Romero-Estévez D, Yánez-Jácome GS, Simbaña-Farinango K, Navarrete H (2021) Comparison of major nutrients and minerals between organic and conventional tomatoes. A review. J Food Compos Anal 100:103922
Schindler M, Solar S, Sontag G (2005) Phenolic compounds in tomatoes. Natural variations and effect of gamma-irradiation. Eur Food Res Technol 221:439–445
Oliveira AB, Moura CFH, Gomes-Filho E, Marco CA, Urban L, Miranda MRA (2013) The impact of organic farming on quality of tomatoes is associated to increased oxidative stress during fruit development. PLoS ONE 8:e56354
Ratto F, Franchini F, Musicco M, Caruso G, Di Santo SG (2021) A narrative review on the potential of tomato and lycopene for the prevention of Alzheimer’s disease and other dementias. Crit Rev Food Sci Nutr. https://doi.org/10.1080/10408398.2021.1880363
Wolak T, Sharoni Y, Levy J, Linnewiel-Hermoni K, Stepensky D, Paran E (2019) Effect of tomato nutrient complex on blood pressure: a double blind, randomized dose–response study. Nutrients 11(5):950
Peng Y, Zhao F, Bai J, Zheng X, Wang W, Sun Q (2018) Detection and classification of tomato seed vitality based on image processing. Nongye Jixie Xuebao/Trans Chin Soc Agric Mach 49(2):327–333
Shrestha S, Deleuran LC, Gislum R (2017) Separation of viable and non-viable tomato (Solanum lycopersicum L.) seeds using single seed near-infrared spectroscopy. Comput Electron Agric 142:348–355
Peng Y, Zhao F, Li L, Xing Y, Fang X (2018) Discrimination of heat-damaged tomato seeds based on near infrared spectroscopy and PCA-SVM method. Nongye Gongcheng Xuebao/Trans Chin Soc Agric Eng 34(5):159–165
Borges SRDS, Silva PPD, Araújo FS, Souza FFDJ, Nascimento WM (2019) Tomato seed image analysis during the maturation. J Seed Sci 41(1):22–31
Szczypiński PM, Strzelecki M, Materka A, Klepaczko A (2009) MaZda—a software package for image texture analysis. Comput Methods Programs Biomed 94(1):66–76
Fernández L, Castillero C, Aguilera JM (2005) An application of image analysis to dehydration of apple discs. J Food Eng 67:185–193
Strzelecki M, Szczypiński P, Materka A, Klepaczko A (2013) A software tool for automatic classification and segmentation of 2D/3D medical images. Nucl Instrum Methods Phys Res 702:137–140
Armi L, Fekri-Ershad S (2019) Texture image analysis and texture classification methods—a review. Int Online J Image Process Pattern Recogn 2(1):1–29
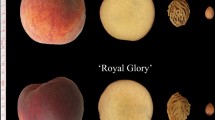
Ropelewska E, Rutkowski KP (2021) Differentiation of peach cultivars by image analysis based on the skin, flesh, stone and seed textures. Eur Food Res Technol 247:2371–2377
Ropelewska E (2020) The use of seed texture features for discriminating different cultivars of stored apples. J Stored Prod Res 88:101668
Pourreza A, Pourreza H, Abbaspour-Fard MH, Sadrnia H (2012) Identification of nine Iranian wheat seed varieties by textural analysis with image processing. Comput Electron Agric 83:102–108
Ropelewska E, Szwejda-Grzybowska J (2021) A comparative analysis of the discrimination of pepper (Capsicum annuum L.) based on the cross-section and seed textures determined using image processing. J Food Process Eng 44:e13694
Bouckaert RR, Frank E, Hall M, Kirkby R, Reutemann P, Seewald A, Scuse D (2016) WEKA manual for version 3-9-1. University of Waikato, Hamilton
Witten IH, Frank E (2005) Data mining: practical machine learning tools and techniques, 525, 2nd edn. Elsevier, San Francisco
Pourdarbani R, Sabzi S, Kalantari D, Hernández-Hernández JL, Arribas JI (2020) A computer vision system based on majority-voting ensemble neural network for the automatic classification of three chickpea varieties. Foods 9:113
Abdullah MZ (2016) Image acquisition systems. In: Sun D-W (ed) Computer vision technology for food quality evaluation, 2nd edn. Academic Press/Elsevier Inc., San Diego, pp 3–43
Priyadharshini K, Akila R (2016) A survey on computer vision Technology for food quality evaluation. Int J Innov Res Comput Commun Eng 4(8):14860–14865
Kapadia VN, Sasidharan N, Kalyanrao P (2017) Seed image analysis and its application in seed science research. Adv Biotechnol Microbiol 7(2):555709
Loddo A, Di Ruberto C (2021) On the efficacy of handcrafted and deep features for seed image classification. J Imaging 7:171
Wang XF, Knoblauch R, Leist N (2000) Varietal discrimination of tomato (Lycopersicon esculentum L.) by ultrathin-layer isoelectric focusing of seed protein. Seed Sci Technol 28(2):521–526
Loddo A, Loddo M, Di Ruberto C (2021) A novel deep learning based approach for seed image classification and retrieval. Comput Electron Agric 187:106269
Škrubej U, Rozman Č, Stajnko D (2015) Assessment of germination rate of the tomato seeds using image processing and machine learning. Europ J Hort Sci 80(2):68–75
Shrestha S, Deleuran LCh, Gislum R (2016) Classification of different tomato seed cultivars by multispectral visible-near infrared spectroscopy and chemometrics. J Spectral Imaging 5:1–8
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts to declare.
Compliance with ethics requirements
This article does not contain any studies with human or animal subject.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ropelewska, E., Piecko, J. Discrimination of tomato seeds belonging to different cultivars using machine learning. Eur Food Res Technol 248, 685–705 (2022). https://doi.org/10.1007/s00217-021-03920-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-021-03920-w