Abstract
Animal microbiota is becoming an object of interest as a source of beneficial bacteria for commercial use. Moreover, the escalating problem of bacterial resistance to antibiotics is threatening animals and humans; therefore, in the last decade intensive search for alternative antimicrobials has been observed. In this study, lactic acid bacteria (LAB) were isolated from suckling and weaned pigs feces (376) and characterized to determine their functional properties and usability as pigs additives. Selection of the most promising LAB was made after each stage of research. Isolates were tested for their antimicrobial activity (376) and susceptibility to antibiotics (71). Selected LAB isolates (41) were tested for the production of organic acids, enzymatic activity, cell surface hydrophobicity and survival in gastrointestinal tract. Isolates selected for feed additive (5) were identified by MALDI-TOF mass spectrometry and partial sequence analysis of 16S rRNA gene, represented by Lentilactobacillus, Lacticaseibacillus (both previously classified as Lactobacillus) and Pediococcus genus. Feed additive prototype demonstrated high viability after lyophilization and during storage at 4 °C and − 20 °C for 30 days. Finally, feed additive was tested for survival in simulated alimentary tract of pigs, showing viability at the sufficient level to colonize the host. Studies are focused on obtaining beneficial strains of LAB with probiotic properties for pigs feed additive.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The health-promoting effects of microorganisms have been the subject of intense research in the recent years, the possibilities of their usage are increasingly described (Braune and Blaut 2016; Bautista-Gallego et al. 2017; Hidalgo-Cantabrana et al. 2018). In animal husbandry, there is a great interest in using probiotic microorganisms as an alternative to antibiotics (Patterson and Burkholder 2003; Cheng et al. 2014). These antimicrobials used as animal growth promoters were banned in 2006 in European Union countries due to the prevalence of antibiotic-resistant microorganisms and possibility of transmission of pathogens to humans (European Parliament and the Council of the European Union 2003; Long et al. 2018). This particular injunction has prompted an increased interest in searching for new health and nutritional additives for animals (Dowarah et al. 2017).
The welfare of farmed animals is one of the elements taken into account in the framework of European Union (EU) policy, whose main goal is to provide consumers with safe food, ensuring that the food chain has a neutral or positive effect on the environment. Standards and legal regulations developed within its framework cover all stages of production, from breeding to consumption. Meanwhile, among the main threats in the food production chain, which pose a risk to human and animal health and life, are pathogenic microorganisms, their metabolites and, on the other hand overused antimicrobial substances (European Commission 2020). Management of animal livestock in an artificial environment requires the prevention of disease spreading as well as rapid weight gain. Therefore, probiotic bacteria have been increasingly used as supplements in the nutrition of farm animals of various species, mainly pigs, poultry, cows, sheep, horses as well as fur animals (Hill et al. 2014; Yang et al. 2015; Dowarah et al. 2018a).
Probiotics are defined by FAO/WHO as “live microorganisms that, when administered in adequate amounts, confer a health benefit on the host” (FAO/WHO 2002). The mechanism of impact of probiotic strains on animals is multidirectional and complex. Probiotic microorganisms change the dynamics of the microbial population in the digestive system by balancing the number of beneficial and harmful microbiota (Mountzouris et al. 2009; Thomas and Versalovic 2010). It is related with the production of antimicrobial substances, such as organic acids or bacteriocins (Shim et al. 2012). Organic acids, in particular lactic and acetic acids, inhibit the growth of pathogens, and reduce pH value of the environment to a lethal level for pathogenic bacteria (Commane et al. 2005; Fayol-Messaoudi et al. 2005; Daşkiran et al. 2012; He et al. 2019). Bacteriocins and other antimicrobial metabolites, such as hydrogen peroxide, also reduce the growth of undesirable microorganisms. In addition, probiotics show the ability to adhere to the intestinal epithelium and they can interfere with the communication processes of pathogenic bacteria, e.g. through the quorum sensing mechanism, what prevents their colonization (Medellin-Peña et al. 2007; Mookiah et al. 2014). The most common change in the qualitative composition of the intestinal microbial population consists in an increase of the number of LAB and a simultaneous decrease in the number of coliform bacteria, including Escherichia coli (Cao et al. 2013; Landy and Kavyani 2013; Mookiah et al. 2014; Zhang and Kim 2014; Spaiser et al. 2015). Literature data also indicate the possibility of non-specific stimulation of the immune system of animals, which is another beneficial effect of the use of probiotic microorganisms (Schrezenmeir and De Vrese 2001; Yousefi et al. 2019). Among the functional properties of probiotic bacteria, recent studies indicate the anti-cancer effect of these microorganisms. It is related to the competition within the microbiome of the digestive tract of animals and the limitation of the growth of bacteria producing fecal enzymes with a carcinogenic effect (Rautray et al. 2011; Siva Kumar et al. 2015). The composition of the bacterial population in the gastrointestinal tract (GIT), supported by the use of probiotics, is often associated with increased productivity of animals, affecting more efficient digestion and absorption of nutrients and increasing immunity. Disease control is also observed (Niba et al. 2009; Hung et al. 2012; Barba-Vidal et al. 2019; Xin et al. 2020).
LAB are considered as the most effective probiotics in swine production (Dowarah et al. 2018b; Sirichokchatchawan et al. 2018; He et al. 2019). However, for using LAB as feed additives some challenges must be met, one of which is that bacterial species must be generally recognised as safe (GRAS), as well as microorganisms must remain viable at the appropriate level during: processing, transport, storage conditions and passage through the digestive system of the animal. Moreover, many scientists underline that bacteria isolated from the host are more effective probiotics than isolates obtained from other sources (Chiang et al. 2015; Bautista-Gallego et al. 2017; Dowarah et al. 2017). Therefore, in the presented study, beneficial microorganisms were isolated from swine feces. Taking into account differentiated microbiota depending on the animal age, bacteria were isolated from feces of suckling and weaned pigs.
Materials and methods
Sample collection
The feces of ten suckling and ten weaned healthy pigs were collected by a veterinarian from a swine farm in Wielkopolska Province, Poland. The health status of the animals was evaluated by the veterinarian on the basis of general condition of piglet and no impairment of health especially lack occurrence of digestive disorders, which were monitored. The samples were stored in sterile conditions and transported to the laboratory in coolers with ice and processed immediately on the same day.
