Abstract
PEGylation has been widely used to improve the biopharmaceutical properties of therapeutic proteins and peptides. Previous studies have used multiple analytical techniques to determine the fate of both the therapeutic molecule and unconjugated poly(ethylene glycol) (PEG) after drug administration. A straightforward strategy utilizing liquid chromatography–mass spectrometry (LC–MS) to characterize high-molecular weight PEG in biologic matrices without a need for complex sample preparation is presented. The method is capable of determining whether high-MW PEG is cleaved in vivo to lower-molecular weight PEG species. Reversed-phase chromatographic separation is used to take advantage of the retention principles of polymeric materials whereby elution order correlates with PEG molecular weight. In-source collision-induced dissociation (CID) combined with selected reaction monitoring (SRM) or selected ion monitoring (SIM) mass spectrometry (MS) is then used to monitor characteristic PEG fragment ions in biological samples. MS provides high sensitivity and specificity for PEG and the observed retention times in reversed-phase LC enable estimation of molecular weight. This method was successfully used to characterize PEG molecular weight in mouse serum samples. No change in molecular weight was observed for 48 h after dosing.

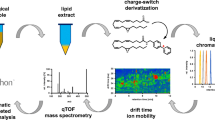
Correlation between log PEG MW and retention time observed by reversed-phase LC-MS with in-source fragmentation



Similar content being viewed by others
References
Kou D, Manius G, Zhan S, Chokshi HP (2009) Size exclusion chromatography with Corona charged aerosol detector for the analysis of polyethylene glycol polymer. J Chromatogr A 1216(28):5424–5428. doi:10.1016/j.chroma.2009.05.043
Elliott VL, Edge GT, Phelan MM, Lian LY, Webster R, Finn RF, Park BK, Kitteringham NR (2012) Evidence for metabolic cleavage of a PEGylated protein in vivo using multiple analytical methodologies. Mol Pharm 9(5):1291–1301. doi:10.1021/mp200587m
Foser S, Schacher A, Weyer KA, Brugger D, Dietel E, Marti S, Schreitmuller T (2003) Isolation, structural characterization, and antiviral activity of positional isomers of monopegylated interferon alpha-2a (PEGASYS). Protein Expr Purif 30(1):78–87
Lee H, Park TG (2003) A novel method for identifying PEGylation sites of protein using biotinylated PEG derivatives. J Pharm Sci 92(1):97–103. doi:10.1002/jps.10270
Lee KC, Moon SC, Park MO, Lee JT, Na DH, Yoo SD, Lee HS, DeLuca PP (1999) Isolation, characterization, and stability of positional isomers of mono-PEGylated salmon calcitonins. Pharm Res 16(6):813–818
Seyfried BK, Siekmann J, Belgacem O, Wenzel RJ, Turecek PL, Allmaier G (2010) MALDI linear TOF mass spectrometry of PEGylated (glyco)proteins. J Mass Spectrom 45(6):612–617. doi:10.1002/jms.1746
Yoo C, Suckau D, Sauerland V, Ronk M, Ma M (2009) Toward top-down determination of PEGylation site using MALDI in-source decay MS analysis. J Am Soc Mass Spectrom 20(2):326–333. doi:10.1016/j.jasms.2008.10.013
Li H, Rose MJ, Holder JR, Wright M, Miranda LP, James CA (2011) Direct quantitative analysis of a 20 kDa PEGylated human calcitonin gene peptide antagonist in cynomolgus monkey serum using in-source CID and UPLC-MS/MS. J Am Soc Mass Spectrom 22(9):1660–1667. doi:10.1007/s13361-011-0180-2
Fekete S, Ganzler K, Fekete J (2010) Simultaneous determination of polysorbate 20 and unbound polyethylene-glycol in protein solutions using new core-shell reversed phase column and condensation nucleation light scattering detection. J Chromatogr A 1217(40):6258–6266. doi:10.1016/j.chroma.2010.08.028
Tang L, Jariwala F, Bolgar M, Lloyd DK (2012) Separation and detection of bis-maleimide-polyethylene glycol and mono-maleimide-polyethylene glycol by reversed-phase high pressure liquid chromatography. J Chromatogr A 1246:117–122. doi:10.1016/j.chroma.2012.03.044
Bovey FA, Mirau PA (1996) NMR of polymers. Academic Press, San Diego
Han J, Danell RM, Patel JR, Gumerov DR, Scarlett CO, Speir JP, Parker CE, Rusyn I, Zeisel S, Borchers CH (2008) Towards high-throughput metabolomics using ultrahigh-field Fourier transform ion cyclotron resonance mass spectrometry. Metabolomics 4(2):128–140. doi:10.1007/s11306-008-0104-8
Hood BL, Zhou M, Chan KC, Lucas DA, Kim GJ, Issaq HJ, Veenstra TD, Conrads TP (2005) Investigation of the mouse serum proteome. J Proteome Res 4(5):1561–1568. doi:10.1021/pr050107r
Bruce SJ, Jonsson P, Antti H, Cloarec O, Trygg J, Marklund SL, Moritz T (2008) Evaluation of a protocol for metabolic profiling studies on human blood plasma by combined ultra-performance liquid chromatography/mass spectrometry: from extraction to data analysis. Anal Biochem 372(2):237–249. doi:10.1016/j.ab.2007.09.037
Huang L, Gough PC, Defelippis MR (2009) Characterization of poly(ethylene glycol) and PEGylated products by LC/MS with postcolumn addition of amines. Anal Chem 81(2):567–577. doi:10.1021/ac801711u
Stadalius MA, Gold HS, Snyder LR (1985) Optimization model for the gradient elution separation of peptide mixtures by reversed-phase high-performance liquid chromatography. Verification of band width relationships for acetonitrile-water mobile phases. J Chromatogr 327:27–45. doi:10.1016/s0021-9673(01)81636-8
Stadalius MA, Quarry MA, Snyder LR (1985) Optimization model for the gradient elution separation of peptide mixtures by reversed-phase high-performance liquid chromatography. Application to method development and the choice of column configuration. J Chromatogr 327:93–113. doi:10.1016/s0021-9673(01)81640-x
Acknowledgements
The authors would like to thank the Pharmaceutical Candidate Optimization—Metabolism and Pharmacokinetics group at Bristol–Myers Squibb for supplying samples. We thank Dr Petia A. Shipkova, Dr Dieter M. Drexler, Dr Purnima Khandelwal, Dr David K. Lloyd, and Dr Adrienne A. Tymiak for helpful discussions and critical review of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Warrack, B.M., Redding, B.P., Chen, G. et al. Determination of the molecular weight of poly(ethylene glycol) in biological samples by reversed-phase LC–MS with in-source fragmentation. Anal Bioanal Chem 405, 4283–4287 (2013). https://doi.org/10.1007/s00216-013-6795-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-013-6795-3