Abstract
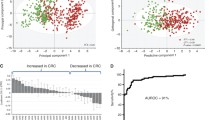
Colorectal cancer (CRC) is one of the most prevalent cancers worldwide and a major cause of human morbidity and mortality. In addition to early detection, close monitoring of disease progression in CRC can be critical for patient prognosis and treatment decisions. Efforts have been made to develop new methods for improved early detection and patient monitoring; however, research focused on CRC surveillance for treatment response and disease recurrence using metabolomics has yet to be reported. In this proof of concept study, we applied a targeted liquid chromatography tandem mass spectrometry (LC-MS/MS) metabolic profiling approach focused on sequential metabolite ratio analysis of serial serum samples to monitor disease progression from 20 CRC patients. The use of serial samples reduces patient to patient metabolic variability. A partial least squares-discriminant analysis (PLS-DA) model using a panel of five metabolites (succinate, N2, N2-dimethylguanosine, adenine, citraconic acid, and 1-methylguanosine) was established, and excellent model performance (sensitivity = 0.83, specificity = 0.94, area under the receiver operator characteristic curve (AUROC) = 0.91 was obtained, which is superior to the traditional CRC monitoring marker carcinoembryonic antigen (sensitivity = 0.75, specificity = 0.76, AUROC = 0.80). Monte Carlo cross validation was applied, and the robustness of our model was clearly observed by the separation of true classification models from the random permutation models. Our results suggest the potential utility of metabolic profiling for CRC disease monitoring.


Similar content being viewed by others
References
Siegel RL, Miller KD, Jemal A (2015) Cancer statistics, 2015. CA Cancer J Clin 65(1):5–29
Zhang A, Sun H, Yan G, Wang P, Han Y, Wang X (2014) Metabolomics in diagnosis and biomarker discovery of colorectal cancer. Cancer Lett 345(1):17–20
Mook ORF, Frederiks WM, Van Noorden CJF (2004) The role of gelatinases in colorectal cancer progression and metastasis. Biochim Biophys Acta (BBA) Rev Cancer 1705(2):69–89
Fletcher RH (1986) Carcinoembryonic antigen. Ann Intern Med 104(1):66–73
Staab HJ, Anderer FA, Stumpf E, Fischer R (1978) Slope analysis of the postoperative CEA time course and its possible application as an aid in diagnosis of disease progression in gastrointestinal cancer. Am J Surg 136(3):322–327
Ludwig JA, Weinstein JN (2005) Biomarkers in cancer staging, prognosis and treatment selection. Nat Rev Cancer 5(11):845–856
Duffy MJ (2001) Carcinoembryonic antigen as a marker for colorectal cancer: is it clinically useful? Clin Chem 47(4):624–630
Zhu J, Djukovic D, Deng L, Gu H, Himmati F, Chiorean EG, Raftery D (2014) Colorectal cancer detection using targeted serum metabolic profiling. J Proteome Res 13(9):4120–4130
Xia J, Mandal R, Sinelnikov IV, Broadhurst D, Wishart DS (2012) MetaboAnalyst 2.0—a comprehensive server for metabolomic data analysis. Nucleic Acids Res 40(W1):W127–W133
Iwanicki-Caron I, Di Fiore F, Roque I, Astruc E, Stetiu M, Duclos A, Tougeron D, Saillard S, Thureau S, Benichou J, Paillot B, Basuyau JP, Michel P (2008) Usefulness of the serum carcinoembryonic antigen kinetic for chemotherapy monitoring in patients with unresectable metastasis of colorectal cancer. J Clin Oncol 26(22):3681–3686
Brereton RG (2006) Consequences of sample size, variable selection, and model validation and optimisation, for predicting classification ability from analytical data. TrAC Trends Anal Chem 25(11):1103–1111
Li H, Wang J, Xu H, Xing R, Pan Y, Li W, Cui J, Zhang H, Lu Y (2013) Decreased fructose-1,6-bisphosphatase-2 expression promotes glycolysis and growth in gastric cancer cells. Mol Cancer 12(1):110
Lu C-W, Lin S-C, Chien C-W, Lin S-C, Lee C-T, Lin B-W, Lee J-C, Tsai S-J (2011) Overexpression of pyruvate dehydrogenase kinase 3 increases drug resistance and early recurrence in colon cancer. Am J Pathol 179(3):1405–1414
Thangaraju M, Carswell KN, Prasad PD, Ganapathy V (2009) Colon cancer cells maintain low levels of pyruvate to avoid cell death caused by inhibition of HDAC1/HDAC3. Biochem J 417(1):379–389
Hsu W-Y, Chen WT-L, Lin W-D, Tsai F-J, Tsai Y, Lin C-T, Lo W-Y, Jeng L-B, Lai C-C (2009) Analysis of urinary nucleosides as potential tumor markers in human colorectal cancer by high performance liquid chromatography/electrospray ionization tandem mass spectrometry. Clin Chim Acta 402(1–2):31–37
Gehrke CW, Kuo KC, Waalkes TP, Borek E (1979) Patterns of urinary excretion of modified nucleosides. Cancer Res 39(4):1150–1153
Hirayama A, Kami K, Sugimoto M, Sugawara M, Toki N, Onozuka H, Kinoshita T, Saito N, Ochiai A, Tomita M, Esumi H, Soga T (2009) Quantitative metabolome profiling of colon and stomach cancer microenvironment by capillary electrophoresis time-of-flight mass spectrometry. Cancer Res 69(11):4918–4925
Acknowledgments
This work was supported by AMRMC grant W81XWH-10-1-0540. The authors would like to thank Dr. Li Yuan Bermel (Purdue University Cancer Center) for assistance with the Cancer Care Engineering project sample collection bank. The China Scholarship Council is also gratefully acknowledged (grant to LD).
Potential conflict of interest
Daniel Raftery holds equity and an executive role at Matrix-Bio, Inc.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1628 kb)
Rights and permissions
About this article
Cite this article
Zhu, J., Djukovic, D., Deng, L. et al. Targeted serum metabolite profiling and sequential metabolite ratio analysis for colorectal cancer progression monitoring. Anal Bioanal Chem 407, 7857–7863 (2015). https://doi.org/10.1007/s00216-015-8984-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-015-8984-8