Isolation of LAB
Piglets feces samples (10 g) were homogenized in stomacher BagMixer Interscience with 90 mL of sterile saline solution, serially diluted and cultured in MRS agar with the addition of 0.5% CaCO3 (Guo et al. 2010; Petsuriyawong and Khunajakr 2011), under microaerophilic conditions at 37 °C for 24 h (Torshizi et al. 2008; Adetoye et al. 2018). Randomly selected separated colonies were passaged onto MRS agar medium by reduction plating technique and incubated to obtain pure cultures. Individual colonies were propagated in MRS broth for further analysis. Isolated bacteria were stored as stocks in MRS broth at − 20 °C, with 80% glycerol in a 1:1 ratio (Adetoye et al. 2018). Every time before studies LAB were cultivated on MRS broth at 37 °C under microaerophilic conditions for 24 h.
Antibacterial activity
Antibacterial susceptibility for 376 LAB isolates was tested using modified well diffusion method (Vu et al. 2002; Aujoulat et al. 2011). The indicator bacteria used for the studies were Gram-positive strains: Clostridium perfringens ATCC® 13124™, Listeria monocytogenes ATCC® 19115™, Staphylococcus aureus ATCC® 33862™ and Gram-negative strains: Aeromonas hydrophila ATCC® 7966™, Campylobacter jejuni ATCC® 33291™, Enterobacter aerogenes PCM 532, Escherichia coli ATCC® 8739™, Proteus vulgaris PCM 542, Pseudomonas aeruginosa ATCC® 9027™, Salmonella enterica ser. Enteritidis ATCC® 13076™, Salmonella enterica ser. Typhimurium PCM 2565, Serratia marcescens PCM 549, Shigella flexneri PCM 89 and Yersinia enterocolitica ATCC® 9610™. Tested indicator bacteria were obtained from American Type Culture Collection (ATCC) and Polish Collection of Microorganisms, Institute of Immunology and Experimental Therapy PAN (PCM). Suspensions in physiological saline with an optical density of 0.5 on the McFarland scale were prepared from 24 h cultures of indicator bacteria. Cultures were carried out using the flood technique, after solidifying the agar wells were cut out with sterile cork borer (diameter 10 mm). Next, freshly cultivated LAB cultures in the amount of 0.1 mL were introduced into wells (MRS broth as a control). The incubation conditions were in accordance with the optimum temperature for the growth of indicator bacteria: 30 °C or 37 °C for 24 h. C. jejuni and C. perfringens were incubated under microaerophilic and anaerobic conditions, respectively. The diameter of growth inhibition was measured, taking into account the well (10 mm) (Chen et al. 2014; Gwiazdowski et al. 2015; Islam et al. 2020). Based on the diameter of the inhibition zones, three activity ranges were established for LAB isolates with specific antibacterial properties: > 20.1 mm—strong activity; 20.0–15.1 mm—moderate activity and < 15.1 mm—weak activity.
Antibacterial activity was re-evaluated for 41 LAB isolates, selected on the basis of antibiotic susceptibility assessment, after 18 months of storage in refrigeration conditions at − 20 °C, with 80% glycerol in a 1:1 ratio. For the in vitro studies additional indicator microorganisms were tested: A. hydrophila, S. flexneri, C. jejuni and P. aeruginosa.
Susceptibility to antibiotics
Antibiotic susceptibility of 71 LAB isolates was determined by the disk diffusion method using commercial antibiotic discs (Oxoid) (Torshizi et al. 2008; Pérez-Sánchez et al. 2011; Petsuriyawong and Khunajakr 2011). Ampicillin (10 mcg), vancomycin (30 mcg), gentamicin (10 mcg), kanamycin (30 mcg), streptomycin (300 mcg), erythromycin (15 mcg), clindamycin (2 mcg), tetracycline (30 mcg) and chloramphenicol (30 mcg) were used for the studies. As a criterion for the selection of antibiotics and their concentrations, data from the EFSA guidelines (European Food Safety Authority (EFSA) 2012) and thematic literature (Pérez-Sánchez et al. 2011; Ripamonti et al. 2011), as well as consultations with producers of feed additives for farm animals and breeders were used. Suspensions in saline solution were prepared from a 24 h LAB culture, setting the optical density at 1.0 on the McFarland scale. Samples were performed using the flood method, and after solidification of the medium, antibiotic discs were applied to the surface. Mueller–Hinton and MRS media were used in the study. Samples were incubated under anaerobic conditions with CO2 at 37 °C for 24 h. Controls were paper discs saturated with saline solution. The diameter of clear zones around the discs were measured, taking into account the size of the antibiotic discs (6 mm). The criteria for analysing the results of the sensitivity of microorganisms to the effects of antibiotics were determined using The Clinical and Laboratory Standards Institute (CLSI) M100 document (29th edition) (Clinical and Laboratory Standards Institute (CLSI) 2019), Charteris et al. (Charteris et al. 1998, 1999) and Han et al. (2015) dividing LAB to the following categories: resistant (R), intermediate susceptibility (I) and sensitive (S).
Susceptibility to antibiotics using the broth microdilution assay following the standard procedure of the ISO 10932|IDF 223:2010 (International Organization for Standardization (ISO) 2010) recommended by the EFSA (European Food Safety Authority (EFSA) 2012), was re-evaluated for five LAB isolates, selected on the basis of antibacterial activity, antibiotic susceptibility and functional properties. Minimum inhibitory concentration (MIC) of the following antibiotics: ampicillin (Sigma–Aldrich, Germany), gentamicin (Sigma–Aldrich, Germany), kanamycin (BioShop, Canada), streptomycin (Fisher Scientific, Belgium), erythromycin (ThermoFisher, Germany), clindamycin (ThermoFisher, Germany), tetracycline (Fisher Scientific, Belgium) and chloramphenicol (BioShop, Canada), was determined (European Food Safety Authority (EFSA) 2012). According to the guidance, determination of vancomycin microbiological cut-off values is not required for obligate heterofermentative Lactobacillus sp. (e.g. L. buchneri), L. paracasei and Pediococcus sp., therefore, it was not included in the research (European Food Safety Authority (EFSA) 2012). Each tested antibiotic was diluted into LSM medium under the appropriate concentration. An aliquot of 100 μL of twofold dilutions of tested antimicrobials in the concentration ranging from 0.008 to 128 μg/mL, depending on the antibiotic, were prepared in U-bottom 96-well microtiter plates. Each antibiotic was diluted into LAB susceptibility test medium (LSM) composed of Iso-Sensitest (Oxoid, Canada) broth and MRS broth (ratio 9:1, respectively) under appropriate concentration. The bacterial suspensions in the LSM from 24 h cultures were standardized to obtain density 1 McFarland’s standard which refers to the spectrophotometric equivalent of 3 × 108 CFU/mL, further diluted 1000-fold (Muñoz-Atienza et al. 2013) Next, the microplates were inoculated with the 100 μL of bacterial suspension and incubated at a temperature 37 °C for 24 h. The negative control was the sterile LSM medium with the addition of antibiotics, the positive control was the standardized bacterial inoculum without the inhibitory agent addition. Control strain used for the tests was E. coli ATCC 25922. After 24 h of incubation, the microorganisms' growth optical density was determined at 600 nm using BioTek Epoch 2 microplate reader. The MIC value was defined as the lowest antibiotic concentration, inhibiting the bacterial growth. The tested LAB results were further on interpreted based on the EFSA guidance (European Food Safety Authority (EFSA) 2012), distinguishing between sustainable and resistant strains.
