Abstract
Exosomes participate in cancer metastasis, but studying them presents unique challenges as a result of their small size and purification difficulties. Asymmetrical field flow fractionation with in-line ultraviolet absorbance, dynamic light scattering, and multi-angle light scattering was applied to the size separation and characterization of non-labeled B16-F10 exosomes from an aggressive mouse melanoma cell culture line. Fractions were collected and further analyzed using batch mode dynamic light scattering, transmission electron microscopy and compared with known size standards. Fractogram peak positions and computed radii show good agreement between samples and across fractions. Ultraviolet absorbance fractograms in combination with transmission electron micrographs were able to resolve subtle heterogeneity of vesicle retention times between separate batches of B16-F10 exosomes collected several weeks apart. Further, asymmetrical field flow fractionation also effectively separated B16-F10 exosomes into vesicle subpopulations by size. Overall, the flow field flow fractionation instrument combined with multiple detectors was able to rapidly characterize and separate exosomes to a degree not previously demonstrated. These approaches have the potential to facilitate a greater understanding of exosome function by subtype, as well as ultimately allow for “label-free” isolation of large scale clinical exosomes for the purpose of developing future exosome-based diagnostics and therapeutics.

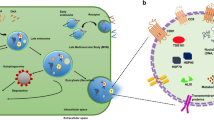
Flow path of exosome sample through the asymmetrical field flow fractionation instrument, detectors, and transmission electron microscope.







Similar content being viewed by others
References
Schorey JS, Bhatnagar S (2008) Exosome function: from tumor immunology to pathogen biology. Traffic (Copenhagen, Denmark) 9(6):871–881. doi:10.1111/j.1600-0854.2008.00734.x
Hood JL, Wickline SA (2012) A systematic approach to exosome-based translational nanomedicine. Wiley Interdiscip Rev Nanomed Nanobiotechnol 4(4):458–467. doi:10.1002/wnan.1174
D'Souza-Schorey C, Clancy JW (2012) Tumor-derived microvesicles: shedding light on novel microenvironment modulators and prospective cancer biomarkers. Genes Dev 26(12):1287–1299. doi:10.1101/gad.192351.112
Thery C, Ostrowski M, Segura E (2009) Membrane vesicles as conveyors of immune responses. Nat Rev Immunol 9(8):581–593. doi:10.1038/nri2567
Peinado H, Aleckovic M, Lavotshkin S, Matei I, Costa-Silva B, Moreno-Bueno G, Hergueta-Redondo M, Williams C, Garcia-Santos G, Ghajar C, Nitadori-Hoshino A, Hoffman C, Badal K, Garcia BA, Callahan MK, Yuan J, Martins VR, Skog J, Kaplan RN, Brady MS, Wolchok JD, Chapman PB, Kang Y, Bromberg J, Lyden D (2012) Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med 18(6):883–891. doi:10.1038/nm.2753
Sokolova V, Ludwig AK, Hornung S, Rotan O, Horn PA, Epple M, Giebel B (2011) Characterisation of exosomes derived from human cells by nanoparticle tracking analysis and scanning electron microscopy. Colloids Surf B Biointerfaces 87(1):146–150. doi:10.1016/j.colsurfb.2011.05.013
Gyorgy B, Szabo TG, Pasztoi M, Pal Z, Misjak P, Aradi B, Laszlo V, Pallinger E, Pap E, Kittel A, Nagy G, Falus A, Buzas EI (2011) Membrane vesicles, current state-of-the-art: emerging role of extracellular vesicles. Cell Mol Life Sci: CMLS 68(16):2667–2688. doi:10.1007/s00018-011-0689-3
van der Pol E, Boing AN, Harrison P, Sturk A, Nieuwland R (2012) Classification, functions, and clinical relevance of extracellular vesicles. Pharmacol Rev 64(3):676–705. doi:10.1124/pr.112.005983
