Abstract
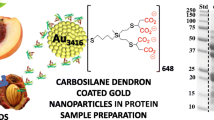
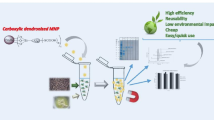
Protein sample preparation is a critical and an unsustainable step since it involves the use of tedious methods that usually require high amount of solvents. The development of new materials offers additional opportunities in protein sample preparation. This work explores, for the first time, the potential application of carboxylate-terminated carbosilane dendrimers to the purification/enrichment of proteins. Studies on dendrimer binding to proteins, based on protein fluorescence intensity and emission wavelengths measurements, demonstrated the interaction between carboxylate-terminated carbosilane dendrimers and proteins at all tested pH levels. Interactions were greatly affected by the protein itself, pH, and dendrimer concentration and generation. Especially interesting was the interaction at acidic pH since it resulted in a significant protein precipitation. Dendrimer-protein interactions were modeled observing stable complexes for all proteins. Carboxylate-terminated carbosilane dendrimers at acidic pH were successfully used in the purification/enrichment of proteins extracted from a complex sample.

Images showing the growing turbidity of solutions containing a mixture of proteins (lysozyme, myoglobin, and BSA) at different protein:dendrimer ratios (1:0, 1:1, 1:8, and 1:20) at acidic pH and SDS-PAGE profiles of the corresponsing supernatants. Comparison of SDS-PAGE profiles for the pellets obtained during the purification of proteins present in a complex sample using a conventional “no-clean” method based on acetone precipitation and the proposed “greener” method using carboxylate-terminated carbosilane dendrimer at a 1:20 protein:dendrimer ratio





Similar content being viewed by others
References
Wang W, Tai F, Chen S. Optimizing protein extraction from plant tissues for enhanced proteomics analysis. J Sep Sci. 2008;31(11):2032–9.
Wu X, Gong F, Wang W. Protein extraction from plant tissues for 2DE and its application in proteomic analysis. Proteomics. 2014;14(6):645–58.
Wang W, Vignani R, Scali M, Cresti M. A universal and rapid protocol for protein extraction from recalcitrant plant tissues for proteomic analysis. Electrophoresis. 2006;27(13):2782–6.
Wongpia A, Mahatheeranont S, Lomthaisong K, Niamsup H. Evaluation of sample preparation methods from rice seeds and seedlings suitable for two-dimensional gel electrophoresis. Appl Biochem Biotechnol. 2015;175(2):1035–51.
Banasiak A. Evolution of the cell wall components during terrestrialization. Acta Soc Bot Pol. 2014;83(4):349–62.
Ward WW, Swiatek G. Protein purification. Curr Anal Chem. 2009;5(2):85–105.
Wisniewski JR, Zielinska DF, Mann M. Comparison of ultrafiltration units for proteomic and N-glycoproteomic analysis by the filter-aided sample preparation method. Anal Biochem. 2011;410(2):307–9.
Feist P, Hummon AB. Proteomic challenges: sample preparation techniques for microgram-quantity protein analysis from biological samples. Int J Mol Sci. 2015;16(2):3537–63.
Jiang L, He L, Fountoulakis M. Comparison of protein precipitation methods for sample preparation prior to proteomic analysis. J Chromatogr A. 2004;1023(2):317–20.
Englard S, Seifter S. Precipitation techniques. In: Deutscher MP, editor. Methods in enzymology, vol. 182. New York: Academic Press, Inc; 1990.
Kalhapure RS, Kathiravan MK, Akamanchi KG, Govender T. Dendrimers—from organic synthesis to pharmaceutical applications: an update. Pharm Dev Technol. 2015;20(1):22–40.
Martinho N, Florindo H, Silva L, Brocchini S, Zloh M, Barata T. Molecular modeling to study dendrimers for biomedical applications. Molecules. 2014;19(12):20424–67.
Jiménez JL, Gómez R, Briz V, Madrid R, Bryszewsk M, de la Mata FJ, et al. Carbosilane dendrimers as carriers of siRNA. J Drug Delivery Sci Technol. 2012;22(1):75–82.
Schlenk C, Frey H. Carbosilane dendrimers—synthesis, functionalization, application. Mon Chem. 1999;130(1):3–14.
Galán M, Sánchez Rodríguez J, Jimenez JL, Relloso M, Maly M, de la Mata FJ, et al. Synthesis of new anionic carbosilane dendrimers via thiol-ene chemistry and their antiviral behaviour. Org Biomol Chem. 2014;12(20):3222–37.
