Abstract
Unraveling the complex, dynamic organization of the cell membrane can provide vital information about many aspects of cellular functions. Reported herein is a method for identifying cytoplasmic proteins that affect cell membrane protein organization. RNA interference (RNAi) is used to reduce the expression of select cytoplasmic proteins and a fluorescence resonance energy transfer (FRET) assay is used to measure changes in receptor microclustering. The advantage of this assay is that it does not require attaching fluorescent tags to the receptor. A change in energy transfer after reducing the expression of a cytoplasmic protein provides information about the protein’s role in altering receptor organization. As a demonstration of the method, cytoplasmic proteins involved in integrin microclustering have been identified. The cytoplasmic proteins targeted in this study include: dreadlock, integrin-linked kinase, paxillin, steamer duck, vinculin, rhea, focal adhesion kinase, and actin 42A. Reducing the expression of vinculin, paxillin, rhea, and focal adhesion kinase increased integrin microclustering, as measured by an increase in energy transfer in cells expressing αPS2CβPS integrins. No change in integrin microclustering was measured in a control cell line. Integrin mutants exhibited different microclustering properties compared to the wild-type integrins after reducing the expression of the listed cytoplasmic proteins. The results demonstrate the utility of this assay format, and provide insight into the function of cytoplasmic proteins in integrin microclustering.

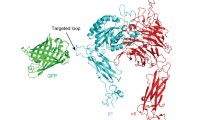
Schematic of a method for identifying cytoplasmic proteins that affect cell membrane protein organization. RNA interference is used to reduce the expression of select cytoplasmic proteins and fluorescence resonance energy transfer assay is used to measure changes in receptor microclustering




Similar content being viewed by others
Abbreviations
- FRET:
-
Fluorescence resonance energy transfer
- RNAi:
-
RNA interference
- GFP:
-
Green fluorescent protein
References
Giancotti FG, Ruoslahti E (1999) Science 285:1028
Phizicky EM, Fields S (1995) Microbiol Rev 59:94
Selvin PR (2000) Nat Struct Biol 7:730
Buensuceso C, de Virgilio M, Shattil SJ (2003) J Biol Chem 278:15217
Kim M, Carman CV, Yang W, Salas A, Springer TA (2004) J Cell Biol 167:1241
Smith EA, Bunch TA, Brower DL (2007) Anal Chem 79:3142
Ginsberg MH, Partridge A, Shattil SJ (2005) Curr Opin Cell Biol 17:509
Martin-Bermudo MD, Brown NH (1996) J Cell Biol 134:217
Helsten TL, Bunch TA, Kato H, Yamanouchi J, Choi SH, Jannuzi AL, Feral CC, Ginsberg MH, Brower DL, Shattil SJ (2008) Mol Biol Cell 19:3589
Subauste MC, Pertz O, Adamson ED, Turner CE, Junger S, Hahn KM (2004) J Cell Biol 165:371
Legate KR, Montanez E, Kudlacek O, Fassler R (2006) Nat Rev Mol Cell Biol 7:20
Vicente-Manzanares M, Choi CK, Horwitz AR (2009) J Cell Sci 122:199
Jannuzi AL, Bunch TA, Brabant MC, Miller SW, Mukai L, Zavortink M, Brower DL (2002) Mol Biol Cell 13:1352
Bunch TA, Grinblat Y, Goldstein LS (1988) Nucleic Acids Res 16:1043
Bunch TA, Brower DL (1992) Development 116:239
Clemens JC, Worby CA, Simonson-Leff N, Muda M, Maehama T, Hemmings BA, Dixon JE (2000) Proc Natl Acad Sci U S A 97:6499
March JC, Bentley WE (2006) Biotechnol Bioeng 95:645
Xia Z, Liu Y (2001) Biophysical J. 81:2395
Zal T, Gascoigne NRJ (2004) Biophys J 86:3923
Chen H, Puhl HL, Koushik SV, Vogel SS, Ikeda SR (2006) Biophysical J 91:L39
Zar J (1999) Biostatistical analysis. Prentice Hall, Upper Saddle River, NJ
Fogerty FJ, Fessler LI, Bunch TA, Yaron Y, Parker CG, Nelson RE, Brower DL, Gullberg D, Fessler JH (1994) Development 120:1747
Bunch TA, Helsten TL, Kendall TL, Shirahatti N, Mahadevan D, Shattil SJ, Brower DL (2006) J Biol Chem 281:5050
FLIGHT [http://flight.licr.org/]
PeptideAtlas [www.mop.unizh.ch/peptideatlas]
Brower D (2003) Curr Opin Cell Biol 15:607
Armknecht S, Boutros M, Kiger A, Nybakken K, Mathey-Prevot B, Perrimon N (2005) Methods Enzymol 392:55
Heidelberg [http://www.dkfz.de/signaling2/rnai/ernai_probes.php]
FlyBase, (http://flybase.bio.indiana.edu).
Kiger AA, Baum B, Jones S, Jones MR, Coulson A, Echeverri C, Perrimon N (2003) J Biol 2:27
Acknowledgments
This work was supported by the Roy J. Carver Charitable Trust (Muscatine IA), National Science Foundation (CHE-0845236) and Iowa State University Office of the Vice President for Research. The authors thank Roger Tsien (Howard Hughes Medical Institute, La Jolla, CA) for the original mCherry plasmid, Atsushi Miyawaki (Riken, Wako-city, Saitama, Japan) for the original Venus plasmid, Man-Yu Yum (Iowa State University) for help with the statistical analysis, Saiju Mathew (Iowa State University) for developing the FRET ImageJ plug-in, and Dilshan Shanaka Harischandra (Iowa State University) for synthesizing the RNA.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary materials
Below is the link to the electronic supplementary material.
ESM
(PDF 793 kb)
Rights and permissions
About this article
Cite this article
Dibya, D., Sander, S. & Smith, E.A. Identifying cytoplasmic proteins that affect receptor clustering using fluorescence resonance energy transfer and RNA interference. Anal Bioanal Chem 395, 2303–2311 (2009). https://doi.org/10.1007/s00216-009-3146-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-009-3146-5