Abstract
Cysteine is widely involved in redox signaling pathways through a number of reversible and irreversible modifications. Reversible modifications (e.g., S-glutathionylation, S-nitrosylation, disulfide bonds, and sulfenic acid) are used to protect proteins from oxidative attack and maintain cellular homeostasis, while irreversible oxidations (e.g., sulfinic acid and sulfonic acid) serve as hallmarks of oxidative stress. Proteomic analysis of cysteine-enriched peptides coupled with reduction of oxidized thiols can be used to measure the oxidation states of cysteine, which is helpful for elucidating the role that oxidative stress plays in biology and disease. As an extension of our previously reported cysDML method, we have developed oxidized cysteine-selective dimethylation (OxcysDML), to investigate the site-specific total oxidation of cysteine residues in biologically relevant samples. OxcysDML employs (1) blocking of free thiols by a cysteine-reactive reagent, (2) enrichment of peptides containing reversibly oxidized cysteine by a solid phase resin, and (3) isotopic labeling of peptide amino groups to quantify cysteine modifications arising from different biological conditions. On-resin enrichment and labeling minimizes sample handing time and improves efficiency in comparison with other redox proteomic methods. OxcysDML is also inexpensive and flexible, as it can accommodate the exploration of various cysteine modifications. Here, we applied the method to liver tissues from a late-stage Alzheimer’s disease (AD) mouse model and wild-type (WT) controls. Because we have previously characterized this proteome using the cysDML approach, we are able here to probe deeper into the redox status of cysteine in AD. OxcysDML identified 1129 cysteine sites (from 527 proteins), among which 828 cysteine sites underwent oxidative modifications. Nineteen oxidized cysteine sites had significant alteration levels in AD and represent proteins involved in metabolic processes. Overall, we have demonstrated OxcysDML as a simple, rapid, robust, and inexpensive redox proteomic approach that is useful for gaining deeper insight into the proteome of AD.

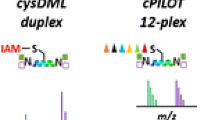
OxcysDML enables the proteome comparision of cysteine reversible oxidations from two biological conditions






Similar content being viewed by others
References
Tornvall U. Pinpointing oxidative modifications in proteins-recent advances in analytical methods. Anal Methods-UK. 2010;2(11):1638–50.
Boronat S, Garcia-Santamarina S, Hidalgo E. Gel-free proteomic methodologies to study reversible cysteine oxidation and irreversible protein carbonyl formation. Free Radic Res. 2015;49(5):494–510.
Verrastro I, Pasha S, Jensen KT, Pitt AR, Spickett CM. Mass spectrometry-based methods for identifying oxidized proteins in disease: advances and challenges. Biogeosciences. 2015;5(2):378–411.
Couvertier SM, Zhou Y, Weerapana E. Chemical-proteomic strategies to investigate cysteine posttranslational modifications. Bba Proteins Proteom. 2014;1844(12):2315–30.
Klomsiri C, Karplus PA, Poole LB. Cysteine-based redox switches in enzymes. Antioxid Redox Signal. 2011;14(6):1065–77.
Garcia-Santamarina S, Boronat S, Hidalgo E. Reversible cysteine oxidation in hydrogen peroxide sensing and signal transduction. Biochemistry. 2014;53(16):2560–80.
Baez NOD, Reisz JA, Furdui CM. Mass spectrometry in studies of protein thiol chemistry and signaling: opportunities and caveats. Free Radical Bio Med. 2015;80:191–211.
Biteau B, Labarre J, Toledano MB. ATP-dependent reduction of cysteine-sulphinic acid by S. cerevisiae sulphiredoxin. Nature. 2003;425(6961):980–4.
Chang TS, Jeong W, Woo HA, Lee SM, Park S, Rhee SG. Characterization of mammalian sulfiredoxin and its reactivation of hyperoxidized peroxiredoxin through reduction of cysteine sulfinic acid in the active site to cysteine. J Biol Chem. 2004;279(49):50994–1001.
