Abstract
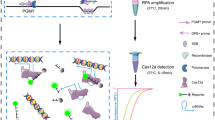
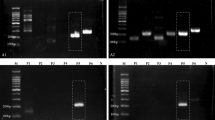
Cervical cancer is the second most common cancer in the world’s woman population with a high incidence in developing countries where diagnostic conditions for the cancer are poor. The main culprit causing the cancer is the human papillomavirus (HPV). HPV is divided into three major groups, i.e., high-risk (HR) group, probable high-risk (pHR) group, and low-risk (LR) group according to their potential of causing cervical cancer. Therefore, developing a sensitive, reliable, and cost-effective point-of-care diagnostic method for the virus genotypes in developing countries even worldwide is of high importance for the cancer prevention and control strategies. Here we present a combined method of isothermal recombinase polymerase amplification (RPA), lateral flow dipstick (LFD), and reverse dot blot (RDB), in quick point-of-care identification of HPV genotypes. The combined method is highly specific to HPV when the conserved L1 genes are used as targeted genes for amplification. The method can be used in identification of HPV genotypes at point-of-care within 1 h with a sensitivity of low to 100 fg of the virus genomic DNA. We have demonstrated that it is an excellent diagnostic point-of-care assay in monitoring the disease without time-consuming and expensive procedures and devices.




Similar content being viewed by others
References
Bosch FX, Lorincz A, Muñoz N, Meijer CJ, Shah KV. The causal relationship between human papillomavirus and cervical cancer. J Clin Pathol. 2002;55(4):245–65.
Muñoz N. Human papillomavirus and cancer: the epidemiological evidence. J Clin Virol. 2000;19(1–2):1–5.
Peng J, Gao L, Guo J, Wang T, Wang L, Yao Q, et al. Type-specific detection of 30 oncogenic human papillomaviruses by genotyping both E6 and L1 genes. J Clin Microbiol. 2013;51(2):402–8.
Urquiza M, Guevara T, Espejo F, Bravo MM, Rivera Z, Patarroyo ME. Two L1-peptides are excellent tools for serological detection of HPV-associated cervical carcinoma lesions. Biochem Biophys Res Commun. 2005;332(1):224–32.
Frías Isaac AM, Avelino Karen YPS, Silva Rafael R, Andrade César AS, Oliveira Maria DL. Trends in biosensors for HPV identification and diagnosis. J Sensors. 2015;2015(5):1–16.
Han J, Swan DC, Smith SJ, Lum SH, Sefers SE, Unger ER, et al. Simultaneous amplification and identification of 25 human papillomavirus types with Templex technology. J Clin Microbiol. 2006;44(11):4157–62.
Jimenez AMJ, Rodrigo MAM, Milosavljevic V, Krizkova S, Kopel P, Heger Z, et al. Gold nanoparticles-modified nanomaghemite and quantum dots-based hybridization assay for detection of HPV. Sens Actuators B Chem. 2017;240:503–10.
Jampasa S, Siangproh W, Laocharoensuk R, Yanatatsaneejit P, Vilaivan T, Chailapakul O. A new DNA sensor design for the simultaneous detection of HPV type 16 and 18 DNA. Sens Actuators B Chem. 2018;265:514–21.
Jiang HL, Zhu HH, Zhou LF, Chen F, Chen Z. Genotyping of human papillomavirus in cervical lesions by L1 consensus PCR and the Luminex xMAP system. J Med Microbiol. 2006;55(6):715–20.
Lee HP, Cho W, Bae JM, Shin JY, Shin SK, Hwang SY, et al. Comparison of the clinical performance of restriction fragment mass polymorphism (RFMP) and Roche linear array HPV test assays for HPV detection and genotyping. J Clin Virol. 2013;57(2):130–5.
Satoh T, Matsumoto K, Fujii T, Sato O, Gemma N, Onuki M, et al. Rapid genotyping of carcinogenic human papillomavirus by loop-mediated isothermal amplification using a new automated DNA test (Clinichip HPV™). J Virol Methods. 2013;188(1–2):83–93.
Lin J, Ma B, Fang J, Wang Y, He H, Lin W, et al. Colorimetric detection of 23 human papillomavirus genotypes by loop-mediated isothermal amplification. Clin Lab. 2017;63(3):495–505.
Hwang SH, Kim DE, Im JH, Kang SJ, Lee DH, Son SJ. Rapid visual identification of PCR amplified nucleic acids by centrifugal gel separation potential use for molecular point-of-care tests. Biosens Bioelectron. 2016;79:829–34.
Ahmed A, van der Linden H, Hartskeerl RA. Development of a recombinase polymerase amplification assay for the detection of pathogenic Leptospira. Int J Environ Res Public Health. 2014;11(5):4953–64.
Zaghloul H, El-shahat M. Recombinase polymerase amplification as a promising tool in hepatitis C virus diagnosis. World J Hepatol. 2014;6(12):916–22.