Hood JL, Pan H, Lanza GM, Wickline SA (2009) Paracrine induction of endothelium by tumor exosomes. Lab Investig; J Technical Methods and Pathology 89(11):1317–1328. doi:10.1038/labinvest.2009.94
Weitz DA Dynamic Light Scattering (QLS, PCS). http://weitzlab.seas.harvard.edu/links/tutorials/dynamiclightscattering.pdf. Accessed 24 Apr 2014
Bobrie A, Colombo M, Raposo G, Thery C (2011) Exosome secretion: molecular mechanisms and roles in immune responses. Traffic (Copenhagen, Denmark) 12(12):1659–1668. doi:10.1111/j.1600-0854.2011.01225.x
van der Pol E, Hoekstra AG, Sturk A, Otto C, van Leeuwen TG, Nieuwland R (2010) Optical and non-optical methods for detection and characterization of microparticles and exosomes. J Thromb Haemost : JTH 8(12):2596–2607. doi:10.1111/j.1538-7836.2010.04074.x
Hood JL, San RS, Wickline SA (2011) Exosomes released by melanoma cells prepare sentinel lymph nodes for tumor metastasis. Cancer Res 71(11):3792–3801. doi:10.1158/0008-5472.can-10-4455
Mϋller G (2012) Novel tools for the study of cell type-specific exosomes and microvesicles. J Bioanal Biomed 4(4):46–60. doi:10.4172/1948-593X.1000063
Hood JL, Scott MJ, Wickline SA (2013) Maximizing exosome colloidal stability following electroporation. Anal Biochem 448C:41–49. doi:10.1016/j.ab.2013.12.001
Pan BT, Johnstone RM (1983) Fate of the transferrin receptor during maturation of sheep reticulocytes in vitro: selective externalization of the receptor. Cell 33(3):967–978
Carayon K, Chaoui K, Ronzier E, Lazar I, Bertrand-Michel J, Roques V, Balor S, Terce F, Lopez A, Salome L, Joly E (2011) Proteolipidic composition of exosomes changes during reticulocyte maturation. J Biol Chem 286(39):34426–34439. doi:10.1074/jbc.M111.257444
Al-Nedawi K, Meehan B, Micallef J, Lhotak V, May L, Guha A, Rak J (2008) Intercellular transfer of the oncogenic receptor EGFRvIII by microvesicles derived from tumour cells. Nat Cell Biol 10(5):619–624. doi:10.1038/ncb1725
Skog J, Wurdinger T, van Rijn S, Meijer DH, Gainche L, Sena-Esteves M, Curry WT Jr, Carter BS, Krichevsky AM, Breakefield XO (2008) Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol 10(12):1470–1476. doi:10.1038/ncb1800
Safaei R, Larson BJ, Cheng TC, Gibson MA, Otani S, Naerdemann W, Howell SB (2005) Abnormal lysosomal trafficking and enhanced exosomal export of cisplatin in drug-resistant human ovarian carcinoma cells. Mol Cancer Ther 4(10):1595–1604. doi:10.1158/1535-7163.mct-05-0102
Mathivanan S, Simpson RJ (2009) ExoCarta: a compendium of exosomal proteins and RNA. Proteomics 9(21):4997–5000. doi:10.1002/pmic.200900351
Mathivanan S, Fahner CJ, Reid GE, Simpson RJ (2012) ExoCarta 2012: database of exosomal proteins, RNA and lipids. Nucleic Acids Res 40(Database issue):D1241–D1244. doi:10.1093/nar/gkr828
Kalra H, Simpson RJ, Ji H, Aikawa E, Altevogt P, Askenase P, Bond VC, Borras FE, Breakefield X, Budnik V, Buzas E, Camussi G, Clayton A, Cocucci E, Falcon-Perez JM, Gabrielsson S, Gho YS, Gupta D, Harsha HC, Hendrix A, Hill AF, Inal JM, Jenster G, Kramer-Albers EM, Lim SK, Llorente A, Lotvall J, Marcilla A, Mincheva-Nilsson L, Nazarenko I, Nieuwland R, Nolte-'t Hoen EN, Pandey A, Patel T, Piper MG, Pluchino S, Prasad TS, Rajendran L, Raposo G, Record M, Reid GE, Sanchez-Madrid F, Schiffelers RM, Siljander P, Stensballe A, Stoorvogel W, Taylor D, Thery C, Valadi H, van Balkom BW, Vazquez J, Vidal M, Wauben MH, Yanez-Mo M, Zoeller M, Mathivanan S (2012) Vesiclepedia: a compendium for extracellular vesicles with continuous community annotation. PLoS Biol 10(12):e1001450. doi:10.1371/journal.pbio.1001450
Thery C, Amigorena S, Raposo G, Clayton A (2006) Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Current protocols in cell biology//editorial board, Juan S Bonifacino [et al.] Chapter 3:Unit 3.22. doi:10.1002/0471143030.cb0322s30