Blennow K, de Leon MJ, Zetterberg H. Alzheimer’s disease. Lancet. 2006;368(9533):387–403.
Rosales-Corral S, Tan DX, Manchester L, Reiter RJ. Diabetes and Alzheimer disease, two overlapping pathologies with the same background: oxidative stress. Oxidative Med Cell Longev. 2015;2015:985845.
Newman SF, Sultana R, Perluigi M, Coccia R, Cai J, Pierce WM, et al. An increase in S-glutathionylated proteins in the Alzheimer’s disease inferior parietal lobule, a proteomics approach. J Neurosci Res. 2007;85(7):1506–14.
Zahid S, Khan R, Oellerich M, Ahmed N, Asif AR. Differential S-nitrosylation of proteins in Alzheimer’s disease. Neuroscience. 2014;256:126–36.
Zareba-Koziol M, Szwajda A, Dadlez M, Wyslouch-Cieszynska A, Lalowski M. Global analysis of S-nitrosylation sites in the wild type and APP transgenic mouse brain—clues for synaptic pathology. Mol Cell Proteomics MCP. 2014;13:2288–305.
Riederer IM, Schiffrin M, Kovari E, Bouras C, Riederer BM. Ubiquitination and cysteine nitrosylation during aging and Alzheimer’s disease. Brain Res Bull. 2009;80(4–5):233–41.
Chen CH, Li W, Sultana R, You MH, Kondo A, Shahpasand K, et al. Pin1 cysteine-113 oxidation inhibits its catalytic activity and cellular function in Alzheimer’s disease. Neurobiol Dis. 2015;76:13–23.
Butterfield DA, Gu L, Domenico FD, Robinson RA. Mass spectrometry and redox proteomics: applications in disease. Mass Spectrom Rev. 2013;33(4):277–301.
Li R, Huang JQ, Kast J. Identification of total reversible cysteine oxidation in an atherosclerosis model using a modified biotin switch assay. J Proteome Res. 2015;14(5):2026–35.
Liu P, Zhang H, Wang H, Xia Y. Identification of redox-sensitive cysteines in the Arabidopsis proteome using OxiTRAQ, a quantitative redox proteomics method. Proteomics. 2014;14(6):750–62.
Gygi SP, Rist B, Gerber SA, Turecek F, Gelb MH, Aebersold R. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat Biotechnol. 1999;17(10):994–9.
Garcia-Santamarina S, Boronat S, Espadas G, Ayte J, Molina H, Hidalgo E. The oxidized thiol proteome in fission yeast—optimization of an ICAT-based method to identify H2O2-oxidized proteins. J Proteome. 2011;74(11):2476–86.
Garcia-Santamarina S, Boronat S, Domenech A, Ayte J, Molina H, Hidalgo E. Monitoring in vivo reversible cysteine oxidation in proteins using ICAT and mass spectrometry. Nat Protoc. 2014;9(5):1131–45.
Guo J, Gaffrey MJ, Su D, Liu T, Camp 2nd DG, Smith RD, et al. Resin-assisted enrichment of thiols as a general strategy for proteomic profiling of cysteine-based reversible modifications. Nat Protoc. 2014;9(1):64–75.
Forrester MT, Thompson JW, Foster MW, Nogueira L, Moseley MA, Stamler JS. Proteomic analysis of S-nitrosylation and denitrosylation by resin-assisted capture. Nat Biotechnol. 2009;27(6):557–9.
Forrester MT, Hess DT, Thompson JW, Hultman R, Moseley MA, Stamler JS, et al. Site-specific analysis of protein S-acylation by resin-assisted capture. J Lipid Res. 2011;52(2):393–8.