Functional properties of LAB isolates
Production of selected organic acids
Determination of lactic, acetic, succinic and propionic acids content in 24 h bacterial cultures of 41 isolates on MRS broth was performed by high-performance liquid chromatography (HPLC) (Lim and Lee 2013; Gwiazdowski et al. 2015). The amount of 2 mL culture samples were centrifuged in sterile Eppendorf centrifuge tubes (10,000 rpm/min, 10 min) in a Centrifuge 5804R, Eppendorf AG. The supernatants were filtered through Millex®-LCR 0.22 Pm filters (Millipore). The tests were performed on a 2695 Waters liquid chromatograph coupled with a 2414 Refractive Index (RI) detector (Waters, Milford, MA, USA). Aminex HPX-87H 300 × 7.8 mm columns (BIO-RAD) were used for the determinations. The eluent was 0.004 M H2SO4 solution, with a flow of 0.75 mL/min. Solutions of lactic, acetic, succinic and propionic acids with a concentration of 0.1 µg/µL were used as standards. The determination was carried out at a temperature of 65 °C. Identification was made using an external standard method with the use of peak areas.
Enzymatic activity assay
The amount of 41 LAB isolates selected for further studies were tested for their amylase, protease and lipase activity based on the modified methods of Taheri et al. (2009), Guo et al.(2010) and Moslehishad et al. (2013). LAB were spot-inoculated on: modified MRS agar with 2% of starch for amylolytic activity, agar with 1% of skimmed milk for proteolytic activity and agar with Tween 80 and calcium chloride for lipolytic activity. Enzymatic properties were assessed on the basis of halo zones (amylolytic) and turbidity zones (lipolytic and proteolytic) around the colony growth after incubation under anaerobic conditions with CO2 at 37 °C for 24 h. To detect clear zones of amylase activity, Lugol’s solution was poured over the plates.
Cell surface hydrophobicity test
The cell surface hydrophobicity was determined for 41 LAB according to Taheri et al. (2009) and modified Pérez-Sánchez et al. (2011). LAB isolates were centrifuged for 5 min at 7500 rpm/min in a Centrifuge 5804R, Eppendorf AG. The supernatant was removed, the bacterial cells were washed twice with phosphate buffer and resuspended in physiological saline to obtain an optical density of 0.5 at 600 nm (OD600). The bacterial cell suspension in the amount of 3 mL was transferred into sterile tubes and 1 mL of toluene was added. Samples were shaken using a Vortex shaker for 90 s. The tubes were then allowed to stand for 15 min to separate the phases. Finally, the absorbance of the aqueous phase was measured at 600 nm. The hydrophobicity of the cells was calculated as the average percentage decrease in density of the LAB suspension (OD600) due to the adhesion of microorganisms to the hydrocarbon used.
Bile salts tolerance
The effect of bile salts on the survival of the 41 LAB isolates was tested according to a modified methods of Lin et al. (2007) and Guo et al. (2010) using microtiter plates. Twofold dilutions of 2% ox bile solution were prepared using MRS broth as diluent, followed by the addition of 100 µl bacterial suspension. The final concentration of ox bile in the cultures was 1.00%, 0.50% and 0.25%. Incubation was carried out under anaerobic conditions obtained using CO2 at 37 °C for 24 h. Spectrophotometric measurements were made using the BioTek Instruments EPOCH 2 microplate reader. Absorbance was measured at 600 nm (OD600) immediately after the samples were prepared and at 1, 3 and 24 h of incubation. The negative control was medium without LAB inoculum. A positive control was bacterial culture in MRS medium without the addition of ox bile. Strains that survived the bile salts exposure in > 50% after 24 h were considered as well tolerating the GIT conditions. In the range from 80 to 50% bile salts tolerance the LAB isolates were considered as well tolerating and above 80% as very well tolerating the GIT component.
Acidic pH tolerance
The impact of the environment pH on the survival of the 41 LAB isolates was tested with modified Guo et al. assay (Guo et al. 2010) using microtiter plates. Bacteria were incubated in the MRS medium with pH set at the levels 2 and 3, using hydrochloric acid (Pérez-Sánchez et al. 2011). Incubation was carried out under anaerobic conditions with CO2 at 37 °C for 24 h. Spectrophotometric measurements were made using the BioTek Instruments EPOCH 2 microplate reader. Absorbance was measured at 600 nm (OD600) immediately after the samples were prepared and after 1, 3 and 24 h of incubation. The negative control was medium without inoculum. The positive control was bacterial culture in MRS medium pH 6.5. Three tolerance ranges were established for LAB survival in 2.0 and 3.0 pH values after 24 h of incubation depending of percentage: > 50% viability—well tolerating isolates; 50–30% viability—moderate tolerating isolates and < 30% viability—weak tolerating isolates.
LAB identification
MALDI-TOF mass spectrometry
Selected, on the basis of obtained results, five LAB isolates were determined using MALDI-TOF Microflex mass spectrometry (Bruker, Germany) according to the standard producers protocol (Dec et al. 2014, 2016). The obtained LAB spectra have been identified using the data stored in BioTyper reference library of MALDI-TOF mass spectra and NCBI (The National Center for Biotechnology Information). Bruker MALDI-TOF BioTyper criteria for interpretation of the results: for the high-confidence of results, the value of the identification index must score ≥ 2. The range from 1.99 to 1.70 indicates low-confidence identification, the value lower than 1.70 equals lack of microorganism identification.
