Abstract
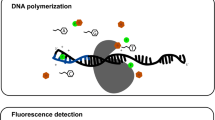
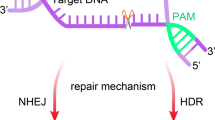
This study develops a new method for detecting target DNA based on Cas9 nuclease, which was named as CARP, representing CRISPR- or Cas9/sgRNAs-associated reverse PCR. This technique detects target DNA in three steps: (1) cleaving the detected DNA sample with Cas9 in complex with a pair of sgRNAs specific to target DNA; (2) ligating the cleaved DNA with DNA ligase; (3) amplifying target DNA with PCR. In the ligation step, the Cas9-cut target DNA was ligated into intramolecular circular or intermolecular concatenated linear DNA. In the PCR step, the ligated DNA was amplified with a pair of reverse primers. The technique was verified by detecting HPV16 and HPV18 L1 genes in nine different human papillomavirus (HPV) subtypes. The technique also detected the L1 and E6–E7 genes of two high-risk HPVs, HPV16 and HPV18, in the genomic DNA of two HPV-positive cervical carcinoma cells (HeLa and SiHa), in which no L1 and E6–E7 genes were detected in the HPV-negative cervical carcinoma cell, C-33a. By performing these proof-of-concept experiments, this study provides a new CRISPR-based DNA detection and typing method. Especially, the CARP method developed by this study is ready for the clinical HPV detection, which was supported by the final clinical sample detection.

CRISPR-associated reverse PCR (CARP) can be used to detect and type target DNA in a simple three-step procedure, cutting, ligation, and amplification.







Similar content being viewed by others
References
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol. 1987;169:5429–33.
Jansen R, Embden J, Gaastra W, Schouls L. Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol. 2002;43:1565–75.
Makarova KS, Aravind L, Wolf YI, Koonin EV. Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR-Cas systems. Biol Direct. 2011;6:38.
Makarova KS, Haft DH, Barrangou R, Brouns SJ, Charpentier E, Horvath P, et al. Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol. 2011;9:467.
Doudna JA, Charpentier E. The new frontier of genome engineering with CRISPR-Cas9. Science. 2014;346:1258096.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337:816–21.
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308.
Wang T, Wei JJ, Sabatini DM, Lander ES. Genetic screens in human cells using the CRISPR-Cas9 system. Science. 2014;343:80–4.
Sander JD, Joung JK. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol. 2014;32:347–55.
Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell. 2014;157:1262–78.
Chavez A, Tuttle M, Pruitt BW, Ewen-Campen B, Chari R, Ter-Ovanesyan D, et al. Comparison of Cas9 activators in multiple species. Nat Methods. 2016;13:563–7.
Liu W, Yu H, Zhou X, Xing D. In vitro evaluation of CRISPR/Cas9 function by an Electrochemiluminescent assay. Anal Chem. 2016;88:8369–74.
Pardee K, Green AA, Takahashi MK, Braff D, Lambert G, Lee JW, et al. Rapid, low-cost detection of Zika virus using programmable biomolecular components. Cell. 2016;165:1255–66.
Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, Joung J, et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science. 2017;356:438–42.
Pinheiro LB, Coleman VA, Hindson CM, Herrmann J, Hindson BJ, Bhat S, et al. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal Chem. 2012;84:1003–11.
Hindson BJ, Ness KD, Masquelier DA, Belgrader P, Heredia NJ, Makarewicz AJ, et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem. 2011;83:8604–10.
Hindson CM, Chevillet JR, Briggs HA, Gallichotte EN, Ruf IK, Hindson BJ, et al. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods. 2013;10:1003–5.
Boshart M, Gissmann L, Ikenberg H, Kleinheinz A, Scheurlen W, zur Hausen H. A new type of papillomavirus DNA, its presence in genital cancer biopsies and in cell lines derived from cervical cancer. EMBO J. 1984;3:1151–7.
Xu F, Cao M, Shi Q, Chen H, Wang Y, Li X. Integration of the full-length HPV16 genome in cervical cancer and Caski and Siha cell lines and the possible ways of HPV integration. Virus Genes. 2015;50:210–20.