Pan KT, Chen YY, Pu TH, Chao YS, Yang CY, Bomgarden RD, et al. Mass spectrometry-based quantitative proteomics for dissecting multiplexed redox cysteine modifications in nitric oxide-protected cardiomyocyte under hypoxia. Antioxid Redox Signal. 2014;20(9):1365–81.
Xiang F, Ye H, Chen R, Fu Q, Li L. N, N-dimethyl leucines as novel isobaric tandem mass tags for quantitative proteomics and peptidomics. Anal Chem. 2010;82(7):2817–25.
Gu L, Evans AR, Robinson RA. Sample multiplexing with cysteine-selective approaches: cysDML and cPILOT. J Am Soc Mass Spectrom. 2015;26(4):615–30.
Qu Z, Meng F, Bomgarden RD, Viner RI, Li J, Rogers JC, et al. Proteomic quantification and site-mapping of S-nitrosylated proteins using isobaric iodoTMT reagents. J Proteome Res. 2014;13(7):3200–11.
Bantscheff M, Lemeer S, Savitski MM, Kuster B. Quantitative mass spectrometry in proteomics: critical review update from 2007 to the present. Anal Bioanal Chem. 2012;404(4):939–65.
Wu R, Dephoure N, Haas W, Huttlin EL, Zhai B, Sowa ME, et al. Correct interpretation of comprehensive phosphorylation dynamics requires normalization by protein expression changes. Mol Cell Proteomics MCP. 2011;10(8):M111 009654.
Kumar V, Kleffmann T, Hampton MB, Cannell MB, Winterbourn CC. Redox proteomics of thiol proteins in mouse heart during ischemia/reperfusion using ICAT reagents and mass spectrometry. Free Radical Bio Med. 2013;58:109–17.
Dayon L, Sonderegger B, Kussmann M. Combination of gas-phase fractionation and MS(3) acquisition modes for relative protein quantification with isobaric tagging. J Proteome Res. 2012;11(10):5081–9.
Wu Y, Wang F, Liu Z, Qin H, Song C, Huang J, et al. Five-plex isotope dimethyl labeling for quantitative proteomics. Chem Commun. 2014;50(14):1708–10.
Wojdyla K, Williamson J, Roepstorff P, Rogowska-Wrzesinska A. The SNO/SOH TMT strategy for combinatorial analysis of reversible cysteine oxidations. J Proteome. 2015;113:415–34.
Su D, Shukla AK, Chen B, Kim JS, Nakayasu E, Qu Y, et al. Quantitative site-specific reactivity profiling of S-nitrosylation in mouse skeletal muscle using cysteinyl peptide enrichment coupled with mass spectrometry. Free Radic Biol Med. 2013;57:68–78.
Evans AR, Gu L, Guerrero Jr R, Robinson RA. Global cPILOT analysis of the APP/PS-1 mouse liver proteome. Proteomics Clin Appl. 2015;9(9–10):872–84.
Chen D, Shah A, Nguyen H, Loo D, Inder KL, Hill MM. Online quantitative proteomics p-value calculator for permutation-based statistical testing of peptide ratios. J Proteome Res. 2014;13(9):4184–91.
Lau HT, Suh HW, Golkowski M, Ong SE. Comparing SILAC- and stable isotope dimethyl-labeling approaches for quantitative proteomics. J Proteome Res. 2014;13(9):4164–74.
Shi R, Kumar C, Zougman A, Zhang Y, Podtelejnikov A, Cox J, et al. Analysis of the mouse liver proteome using advanced mass spectrometry. J Proteome Res. 2007;6(8):2963–72.
Go YM, Roede JR, Orr M, Liang Y, Jones DP. Integrated redox proteomics and metabolomics of mitochondria to identify mechanisms of cd toxicity. Toxicol Sci Off J Soc Toxicol. 2014;139(1):59–73.
Bairoch A, Apweiler R, Wu CH, Barker WC, Boeckmann B, Ferro S, et al. The Universal Protein Resource (UniProt). Nucleic Acids Res. 2005;33(Database issue):D154–9.