Ma B, Fang J, Wang Y, He H, Dai M, Lin W, et al. Isothermal method of a recombinase polymerase amplification assay for the detection of most common high-risk human papillomavirus type 16 and type 18 DNA. Clin Lab. 2017;63(1):27–38.
Boyle DS, McNerney R, Teng Low H, Leader BT, Pérez-Osorio AC, Meyer JC, et al. Rapid detection of Mycobacterium tuberculosis by recombinase polymerase amplification. PLoS One. 2014;9(8):e103091.
Niemz A, Boyle DS. Nucleic acid testing for tuberculosis at the point-of-care in high-burden countries. Expert Rev Mol Diagn. 2012;12(7):687–701.
Tian T, Bi Y, Xu X, Zhu Z, Yang C. Integrated paper-based microfluidic devices for point-of-care testing. Anal Methods. 2018;10(29):3567–81.
Piepenburg O, Williams CH, Stemple DL, Armes NA. DNA detection using recombination proteins. PLoS One. 2006;4(7):e204.
Abd El Wahed A, Patel P, Faye O, Thaloengsok S, Heidenreich D, Matangkasombut P, et al. Recombinase polymerase amplification assay for rapid diagnostics of dengue infection. Plos One. 2015;10(6):e0129682.
Clancy E, Higgins O, Forrest MS, Boo TW, Cormican M, Barry T, et al. Development of a rapid recombinase polymerase amplification assay for the detection of Streptococcus pneumoniae in whole blood. BMC Infect Dis. 2015;15:481.
Yang Y, Qin X, Wang G, Zhang Y, Shang Y, Zhang Z. Development of a fluorescent probe-based recombinase polymerase amplification assay for rapid detection of Orf virus. Virol J. 2015;12:206.
Choi G, Jung JH, Park BH, Oh SJ, Seo JH, Choi JS, et al. A centrifugal direct recombinase polymerase amplification (direct-RPA) microdevice for multiplex and real-time identification of food poisoning bacteria. Lab Chip. 2016;16(12):2309–16.
Kersting S, Rausch V, Bier FF, von Nickisch-Rosenegk M. Rapid detection of Plasmodium falciparum with isothermal recombinase polymerase amplification and lateral flow analysis. Malar J. 2014;13:99.
Rosser A, Rollinson D, Forrest M, Webster BL. Isothermal recombinase polymerase amplification (RPA) of Schistosoma haematobium DNA and oligochromatographic lateral flow detection. Parasit Vectors. 2015;8:446.
Del Río JS, Lobato IM, Mayboroda O, Katakis I, O'Sullivan CK. Enhanced solid-phase recombinase polymerase amplification and electrochemical detection. Anal Bioanal Chem. 2017;409(12):3261–9.
Nybond S, Réu P, Rhedin S, Svedberg G, Alfvén T, Gantelius J, et al. Adenoviral detection by recombinase polymerase amplification and vertical flow paper microarray. Anal Bioanal Chem. 2018. https://doi.org/10.1007/s00216-018-1503-y.
Chen Y, Cheng N, Xu Y, Huang K, Luo Y, Xu W. Point-of-care and visual detection of P. aeruginosa and its toxin genes by multiple LAMP and lateral flow nucleic acid biosensor. Biosens Bioelectron. 2016;81:317–23.
Toubanaki DK, Margaroni M, Karagouni E. Nanoparticle-based lateral flow biosensor for visual detection of fish nervous necrosis virus amplification products. Mol Cell Probes. 2015;29(3):158–66.
Santos VO, Pelegrini PB, Mulinari F, Lacerda AF, Moura RS, Cardoso LPV, et al. Development and validation of a novel lateral flow immunoassay device for detection of aflatoxins in soy-based foods. Anal Methods. 2017;9(18):2715–22.
Geraets DT, van Baars R, Alonso I, Ordi J, Torné A, Melchers WJ, et al. Clinical evaluation of high-risk HPV detection on self-samples using the indicating FTA-elute solid-carrier cartridge. J Clin Virol. 2013;57(2):125–9.
Acknowledgments
We thank Wenzhou People’s Hospital and Mei Zhong Medical Laboratory for providing the cervical scrape samples.
Funding
The work was supported by the National Key Research and Development Program of China (2017YFF0210200, 2018YFF0215205) and the Public Projects of Zhejiang Province (LGC19C200006).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The ethical committee of Wenzhou People’s Hospital approved the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 150 kb)
Rights and permissions
About this article
Cite this article
Ma, B., Fang, J., Lin, W. et al. A simple and efficient method for potential point-of-care diagnosis of human papillomavirus genotypes: combination of isothermal recombinase polymerase amplification with lateral flow dipstick and reverse dot blot. Anal Bioanal Chem 411, 7451–7460 (2019). https://doi.org/10.1007/s00216-019-02113-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-02113-5