Genetic identification
Identification of five selected LAB isolates was carried out based on the partial sequence analysis of the 16S rRNA gene, which required the isolation of genomic DNA, PCR with appropriately selected primers, sequencing and the analysis of the obtained sequences. 16S rRNA genes of the selected LAB strains were amplified using primers 1492r (ggT TAC CTT gTT ACg ACT T) and S-D-Bact-0008 (AgA gTT TgA TCM Tgg CTC AG) (Leser et al. 2002; Pang et al. 2011). The 1500 bases sequences were edited, combined and generated using the GeneDoc 2.700. Obtained sequences were analyzed using the BLAST (Megablast algorithm) Tool (https://blast.ncbi.nlm.nih.gov/) and submitted in GenBank. The unrooted phylogenetic tree was constructed to determine the closest LAB species by the neighbor-joining method (Saitou and Imanishi 1989) using the MEGA X software (Kumar et al. 2018). For the construction of the phylogenetic tree, 16S rRNA sequences of L. paracasei (GeneBank ID: MZ411515.1, M411523.1, MZ411532.1, NR_025880.1), L. paracasei subsp. paracasei (GeneBank ID: LC096209.1), L. paracasei subsp. tolerans (GeneBank ID: LC065035.1), P. pentosaceus (GeneBank ID: MW025983.1, KX886792.1), L. buchneri (GeneBank ID: MW025972.1, NR_041293.1) and E. coli (GeneBank ID: X80725) were obtained from NCBI database.
Feed additive preparation
Preparation of freeze-dried bacterial cultures
Lyophilization of selected bacterial isolates cultured on a skim milk carrier was carried out on a medium enriched with maltodextrin and trehalose (Meng et al. 2008; Strasser et al. 2009; Lo Verso et al. 2018). Experimental work was performed using a shelf freeze dryer, Alpha 1 D series. Samples of 24 h cultures of each isolate separately, in the amount of 20 mL, were transferred to sterile disposable containers of polypropylene with a capacity of 60 mL. The test samples were then kept in the freezer at − 20 °C for 72 h, followed by lyophilization. The process of vacuum freeze drying took place at constant parameters: pressure 63 Pa, time 96 h, temperature of the heating shelves of the freeze dryer 30 °C.
Assessment of the stability of freeze-dried bacterial cultures and additive during storage
Samples of five lyophilized bacterial cultures were grounded and mixed thoroughly in equal proportions, giving the prototype of feed additive. The stability of five freeze-dried bacterial cultures and the feed additive was determined by a plating method comparing the number of bacteria immediately after freeze drying and after 30 days of storage at 4 °C and − 22 °C. Samples were kept in sealed, sterile containers to assess storage stability (Fonseca et al. 2006, 2015a; Strasser et al. 2009).
Evaluation of the survival of LAB present in feed additive in the simulated digestive system
The survival of LAB strains in a simulated digestive system was determined using in vitro pig model. The survival of bacteria was evaluated by transferring test samples: (A) lyophilized mix of five LAB cultures contained addiction of feed and (B) lyophilized mix of five LAB cultures, through the simulated GIT. In the experiments, commercial feed was used—a universal protein concentrate for pigs during the fattening period. This mix is made on the basis of, among others, cereals, high-protein extraction shots, vitamin and mineral premix, mineral additives as well as biologically active compounds and other components. Sample A was 5 g of protein concentrate with 10% of the freeze-dried LAB, sample B was 5 g of the freeze-dried LAB. The in vitro digestion model included a three-stage digestion process: in the mouth (simulated saliva solution: sodium chloride, potassium chloride, calcium chloride, sodium bicarbonate, α-amylase, water) (Avantaggiato et al. 2003; Versantvoort et al. 2005), stomach (0.1 M phosphate buffer; 0.2 M hydrochloric acid solution, water pepsin 25 mg/cm3 solution) and small intestine (water pancreatin 100 mg/cm3 solution, 0.6 M sodium hydroxide solution, 0.2 M phosphate buffer) (Boisen and Fernández 1997). Samples A and B were pre-digested at 39 °C in simulated solution of saliva for 5 min, then in simulated gastric juice for 2 h and intestinal juice for 4 h. At each stage, the test mixture was incubated at 39 °C under continuous mixing at 150 rpm on a rotary shaker to ensure homogeneous distribution of all components (Boisen and Fernández 1997; Avantaggiato et al. 2003; Versantvoort et al. 2005). At the beginning of each section of the GIT and after the retention time, the viability of LAB was determined.
Statistical analysis
Statistical analysis of the results was carried out using SPSS Statistics 23 and Microsoft Excel®. All the experiments were carried out in three parallel replicates. The standard deviation value was calculated for the obtained results. One-way analysis of variance (ANOVA) was performed to compare the mean values. The significance of differences between the mean values of the results was determined by the Tukey post hoc test. All statistical hypotheses were verified at the significance level p < 0.05.
Results
Isolation of LAB
A total of 244 microorganisms were isolated from the feces of suckling piglets and 132 from weaned piglets. All isolates were preliminary identified as LAB based on the criteria of being Gram-positive and catalase negative rods and cocci, hydrolysing CaCO3 on the MRS agar plates. The number of LAB in tested feces samples ranged from 7.08 to 9.85 log CFU/g from suckling piglets and from 8.46 up to 9.99 log CFU/g from weaned piglets.
Antibacterial activity
Antibacterial activity was assessed towards chosen pathogens of the intestinal tract of pigs, human pathogens and some potentially pathogenic microorganisms. The results of in vitro experiments showed a large diversity of the 376 LAB isolates antibacterial effect, both taking into account their activity spectrum and the degree of inhibition growth of indicator microorganisms (Table S1 in Supplementary material). Out of all the tested bacterial isolates, 87 did not exhibit any antagonistic properties against the indicator microorganisms. Most isolates that showed antibacterial activity were characterized by a weak interaction (8.24–21.01% depending on the indicator strain). Only a small group of isolates was characterized by moderate (0.27–11.17%) and strong (0.00–4.52%) activity towards indicator microorganisms. Based on the results of antibacterial activity, 71 LAB isolates with the strongest antimicrobial effect towards different indicator bacteria were selected for further tests.
