Abstract
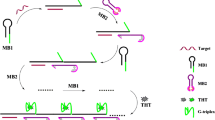
A powerful double-hairpin molecular beacon (DHMB) was developed for cancer-related KRAS gene detection based on the one-to-two stoichiometry. During target DNA detection, DHMB can execute signal transduction even if no any exogenous element is involved. Unlike the conventional molecular beacon based on the one-to-one interaction, one target DNA not only hybridizes with one DHMB and opens its hairpin but also promotes the interaction between two DHMBs, causing the separation of two fluorophores from quenchers. This leads to an enhanced fluorescence signal. As a result, the target KRAS gene is able to be detected within a wide dynamic range from 0.05 to 200 nM with the detection limit of 50 pM, indicating a dramatic improvement compared with traditional molecular beacons. Moreover, the point mutations existing in target DNAs can be easily screened. The potential application for target species in real samples was indicated by the analysis of PCR amplicons of DNAs from the DNA extracted from SW620 cell. Besides becoming a promising candidate probe for molecular biology research and clinical diagnosis of genetic diseases, the DHMB is expected to provide a significant insight into the design of DNA probe-based homogenous sensing systems.

A powerful double-hairpin molecular beacon (DHMB) was developed for cancer-related gene KRAS detection based on the one-to-two stoichiometry. Without the help of any exogenous probe, the point mutation is easily screened, and the target DNA can be quantified down to 50 pM, indicating a dramatic improvement compared with traditional molecular beacons






Similar content being viewed by others
References
Saito H, Sekizawa A, Morimoto T, Suzuki M, Yanaihara T. Prenatal DNA diagnosis of a single-gene disorder from maternal plasma. Lancet. 2000;356(9236):1170.
Williamson R, Eskdale J, Coleman D, Niazi M, Loeffler F, Modell B. Direct gene analysis of chorionic villi: a possible technique for first-trimester antenatal diagnosis of haemoglobinopathies. Lancet. 1981;318(8256):1125–7.
Meng H-M, Zhang X, Lv Y, Zhao Z, Wang N-N, Fu T, et al. DNA dendrimer: an efficient nanocarrier of functional nucleic acids for intracellular molecular sensing. ACS Nano. 2014;8(6):6171–81.
Yang Y, Huang J, Yang X, Quan K, Wang H, Ying L, et al. FRET nanoflares for intracellular mRNA detection: avoiding false positive signals and minimizing effects of system fluctuations. J Am Chem Soc. 2015;137(26):8340–3.
Qiu L, Wu C, You M, Han D, Chen T, Zhu G, et al. A targeted, self-delivered, and photocontrolled molecular beacon for mRNA detection in living cells. J Am Chem Soc. 2013;135(35):12952–5.
Deng R, Tang L, Tian Q, Wang Y, Lin L, Li J. Toehold-initiated rolling circle amplification for visualizing individual microRNAs in situ in single cells. Angew Chem Int Ed. 2014;53(9):2389–93.
Xuan F, Fan TW, Hsing I-M. Electrochemical interrogation of kinetically-controlled dendritic DNA/PNA assembly for immobilization-free and enzyme-free nucleic acids sensing. ACS Nano. 2015;9(5):5027–33.
Wang F, Lu C-H, Liu X, Freage L, Willner I. Amplified and multiplexed detection of DNA using the dendritic rolling circle amplified synthesis of DNAzyme reporter units. Anal Chem. 2014;86(3):1614–21.
Xuan F, Hsing I-M. Triggering hairpin-free chain-branching growth of fluorescent DNA dendrimers for nonlinear hybridization chain reaction. J Am Chem Soc. 2014;136(28):9810–3.
Wang F, Elbaz J, Orbach R, Magen N, Willner I. Amplified analysis of DNA by the autonomous assembly of polymers consisting of DNAzyme wires. J Am Chem Soc. 2011;133(43):17149–51.
Li Z, Miao X, Xing K, Zhu A, Ling L. Enhanced electrochemical recognition of double-stranded DNA by using hybridization chain reaction and positively charged gold nanoparticles. Biosens Bioelectron. 2015;74:687–90.
Miao XM, Xiong C, Wang WW, Ling LS, Shuai XT. Dynamic-light-scattering-based sequence-specific recognition of double-stranded DNA with oligonucleotide-functionalized gold nanoparticles. Chem-Eur J. 2011;17(40):11230–6.
Cheglakov Z, Cronin TM, He C, Weizmann Y. Live-cell microRNA imaging using cascade hybridization reaction. J Am Chem Soc. 2015.