Pace NJ, Weerapana E. Zinc-binding cysteines: diverse functions and structural motifs. Biogeosciences. 2014;4(2):419–34.
Morris JK, Honea RA, Vidoni ED, Swerdlow RH, Burns JM. Is Alzheimer’s disease a systemic disease? Bba Mol Basis Dis. 2014;1842(9):1340–9.
Robinson RA, Cao Z, Williams C. Oxidative stress in CD90+ T-cells of AbetaPP/PS-1 transgenic mice. J Alzheimer’s Dis JAD. 2013;37(4):661–6.
Swomley AM, Forster S, Keeney JT, Triplett J, Zhang Z, Sultana R, et al. Abeta, oxidative stress in Alzheimer disease: evidence based on proteomics studies. Bba Mol Basis Dis. 2014;1842(8):1248–57.
Butterfield DA, Perluigi M, Reed T, Muharib T, Hughes CP, Robinson RA, et al. Redox proteomics in selected neurodegenerative disorders: from its infancy to future applications. Antioxid Redox Signal. 2012;17(11):1610–55.
Di Domenico F, Cenini G, Sultana R, Perluigi M, Uberti D, Memo M, et al. Glutathionylation of the pro-apoptotic protein p53 in Alzheimer’s disease brain: implications for AD pathogenesis. Neurochem Res. 2009;34(4):727–33.
Hooijmans CR, Rutters F, Dederen PJ, Gambarota G, Veltien A, van Groen T, et al. Changes in cerebral blood volume and amyloid pathology in aged Alzheimer APP/PS1 mice on a docosahexaenoic acid (DHA) diet or cholesterol enriched typical western diet (TWD). Neurobiol Dis. 2007;28(1):16–29.
Sagare A, Deane R, Bell RD, Johnson B, Hamm K, Pendu R, et al. Clearance of amyloid-beta by circulating lipoprotein receptors. Nat Med. 2007;13(9):1029–31.
Abdul HM, Sultana R, Clair DKS, Markesbery WR, Butterfield DA. Oxidative damage in brain from human mutant APP/PS-1 double knock-in mice as a function of age. Free Radical Bio Med. 2008;45(10):1420–5.
Sinclair AJ, Bayer AJ, Johnston J, Warner C, Maxwell SRJ. Altered plasma antioxidant status in subjects with Alzheimer’s disease and vascular dementia. Int J Geriatr Psych. 1998;13(12):840–5.
Smith MA, Rottkamp CA, Nunomura A, Raina AK, Perry G. Oxidative stress in Alzheimer’s disease. Biochim Biophys Acta. 2000;1502(1):139–44.
Cichoz-Lach H, Michalak A. Oxidative stress as a crucial factor in liver diseases. World J Gastroenterol. 2014;20(25):8082–91.
Fransen M, Nordgren M, Wang B, Apanasets O. Role of peroxisomes in ROS/RNS-metabolism: implications for human disease. Biochim Biophys Acta. 2012;1822(9):1363–73.
Acknowledgments
This research was supported by the University of Pittsburgh Start-up Funds. The authors acknowledge Xi Wang for assistance with statistical testing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Animal protocols were approved by the Institutional Animal Care and Use Committee at the University of Pittsburgh.
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Published in the topical collection featuring Young Investigators in Analytical and Bioanalytical Science with guest editors S. Daunert, A. Baeumner, S. Deo, J. Ruiz Encinar, and L. Zhang.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 389 kb)
Rights and permissions
About this article
Cite this article
Gu, L., Robinson, R.A.S. A simple isotopic labeling method to study cysteine oxidation in Alzheimer’s disease: oxidized cysteine-selective dimethylation (OxcysDML). Anal Bioanal Chem 408, 2993–3004 (2016). https://doi.org/10.1007/s00216-016-9307-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9307-4