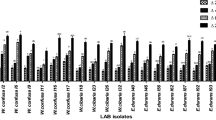
After 18 months of storage, 41 LAB isolates, selected on the basis of susceptibility to antibiotics, were redefined for maintaining their antibacterial activity towards an extended group of indicator bacteria (Fig. 1). LAB isolates showed a similar antagonistic effect against indicator bacteria, compared to the results obtained in previous antimicrobial tests. Only in the case of antagonistic effect towards C. perfringens, after a storage period, a significant reduction in the activity of most LAB isolates was observed. Out of the 41 tested isolates, only 2 of them inhibited the growth of C. perfringens to a small extent. The results of the experiments also showed a diverse antimicrobial effect of the tested LAB, taking into account both their activity spectrum and the degree of growth inhibition. Furthermore, 4 indicator microorganisms were added for antibacterial evaluation. A. hydrophila and P. aeruginosa were susceptible to 40 out of 41 tested isolates, with growth inhibition ranging from low to high. All tested isolates showed antagonistic activity against S. flexneri, where the values of growth inhibition zones ranged from low impact to moderate. Inhibition of C. jejuni growth was also observed by 30 tested LAB, where high antimicrobial activity was observed in several isolates. In summary, the conducted research indicates the ability of LAB to inhibit the growth of A. hydrophila, S. flexneri, C. jejuni and P. aeruginosa. In most cases, the antibacterial activity of the tested isolates was maintained after 18 months, although, as mentioned, a reduction, disappearance and even increase in the activity was observed in relation to selected indicator microorganisms. Detailed results concerning antibacterial activity are presented for five finally chosen LAB isolates in Table 3.
Susceptibility to antibiotics
The assessment of susceptibility to nine antibiotics was the next step of LAB selection. Using the antibiogram method to determine the drug resistance of LAB, the following levels of action were specified: sensitive isolate, intermediate sensitive isolate or resistant isolate, depending on the size of the growth inhibition zones. The sensitivity of selected 71 LAB isolates to chosen antibiotics was varied (Table S2 in Supplementary material). All the tested isolates were vulnerable to the effects of clindamycin and erythromycin (in the tested concentrations), except for one isolate derived from the feces of suckling piglets, which showed medium sensitivity to erythromycin. Most isolates were also sensitive to chloramphenicol as only one isolate showed moderate sensitivity. Among LAB, five showed intermediate susceptibility towards tetracycline and one resistance. 52 isolates showed high sensitivity to 10 mcg ampicillin, while 18 isolates—medium sensitivity, as well as lack of sensitivity was observed for 1 isolate. The number of LAB isolates showing high or medium sensitivity to streptomycin at 300 mcg was 67. In case of gentamicin, a wide variation of isolates sensitivity to this antibiotic was observed. 50 LAB isolates were sensitive, including 12 vulnerability, while 20 isolates were insensitive to gentamicin at the tested concentration. The highest number of resistant isolates was found for kanamycin (33) and vancomycin (69) at 30 mcg. Based on the obtained results, 41 LAB isolates, which had no or low acceptable antibiotic resistance, were selected for further studies. Among the bacteria selected after this stage, 29 isolates came from the digestive tract of suckling piglets and 12 isolates from the weaned piglets.
Production of selected organic acids
The selected 41 LAB were characterized by a diverse content of lactic acid in culture, which was the dominant product and ranged from 7.67 to 40.68 mg/mL. Among LAB isolates, 17 had a high level of lactic acid production, in the range from 26.31 to 40.68 mg/mL. In cultures of 22 isolates, the amount of lactic acid was at an average level, from 15.04 to 24.62 mg/mL, and in 2 cases the content of this metabolite was low and ranged from 7.67 to 12.76 mg/mL. A small amount of acetic acid was found in the LAB cultures, the concentration was between 2.59 and 4.79 mg/mL. It should be emphasized that the acetic acid content varied significantly in most cases. Tested LAB isolates did not produce propionic and succinic acids.
Functional properties of LAB
Enzymatic activities
Enzymatic activity tests included proteolytic, amylolytic and lipolytic activity (Table 1). The results of the studies showed that all tested LAB isolates caused the degradation of milk casein (clear zones around the growth). Among tested LAB, 46% showed starch degradation (clear zone after flooding with Lugol's liquid). Amylolytic properties were noted for 13 isolates derived from the suckling piglets feces and 6 isolates isolated from the weaned piglets feces. When assessing the ability of the tested LAB to produce lipases, no halo zones were found around the bacterial colonies, which means that LAB isolates did not exhibit lipolytic properties.
Cell surface hydrophobicity test
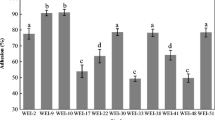
The ability of LAB cells to adhere to hydrocarbons as an indicator of their hydrophobicity, and also the criterion of the ability of adhesion to intestinal epithelial cells was tested. The results are expressed in the form of percentage changes in the density of the bacterial cell suspension after adhesion to the hydrocarbon used. Above 40% of the value of toluene adhesion was assumed as a high degree of cell hydrophobicity, ranging from 20% to 40%, as medium, while low level, below 20%, was determined (Table 1). The results regarding the assessment of the hydrophobicity of LAB derived from piglets feces indicate a large variation in this parameter. Out of the 41 tested isolates, 7 were characterized by high and 11—by medium hydrophobicity values from 22% to 63%. The remaining 22 isolates had a low toluene adhesion value, no more than 10%.
Survival of LAB in the GIT conditions: bile salts and acidic pH tolerance
The presence of bile salts in the samples had a different effect on the growth of the tested isolates (Table 1). Detailed data concerning selected strains are presented in Table 5 and on Fig. 3. Tolerance ranges presented in the Table 1 are established by the Authors based on the thematic literature data (García-Hernández et al. 2016) as well as obtained results. Bile salts tolerance, expressed as a percentage of population growth in the presence of this component relative to the control sample, depended on the tested LAB isolate as well as the concentration of ox bile salts in the culture. As a rule, the increase in bile salt content caused a decrease in bacterial survival rate; however, for many isolates, there were no significant differences in growth in the presence of 0.5 and 1%. Several isolates (e.g. KK 033) showed very high tolerance to the presence of bile salts, resulting in intense growth comparable to bacterial growth in control samples. Significant growth inhibition of some LAB (e.g. DG 068) was also noted. Similar relationships were observed in the group of isolates derived from the suckling and weaned piglets feces. For some isolates, the effect of bile salts concentration on population growth was visible in the first hours. The growth of most isolates remained at a similar level during the first hours of incubation, regardless of concentration, while differences in growth were only revealed in the following hours. Isolates that showed low tolerance of this component in the medium were characterized by weak growth during the first hours.
Taking into account the pH range prevailing in the stomach of various monogastric animals, the survival of the tested LAB isolates was tested in MRS medium, whose pH was set at 2.0 and 3.0. For comparison, cultures were run in parallel under optimal conditions at a pH of 6.5. It was found that the OD600 value after 24 h incubation for all isolates was significantly lower in the medium at pH 2.0 and 3.0 compared to the control value. However, a diverse tolerance of the tested LAB to low pH values was found (Table 1). Statistical analysis showed significant differences between isolates. Comparing bacterial growth at pH 2 and pH 3, in a few cases significant differences in OD600 values can be seen, which illustrates the growth of isolates under these conditions. The weaker growth was also observed at pH 2 compared to pH 3.