Wang JM, Wolf RM, Caldwell JW, Kollman PA, Case DA. Development and testing of a general amber force field. J Comput Chem. 2004;25(9):1157–74.
Bayly CI, Cieplak P, Cornell WD, Kollman PA. A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges—the RESP model. J Phys Chem. 1993;97(40):10269–80.
Dupradeau F, Pigache A, Zaffran T, Savineau C, Lelong R, Grivel N, et al. The R.ED. tools: advances in RESP and ESP charge derivation and force field library building. Phys Chem Chem Phys. 2010;12(28):7821–39.
Gordon MS, Schmidt MW. Advances in electronic structure theory: GAMESS a decade later. In: Frenking G, Dykstra CE, editors. Theory and applications of computational chemistry: the first forty years. Amsterdam: Elsevier; 2005. p. 1167–89.
Bujacz A. Structures of bovine, equine and leporine serum albumin. Acta Crystallogr Sect D: Biol Crystallogr. 2012;68:1278–89.
Botha S, Nass K, Barends TRM, Kabsch W, Latz B, Dworkowski F, et al. Room-temperature serial crystallography at synchrotron X-ray sources using slowly flowing free-standing high-viscosity microstreams. Acta Crystallogr Sect D: Biol Crystallogr. 2015;71:387–97.
Hubbard SR, Hendrickson WA, Lambright DG, Boxer SG. X-ray crystal-structure of a recombinant human myoglobin mutant at 2.8 a resolution. J Mol Biol. 1990;213(2):215–8.
Case DA, Babin V, Berryman JT, Betz RM,. Cai Q, Cerutti DS, et al. AMBER 14. 2014.
Goetz AW, Williamson MJ, Xu D, Poole D, Le Grand S, Walker RC. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. Generalized born. J Chem Theory Comput. 2012;8(5):1542–55.
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, et al. UCSF chimera—a visualization system for exploratory research and analysis. J Comput Chem. 2004;25(13):1605–12.
González-García E, Marina ML, García MC. Plum (Prunus domestica L.) by-product as a new and cheap source of bioactive peptides: extraction method and peptides characterization. J Funct Foods. 2014;11:428–37.
Bradford MM. Rapid and sensitive method for quantitation of microgram quantities of protein utilizing principle of protein-dye binding. Anal Biochem. 1976;72(1–2):248–54.
Eftink MR. The use of fluorescence methods to monitor unfolding transitions in proteins. Biophys J. 1994;66(2):482–501.
Marden MC, Hoa GHB, Stetzkowskimarden F. Heme protein fluorescence versus pressure. Biophys J. 1986;49(3):619–27.
Hashimoto S, Fukasaka J, Takeuchi H. Structural study on acid-induced unfolding intermediates of myoglobin by using UV resonance Raman scattering from tryptophan residues. J Raman Spectrosc. 2001;32(6–7):557–63.
Baler K, Martin OA, Carignano MA, Ameer GA, Vila JA, Szleifer I. Electrostatic unfolding and interactions of albumin driven by pH changes: a molecular dynamics study. J Phys Chem B. 2014;118(4):921–30.
Lee JW, Kim HI. Investigating acid-induced structural transitions of lysozyme in an electrospray ionization source. Analyst. 2015;140(2):661–9.
Acknowledgments
This work was supported by the Ministry of Economy and Competitiveness (ref. AGL2012-36362 and CTQ-2014-54004-P), the Comunidad de Madrid and European funding from FEDER program (ref. S2013/ABI-3028, AVANSECAL), and Consortium NANODENDMED ref. S2011/BMD-2351 (CAM). The financial support of the Czech Science Foundation (project no. GA15-05903S) is also acknowledged. E.G.-G. thanks the University of Alcalá for her pre-doctoral contract. CIBER-BBN is an initiative funded by the VI National R&D&i Plan 2008–2011, Iniciativa Ingenio 2010, Consolider Program, CIBER Actions and financed by the Instituto de Salud Carlos III with assistance from the European Regional Development Fund.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1498 kb)
Rights and permissions
About this article
Cite this article
González-García, E., Maly, M., de la Mata, F.J. et al. Proof of concept of a “greener” protein purification/enrichment method based on carboxylate-terminated carbosilane dendrimer-protein interactions. Anal Bioanal Chem 408, 7679–7687 (2016). https://doi.org/10.1007/s00216-016-9864-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9864-6