Taylor D, Zacharias W, Gercel-Taylor C (2011) Exosome Isolation for Proteomic Analyses and RNA Profiling. In: Simpson RJ, Greening DW (eds) Serum/Plasma Proteomics, vol 728. Methods in Molecular Biology. Humana, pp 235–246. doi:10.1007/978-1-61779-068-3_15
Chen C, Skog J, Hsu CH, Lessard RT, Balaj L, Wurdinger T, Carter BS, Breakefield XO, Toner M, Irimia D (2010) Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab Chip 10(4):505–511. doi:10.1039/b916199f
Raj DAA, Fiume I, Capasso G, Pocsfalvi G (2012) Urinary exosomes for protein biomarker research. In: Man TK, Prof (ed) Proteomics—human diseases and protein functions. In Tech, pp 49–64
Lesieur S, Grabielle-Madelmont C, Paternostre M, Ollivon M (1993) Study of size distribution and stability of liposomes by high performance gel exclusion chromatography. Chem Phys Lipids 64(1):57–82
Ruysschaert T, Marque A, Duteyrat JL, Lesieur S, Winterhalter M, Fournier D (2005) Liposome retention in size exclusion chromatography. BMC Biotechnol 5:11. doi:10.1186/1472-6750-5-11
Ingebrigtsen L, Brandl M (2002) Determination of the size distribution of liposomes by SEC fractionation, and PCS analysis and enzymatic assay of lipid content. AAPS PharmSciTech 3(2):E7. doi:10.1208/pt030207
Lai RC, Arslan F, Lee MM, Sze NS, Choo A, Chen TS, Salto-Tellez M, Timmers L, Lee CN, El Oakley RM, Pasterkamp G, de Kleijn DP, Lim SK (2010) Exosome secreted by MSC reduces myocardial ischemia/reperfusion injury. Stem Cell Res 4(3):214–222. doi:10.1016/j.scr.2009.12.003
Oshima K, Aoki N, Kato T, Kitajima K, Matsuda T (2002) Secretion of a peripheral membrane protein, MFG-E8, as a complex with membrane vesicles. Eur J Biochem FEBS 269(4):1209–1218
Chen TS, Arslan F, Yin Y, Tan SS, Lai RC, Choo AB, Padmanabhan J, Lee CN, de Kleijn DP, Lim SK (2011) Enabling a robust scalable manufacturing process for therapeutic exosomes through oncogenic immortalization of human ESC-derived MSCs. J Transl Med 9:47. doi:10.1186/1479-5876-9-47
Rekker K, Saare M, Roost AM, Kubo AL, Zarovni N, Chiesi A, Salumets A, Peters M (2014) Comparison of serum exosome isolation methods for microRNA profiling. Clin Biochem 47(1–2):135–138. doi:10.1016/j.clinbiochem.2013.10.020
Yamada T, Inoshima Y, Matsuda T, Ishiguro N (2012) Comparison of methods for isolating exosomes from bovine milk. J Vet Med Sci///Japanese Society of Veterinary Science 74(11):1523–1525
Cheruvanky A, Zhou H, Pisitkun T, Kopp JB, Knepper MA, Yuen PS, Star RA (2007) Rapid isolation of urinary exosomal biomarkers using a nanomembrane ultrafiltration concentrator. Am J Physiol Renal Physiol 292(5):F1657–F1661. doi:10.1152/ajprenal.00434.2006
Merchant ML, Powell DW, Wilkey DW, Cummins TD, Deegens JK, Rood IM, McAfee KJ, Fleischer C, Klein E, Klein JB (2010) Microfiltration isolation of human urinary exosomes for characterization by MS. Proteomics Clin Appl 4(1):84–96. doi:10.1002/prca.200800093
Oh S, Kang D, Ahn SM, Simpson RJ, Lee BH, Moon MH (2007) Miniaturized asymmetrical flow field-flow fractionation: application to biological vesicles. J Sep Sci 30(7):1082–1087
Kang D, Oh S, Ahn S-M, Lee B-H, Moon MH (2008) Proteomic analysis of exosomes from human neural stem cells by flow field-flow fractionation and nanoflow liquid chromatography-tandem mass spectrometry. J Proteome Res 7(8):3475–3480
Lasser C, Eldh M, Lotvall J (2012) Isolation and characterization of RNA-containing exosomes. J Visualized Experiments : JoVE 59:e3037. doi:10.3791/3037
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9(6):654–659. doi:10.1038/ncb1596