Identification of selected LAB isolates
The conducted research including production of organic acids, enzymatic activity, cell surface hydrophobicity as well as bile salts and acidic pH tolerance, allowed the selection of five LAB isolates demonstrating high antimicrobial activity, sensitivity to antibiotics, high production values of lactic acid as well as good survival in GIT conditions. Moreover, the obtained results of five selected LAB isolates were meant to complement each other in terms of the examined features. Selected LAB isolates were identified by MALDI-TOF mass spectrometry and sequencing of the 16S rRNA gene. The partial sequencing results obtained using two complementary primers 1492r and S-D-Bact-0008, after initial editing in GeneDoc 2.7.000, were combined. Sequence similarity searching was carried out using the BLAST algorithm. Results covering LAB genus and identification value are compiled in Table 2. In the group of LAB derived from the suckling piglets feces, isolates belonged to the following species Lacticaseibacillus paracasei, Lentilactobacillus buchneri and Pediococcus pentosaceus. One selected isolate obtained from weaned piglets showed homology to L. paracasei. The data restrictions of GeneBank were observed for three strains which have been identified as L. paracasei species, with differences on subspecies level identification. Query coverage of the presented LAB identification results obtained using BLAST was 99–100%. The combined partial sequences of isolates have been deposited in the GenBank.
For LAB strains identification and specific species confirmation, molecular phylogeny analysis was performed, the phylogenetic tree was constructed based on 16S rRNA sequences using by neighbor-joining method (no. of bootstrap replications = 2000) (Fig. 2) (Felsenstein 1985; Saitou and Imanishi 1989; Tamura et al. 2004; Kumar et al. 2018). Following the phylogenetic analysis, LAB strains KK 008, KK 033, and KK 160 were placed in the cluster making up the Lacticaseibacillus genus, subgroup L. paracasei. The strain DG 068 was placed in the Lentilactobacillus cluster, L. buchneri subgroup. The strain DG 059 was positioned in the Pediococcus genus cluster, subgroup P. pentosaceus. The conducted phylogenetic analysis also confirmed that the results of identification studies for the isolated LAB strains were correct.
Characteristics of five finally selected LAB strains
The results of laboratory experiments carried out to the point of feed additive composition, obtained for five selected bacterial strains, are summarized in Tables 3, 4, 5, as well as on Fig. 3. The feed additive developed for pigs supplementation included the following bacteria isolates: L. paracasei KK 008, L. paracasei KK 033, L. paracasei KK 160, P. pentosaceus DG 059 and L. buchneri DG 068. Selected isolates showed a wide range of antibacterial activity, which was tested twice: first, as the bacteria were isolated and identified as LAB and second after 18 months of storage. Differences in demonstrated antagonistic activity between the months have been observed. Both weaker antibacterial activity or its loss as well as gaining and greater antagonistic properties were spotted. None of five selected LAB isolates inhibited the growth of Y. enterocolitica. Furthermore, LAB isolates after 18 months of storage were tested positive for the inhibition of A. hydrophila, C. jejuni, P. aeruginosa and S. fexneri.
Antibiotic susceptibility of five selected LAB strains was determined both by disc diffusion technique as well as by microdilution method. On the basis of the disc diffusion assay results, it can be concluded that selected LAB strains showed high or medium sensitivity to eight out of nine tested antibiotics in tested concentrations. All selected LAB isolates were sensitive to erythromycin, clindamycin, tetracycline and chloramphenicol in tested concentrations. Differences in the results of LAB isolates susceptibility to tested antibiotics were observed for ampicillin, gentamycin and streptomycin. L. paracasei isolates no. KK 008 and KK 033 were susceptible to these three antibiotics. P. pentosaceus DG 059 exhibited susceptibility towards ampicillin and intermediate susceptibility for the gentamycin and streptomycin in tested discs concentrations. L. buchneri DG 068 was susceptible to gentamycin and intermediate susceptible to ampicillin and streptomycin in tested concentrations. Finally, L. paracasei KK 160 demonstrated susceptibility towards ampicillin and streptomycin whereas towards gentamycin these bacteria were intermediate susceptible. Kanamycin (30 mcg) was the only antibiotic for which an intermediate susceptibility of the five LAB isolates was observed. All of the selected isolates were resistant to vancomycin (30 mcg). The results of antibiotics MIC values confirmed previously obtained results. According to the results, all five tested LAB strains were susceptible to ampicillin, gentamicin, kanamycin, streptomycin, erythromycin, clindamycin, tetracycline and chloramphenicol. The MIC values of all five selected LAB strains were lower than the EFSA breakpoints (Table 4). In the EFSA guidelines, cut-off values for L. buchneri are not discussed individually, therefore, the respective MIC values were interpreted using breakpoints given for the obligate fermentative Lactobacillus bacteria (European Food Safety Authority (EFSA) 2012).
Selected LAB strains showed proteolytic activity, while only two of them: L. paracasei KK 008 and KK 160 demonstrated amylolytic activity. None of the five selected LAB exhibited lipolytic properties. Results concerning the production of dominant organic acid, cells hydrophobicity and survival of selected LAB in the GIT conditions are presented in Table 5. Lactic acid production by the studied bacteria ranged from 15.15 to 31.69 mg/mL, depending on the strain. The possibility of adhesion to the epithelium was estimated based on the assessment of the hydrophobicity of cells from 22.44 to 41.61%. Selected isolates were characterized by good tolerance of low medium pH: 2.0–3.0 and bile salts concentration: 0.25–1.0%.
The results of in vitro LAB survival in the GIT conditions demonstrating tolerance of acidic pH and ox bile, are presented on Fig. 3. Growth rate in unfavorable conditions was observed between 3 and 24th h. All the finally selected LAB showed the capacity to survive the tested values of acidic pH: however, their growth was strongly reduced. In case of four out of five LAB, their viability degree was better in pH 3 compared to pH 2. Isolates no. KK 008, KK 033 and KK 160 showed good tolerance of all tested ox bile concentrations, while isolate no. DG 059 only in concentration 0.25%. L. buchneri DG 068 was the only vulnerable isolate to the tested concentrations of bile salts.