Kilic G, Cardillo M, Ozdemirli M, Arun B. Human papillomavirus 18 oncoproteins E6 and E7 enhance irradiation- and chemotherapeutic agent-induced apoptosis in p53 and Rb mutated cervical cancer cell lines. Eur J Gynaecol Oncol. 1999;20:167–71.
zur Hausen H. Human papillomaviruses in the pathogenesis of anogenital cancer. Virology. 1991;184:9–13.
zur Hausen H. Papillomaviruses in the causation of human cancers—a brief historical account. Virology. 2009;384:260–5.
Walboomers JM, Jacobs MV, Manos MM, Bosch FX, Kummer JA, Shah KV, et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. J Pathol. 1999;189:12–9.
Faridi R, Zahra A, Khan K, Idrees M. Oncogenic potential of human papillomavirus (HPV) and its relation with cervical cancer. Virol J. 2011;8:269.
Farhat S, Scott M, Ma Y, Moscicki A-B. Development of a novel liquid bead array human papillomavirus genotyping assay (PGMY-LX) and comparison with linear array for continuity in longitudinal cohort studies. J Clin Microbiol. 2015:JCM. 03100–14.
Tommasino M, editor. The human papillomavirus family and its role in carcinogenesis. Semin Cancer Biol; 2014: Elsevier.
Han LL, Maimaitiming T, Husaiyin S, Wang L, Wusainahong K, Ma CH, et al. Comparative study of HPV16 integration in cervical lesions between ethnicities with high and low rates of infection with high-risk HPV and the correlation between integration rate and cervical neoplasia. Experimental and Therapeutic Medicine. 2015;10:2169–74.
Rao A, Young S, Erlich H, Boyle S, Krevolin M, Sun R, et al. Development and characterization of the cobas human papillomavirus test. J Clin Microbiol. 2013;51:1478–84.
Abreu AL, Souza RP, Gimenes F, Consolaro ME. A review of methods for detect human papillomavirus infection. Virol J. 2012;9:262.
Doorbar J, Quint W, Banks L, Bravo IG, Stoler M, Broker TR, et al. The biology and life-cycle of human papillomaviruses. Vaccine. 2012;30:F55–70.
Tao P, Zheng W, Wang Y, Bian M-L. Sensitive HPV genotyping based on the flow-through hybridization and gene chip. BioMed Research International. 2012;2012.
Syrjänen S, Lodi G, von Bültzingslöwen I, Aliko A, Arduino P, Campisi G, et al. Human papillomaviruses in oral carcinoma and oral potentially malignant disorders: a systematic review. Oral Dis. 2011;17:58–72.
Karlsen F, Kalantari M, Jenkins A, Pettersen E, Kristensen G, Holm R, et al. Use of multiple PCR primer sets for optimal detection of human papillomavirus. J Clin Microbiol. 1996;34:2095–100.
Depuydt CE, Boulet GA, Horvath CA, Benoy IH, Vereecken AJ, Bogers JJ. Comparison of MY09/11 consensus PCR and type-specific PCRs in the detection of oncogenic HPV types. J Cell Mol Med. 2007;11:881–91.
Woodman CB, Collins SI, Young LS. The natural history of cervical HPV infection: unresolved issues. Nat Rev Cancer. 2007;7:11–22.
Pattanayak V, Lin S, Guilinger JP, Ma E, Doudna JA, Liu DR. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat Biotechnol. 2013;31:839–43.
Mali P, Aach J, Stranges PB, Esvelt KM, Moosburner M, Kosuri S, et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat Biotechnol. 2013;31:833–8.
Wu X, Scott DA, Kriz AJ, Chiu AC, Hsu PD, Dadon DB, et al. Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nat Biotechnol. 2014;32:670–6.
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol. 2013;31:827–32.
Ran FA, Hsu PD, Lin C-Y, Gootenberg JS, Konermann S, Trevino AE, et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell. 2013;154:1380–9.
Montague TG, Cruz JM, Gagnon JA, Church GM, Valen E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 2014;42:W401–W7.
Acknowledgments
This work was supported by the grants from the National Natural Science Foundation of China (grant 61571119).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Informed consent
All procedures used in the present study were performed according to the Declaration of Helsinki. The Ethics Committee of Jinling Hospital (Nanjing, China) approved the present study. Written informed consent was obtained from all participants.
Rights and permissions
About this article
Cite this article
Zhang, B., Wang, Q., Xu, X. et al. Detection of target DNA with a novel Cas9/sgRNAs-associated reverse PCR (CARP) technique. Anal Bioanal Chem 410, 2889–2900 (2018). https://doi.org/10.1007/s00216-018-0873-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-018-0873-5