Jorgensen M, Baek R, Pedersen S, Sondergaard EK, Kristensen SR, Varming K (2013) Extracellular Vesicle (EV) Array: microarray capturing of exosomes and other extracellular vesicles for multiplexed phenotyping. J Extracellular Vesicles 2. doi: 10.3402/jev.v2i0.20920
Tauro BJ, Greening DW, Mathias RA, Ji H, Mathivanan S, Scott AM, Simpson RJ (2012) Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods (San Diego, Calif) 56(2):293–304. doi:10.1016/j.ymeth.2012.01.002
Wahlund KG, Giddings JC (1987) Properties of an asymmetrical flow field-flow fractionation channel having one permeable wall. Anal Chem 59(9):1332–1339
Qureshi RN, Kok WT (2011) Application of flow field-flow fractionation for the characterization of macromolecules of biological interest: a review. Anal Bioanal Chem 399(4):1401–1411. doi:10.1007/s00216-010-4278-3
Korgel BA, van Zanten JH, Monbouquette HG (1998) Vesicle size distributions measured by flow field-flow fractionation coupled with multiangle light scattering. Biophys J 74(6):3264–3272. doi:10.1016/s0006-3495(98)78033-6
Paulaitis M, Guzman N, Agarwal K, Saji M (2010) Poster Session 3—tumor cell and molecular biology: MicroRNAs Abstract P3-09-04: exosome-specific microRNA signatures in combination with characteristic surface markers on the circulating exosomes themselves provide new insights in the EMT. Cancer Res 70 (24). doi:10.1158/0008-5472.SABCS10-P3-09-04
Roda B, Zattoni A, Reschiglian P, Moon MH, Mirasoli M, Michelini E, Roda A (2009) Field-flow fractionation in bioanalysis: a review of recent trends. Anal Chim Acta 635(2):132–143. doi:10.1016/j.aca.2009.01.015
Mitrus I, Bryndza E, Kazura M, Smagur A, Sochanik A, Cichon T, Szala S (2012) Properties of B16-F10 murine melanoma cells subjected to metabolic stress conditions. Acta Biochim Pol 59(3):363–366
Wyatt Technology Corporation (2010) DYNAMICS User's Guide. In., Version 7.1 (M1400 Rev. K) edn., pp Ch8-8, Appendix A
Wyatt Technology Corporation (2008) Astra V User's Guide. In, vol Version 5.3.4 (M1000 Rev. H). pp Appendix D-F
Belnap D (2013) Negative Stain Procedure. http://www.cores.utah.edu/wp-content/uploads/2013/06/negative-stain.pdf. Accessed 13 Nov 2013
Wyatt Technology Corporation (2011) Introduction to light scattering. Light Scattering University. Santa Barbara California
Pencer J, White GF, Hallett FR (2001) Osmotically induced shape changes of large unilamellar vesicles measured by dynamic light scattering. Biophys J 81(5):2716–2728. doi:10.1016/s0006-3495(01)75914-0
Pencer J, Hallett FR (2003) Effects of vesicle size and shape on static and dynamic light scattering measurements. Langmuir 19(18):7488–7497. doi:10.1021/la0345439
Acknowledgments
This research was supported in part by the US National Science Foundation Division of Graduate Education under grant DGE-0903715 and in part by the National Institutes of Health under grant 1R21GM107894-01. We would also like to thank Michael J. Scott at C-TRAIN for assisting in exosome collections and shipments between C-TRAIN at Washington University and the University of Utah laboratories. Additional grant support for C-TRAIN investigators included the Elsa U. Pardee Foundation (J. L. Hood) and the National Institutes of Health R01HL073646-08 (S. A. Wickline).
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in the topical collection Field- and Flow-based Separations with guest editors Gaetane Lespes, Catia Contado, and Bruce Gale.
Rights and permissions
About this article
Cite this article
Petersen, K.E., Manangon, E., Hood, J.L. et al. A review of exosome separation techniques and characterization of B16-F10 mouse melanoma exosomes with AF4-UV-MALS-DLS-TEM. Anal Bioanal Chem 406, 7855–7866 (2014). https://doi.org/10.1007/s00216-014-8040-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-014-8040-0