Preparation and stability of feed additive
Preparation of feed additive using selected LAB isolates
Selected LAB isolates, with potentially probiotic properties whose biological activity could complement each other were used for the composition of feed additive prototype for pigs (Fig. 4). Selected LAB strains were cultivated on a medium with milk enriched with trehalose and maltodextrin and freeze-dried. The largest decrease in the population size was observed for L. buchneri DG 068 samples (0.34 log CFU/mL). In the remaining samples, the difference was small and ranged from 0.08 to 0.23 log units. The exception was the population of P. pentosaceus DG 059, which increased after the fixation process (9.5–9.66 log CFU/mL). The tested LAB showed a high survival rate during freeze-drying, which confirms a proper selection of the medium.
Growth rate of selected LAB strains at each stage of feed additive formulation: KK 008 L. paracasei; KK 033, L. paracasei; DG 059, P. pentosaceus; DG 068, L. buchneri; KK 160, L. paracasei; feed additive, freeze-dried bacterial cultures of KK 008, KK 033, DG 059, DG 068 and KK 160, ground and mixed thoroughly in equal proportions. Data were analysed by Tukey’s test at p < 0.05 (a, b means marked with different letters in bars differ significantly)
The stability of freeze-dried LAB cultures and additive during storage
An important parameter determining the technological usefulness of microorganisms is the stability of the population during storage. The freeze-dried culture samples of selected five LAB, as well as the prepared feed additive, were stored at 4 °C and – 20 °C for 30 days. Next, the number of LAB population was determined. The number of LAB in the mixed feed additive remained stable during storage in different conditions, while in LAB samples statistically significant differences were observed. In all the samples stored at 4 °C the number of LAB was significantly lower than in freeze-dried samples. Among samples stored at − 20 °C, the decrease of population was observed only for L. paracasei KK 033 and KK 160.
Survival of LAB present in feed additive in the simulated digestive system
Composed prototype of feed additive for pigs has been passaged through the in vitro digestive tract. The experimental conditions were similar to those prevailing in the GIT of monogastric animals. Samples of freeze-dried bacterial cultures of KK 008, KK 033, DG 059, DG 068 and KK 160, ground and mixed thoroughly in equal proportions with commercial feed addition (A) and without the feed (B) were transferred through solutions simulating: saliva, gastric juice and intestinal juice (Fig. 5). The initial number of bacteria tested in samples differed by one logarithmic unit, due to the fact that sample A (8.38 ± 0.01 log CFU) contained addiction of feed, while sample B (9.28 ± 0.03 log CFU) was freeze-dried LAB. After a few minutes of simulated solutions of saliva and gastric juice impact, the number of LAB decreased by 4 and 3 log units (sample A and B, respectively). After two hours of incubation in solution simulating gastric juice, a further decrease in cell population was observed in both samples, by 1 and 3 log units (sample A and B, respectively). After changing the environment to mixture simulating intestinal juice, the number of bacteria equaled in both samples. An increase in LAB population in sample B was observed, while in the sample A this parameter remained at a comparable level. After three more hours of incubation in solution simulating intestinal juice, a significant increase in the number of bacteria was observed in both samples.
Number of LAB during the passage in the in vitro digestive tract of pigs. Five selected and freeze-dried LAB: KK 008, L. paracasei; KK 033, L. paracasei; DG 059, P. pentosaceus; DG 068, L. buchneri; KK 160, L. paracasei. Data were analysed by Tukey’s test at p < 0.05 (a–e means marked with different letters differ significantly)
Discussion
LAB are the largest and best characterized group of probiotics. Their beneficial effect is associated with the ability to inhibit pathogens growth, influence the balance of gastrointestinal microbiota and support the immune system. Despite a lot of research concerning probiotics, there is a need to search for new strains as probiotic properties are strain-specific (Casarotti et al. 2017) and many authors suggest linking the origin of the strain and its intended use (Chiang et al. 2015; Bautista-Gallego et al. 2017; Dowarah et al. 2017). In the presented work, evaluation of functional properties of LAB strains isolated from pig feces obtained from suckling and weaned pigs has been carried out according to guideline recommended by FAO/WHO (2002) together with a construction of feed additive prototype. Out of the initial number of 376 isolates, five strains which demonstrated the best properties (including broad range of antimicrobial activity, high production values of organic acids, good survival under GI conditions as well as cells hydrophobicity) have been chosen. The research covered the sequence of experiments with antimicrobial activity as a first criterion. Out of 376 isolates, 87 demonstrated no antibacterial properties. Similar results were reported by other authors who observed up to half of isolates without activity during their screening tests (Guo et al. 2010; Lo Verso et al. 2018). Taking into account that only a part of isolates inhibited the growth of pathogenic bacteria, this step enabled selection of the most promising LAB isolate, tested next for their antibiotic sensitivity. According to EFSA, studies on antibiotic resistance profiles and transmission mechanisms are necessary to give QPS status to probiotic strains intended for industrial use (European Food Safety Authority Panel on Biological Hazards (BIOHAZ) 2013). In the presented study high susceptibility of almost all the tested LAB isolates to chloramphenicol, clindamycin, erythromycin and tetracycline was detected. Over 90% of isolates were highly or intermediate sensitive to streptomycin and ampicillin, more than 70% were sensitive to gentamicin, while high and moderate susceptibility to kanamycin was demonstrated only by 16.44% and 38.36%, respectively. Only two isolates showed sensitivity to vancomycin. These results are consistent with literature data. Moreover, re-evaluation of five finally selected LAB isolates using broth microdilution assay confirmed their susceptibility towards ampicillin, gentamicin, kanamycin, streptomycin, erythromycin, clindamycin, tetracycline and chloramphenicol on the basis of EFSA FEEDAP Panel guidance (European Food Safety Authority (EFSA) 2012). In the present study for microdilution antibiotic testing, LSM was used to avoid the antibiotic-inhibitory activity by the medium components, observed for MRS (Huys et al. 2002; Klare et al. 2005). LAB to grow require carbohydrates, vitamins, minerals and amino acids, so the usage of Mueller Hinton or Iso-Sensitest media is inadequate. Therefore, for nonenterococcal LAB, the combination of Iso-Sensitest and MRS media was obtained by Klare et al. (2005) and further on become a reference medium for microdilution antibiotics tests (International Organization for Standardization (ISO) 2010). In the research of de Souza et al. (2019) all the tested LAB strains were susceptible to ampicillin, erythromycin, clindamycin, tetracycline and chloramphenicol, while all LAB strains were resistant to vancomycin and 85% of strains were resistant to kanamycin. Similarly, Birri et al. (2012) reported that all the tested L. casei strains were resistant to vancomycin and susceptible to erythromycin, chloramphenicol and tetracycline. In the study of Jeronymo-Ceneviva et al. (2014) 100% of LAB strains isolated from cheese showed resistance to vancomycin. In turn, Casarotti et al. (2017) reported that all the L. delbrueckii subsp. bulgaricus strains were susceptible to vancomycin and gentamicin, whereas other strains, belonging to L. rhamnosus, L. casei and L. fermentum species, were resistant. The resistance of LAB to vancomycin, according to some authors, will not be transferable to pathogenic bacteria as it is an intrinsic feature, codified by chromosomic genes (Divya et al. 2012) and the replacement of terminal d-alanine by d-lactate or d-serine in the muramyl-pentapeptide prevents vancomycin binding (Gueimonde et al. 2013). As the antibiotic sensitivity was an important criterion, it allowed to choose 41 isolates for further characterization including organic acids production, enzymatic activity, hydrophobicity as well as tolerance of acid and bile salts.