Tyagi S, Marras SA, Kramer FR. Wavelength-shifting molecular beacons. Nat Biotechnol. 2000;18(11):1191–6.
Tyagi S, Kramer FR. Molecular beacons: probes that fluoresce upon hybridization. Nat Biotechnol. 1996;14(3):303–8.
Tyagi S, Bratu DP, Kramer FR. Multicolor molecular beacons for allele discrimination. Nat Biotechnol. 1998;16(1):49–53.
Zhou J, Lu Q, Tong Y, Wei W, Liu S. Detection of DNA damage by using hairpin molecular beacon probes and graphene oxide. Talanta. 2012;99:625–30.
Xiong Y, Wei M, Wei W, Yin L, Pu Y, Liu S. Detection of DNA damage based on metal-mediated molecular beacon and DNA strands displacement reaction. Spectrochim Acta A. 2014;118:806–10.
Johnston AP, Caruso F. A molecular beacon approach to measuring the DNA permeability of thin films. J Am Chem Soc. 2005;127(28):10014–5.
Lu L-M, Zhang X-B, Kong R-M, Yang B, Tan W. A ligation-triggered DNAzyme cascade for amplified fluorescence detection of biological small molecules with zero-background signal. J Am Chem Soc. 2011;133(30):11686–91.
Xue Q, Wang L, Jiang W. Label-free molecular beacon-based quadratic isothermal exponential amplification: a simple and sensitive one-pot method to detect DNA methyltransferase activity. Chem Commun. 2015;51(70):13538–41.
Ye T, Liu Y, Luo M, Xiang X, Ji X, Zhou G, et al. Metal–organic framework-based molecular beacons for multiplexed DNA detection by synchronous fluorescence analysis. Analyst. 2014;139(7):1721–5.
Miao X, Guo X, Xiao Z, Ling L. Electrochemical molecular beacon biosensor for sequence-specific recognition of double-stranded DNA. Biosens Bioelectron. 2014;59:54–7.
Li JJ, Chu Y, Lee BY-H, Xie XS. Enzymatic signal amplification of molecular beacons for sensitive DNA detection. Nucleic Acids Res. 2008;36(6), e36.
Nilsson M, Gullberg M, Dahl F, Szuhai K, Raap AK. Real-time monitoring of rolling-circle amplification using a modified molecular beacon design. Nucleic Acids Res. 2002;30(14):e66-e.
Zuo X, Xia F, Xiao Y, Plaxco KW. Sensitive and selective amplified fluorescence DNA detection based on exonuclease III-aided target recycling. J Am Chem Soc. 2010;132(6):1816–8.
Peyssonnaux C, Eychène A. The Raf/MEK/ERK pathway: new concepts of activation. Biol Cell. 2001;93(1-2):53–62.
Singer G, Kurman RJ, Chang H-W, Cho SK, Shih I-M. Diverse tumorigenic pathways in ovarian serous carcinoma. Am J Pathol. 2002;160(4):1223–8.
Martinez K, Estevez M-C, Wu Y, Phillips JA, Medley CD, Tan W. Locked nucleic acid based beacons for surface interaction studies and biosensor development. Anal Chem. 2009;81(9):3448–54.
Liu X, Tan W. A fiber-optic evanescent wave DNA biosensor based on novel molecular beacons. Anal Chem. 1999;71(22):5054–9.
Qiu L-P, Wu Z-S, Shen G-L, Yu R-Q. Highly sensitive and selective bifunctional oligonucleotide probe for homogeneous parallel fluorescence detection of protein and nucleotide sequence. Anal Chem. 2011;83(8):3050–7.
Hu J, Wen C-Y, Zhang Z-L, Xie M, Hu J, Wu M, et al. Optically encoded multifunctional nanospheres for one-pot separation and detection of multiplex DNA sequences. Anal Chem. 2013;85(24):11929–35.
Acknowledgments
This work was supported by National Natural Science Foundation of China (NSFC) (grant no: 21275002), Zhejiang Province Natural Science Foundation of China (LY16C07002), and Independent Research Project of State Key Laboratory of Photocatalysis on Energy and Environment (No. 2014CO1).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have declared that no competing interest exists.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 350 kb)
Rights and permissions
About this article
Cite this article
Xu, H., Zhang, R., Li, F. et al. Double-hairpin molecular-beacon-based amplification detection for gene diagnosis linked to cancer. Anal Bioanal Chem 408, 6181–6188 (2016). https://doi.org/10.1007/s00216-016-9729-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9729-z