Determination of the content of organic acids, such as lactic and acetic acids, which are the main metabolites of LAB, was, on the one hand, confirmation that the tested isolates belong to this group of microorganisms, and, on the other hand was one of the selection factors. Moreover, propionic acid content was also determined, having considered that some heterofermentative or optionally heterofermentative LAB may produce it (Ripamonti et al. 2011). High level of lactic acid production is a desirable feature for commercial-use strains selection, as their functional properties, e.g. antimicrobial activity, are dependent, among others, on the ability of bacteria to acidify their habitat. Although differentiated mechanisms may play a role in the inhibition of pathogens, such as competitive exclusion and production of antimicrobial substances including bacteriocins or hydrogen peroxide, organic acids are important metabolites involved in the antagonistic activity (Niku-Paavola et al. 1999; Arena et al. 2016; Sirichokchatchawan et al. 2018).
The functional characteristics of LAB isolates included the enzymatic properties as a particularly desirable feature in supplements intended for animals, supporting digest feed (Lee et al. 2001). Many authors emphasize the desirability of testing microorganisms in terms of determining enzymatic properties during the selection of strains for animal feed supplementation. In the presented study, rather weak enzymatic activity of LAB isolates has been stated. None of the tested bacteria showed lipolytic activity and some of them showed only weak proteolytic and amylolytic properties. Similar amylolytic activity results were obtained by Guo et al. (2010), where only 3 LAB isolates out of 150 tested showed high capacity of starch digestion. Literature data indicate that the enzymatic properties of LAB are very diverse. Similar results were obtained by Kim et al. (2007) who tested 252 LAB isolates derived from faeces and intestines of piglets in terms of their ability to produce proteases, amylases, lipases and phytases. 81% of the strains tested had proteolytic properties, 22% of isolates derived from feces and 23% obtained from intestines had amylolytic properties. However, the authors obtained different results for lipolytic properties, which they found in 15% of faeces isolates and 17% obtained from intestines. Differences in the occurrence of certain enzymatic properties among LAB may result, among others, from their origin. For example, authors describe strains that have strong amylolytic properties that come from fermented starchy foods (Sanni et al. 2002). Another problem affecting the biochemical properties of bacteria may be the degree of binding of enzymes to the cell. (Lee et al. 2001) during selection of potentially probiotic bacteria found that the amylolytic activity of L. acidophilus L23 was closely related to the presence of cells, which indicates that the enzyme was associated with the cell wall. In turn, another strain, L. fermentum L9, produced extracellular amylolytic enzymes.
Among the most important parameters determining the usefulness of LAB as potential probiotics, the hydrophobicity as well as tolerance of low pH and bile salts should be taken into account. Cell surface hydrophobicity affects the overall adhesion capacity and can facilitate the contact between bacterial and host epithelial cells (Kos et al. 2003; Sánchez-Ortiz et al. 2015). According to literature data, high hydrophobicity indicates that bacteria can bind better to the intestinal mucosa (Todorov et al. 2011; de Souza et al. 2019), therefore, the adhesion to hydrocarbons is used as a biochemical marker allowing to evaluate this feature (Gandomi et al. 2019). In the presented work various hydrophobicity among tested LAB isolates was observed with the dominance of isolates demonstrating low degree of adhesion to toluene and only 7 out of 41 isolates showing high hydrophobicity. These results are consistent with data reported by other authors. (Angmo et al. 2016) have stated extremely variable results of tests provided for 25 isolates, where only 3 demonstrated high hydrophobicity. Generally low degree of hydrophobicity with 5 out of 19 strains obtained from Mozzarella cheese showed values around 60% observed (de Souza et al. 2019). In turn, (Jeronymo-Ceneviva et al. 2014) noticed high hydrophobicity value for all four tested LAB strains isolated from water-buffalo Mozzarella cheese. LAB isolates tolerance to ox bile and acidic pH was studied to predict bacterial survival after oral administration. Extreme conditions of the medium set at the 2 and 3 pH as well as 1.00%, 0.50% and 0.25% ox bile concentration were used to reflect stomach prevailing conditions (Lin et al. 2007; Guo et al. 2010; Pérez-Sánchez et al. 2011b). Guo et al.(2010) reported that, piglets originating, LAB isolates losses the viability due to the high acidity of the medium as well as tested bile concentrations. This phenomenon was also observed in our studies in case of a few isolates. However, in this paper, the majority of the tested LAB isolates revealed tolerance to acidic pH and tested ox bile concentrations for 3 h, no significant decrease of viable counts was observed, even bacterial growth was detected after 24 h. All tested by García-Hernández et al. LAB isolates from broiler chicken showed resistance to acidic environment (pH 2.5), although bacterial growth was depressed. Bile salts concentration of 0.3% occurred in obtaining very different results of LAB survival after 3 h of exposure and bacterial final growth after 12 h of incubation. The strain LB-31 showed the highest value of 82.13% for initial survival, but the lowest relative final growth. Obtained by several studies results suggest that the acid tolerance of LAB isolates was strain-specific (Mishra and Prasad 2005; Guo et al. 2010).
Freezing with cryoprotectants at low temperatures is a common method applied to preserve the viability and properties of LAB during long-term storage. One of the most effective cryoprotectant, which is widely used in frozen concentrates, is glycerol. Although glycerol prevents bacteria from freezing damage, their viability after storage decreases. The mechanisms of bacterial freezing damage are not completely understood. Mainly low freezing temperature, ice crystal formation as well as glass transitions are causing cellular damage due to occurring cell stresses including cold, osmotic, mechanical as well as oxidant stress. During freezing, damages of membrane lipids, cell proteins and DNA is observed, leading to cell functionality losses and finally loss of viability (Fonseca et al. 2006, 2015b; Smirnova et al. 2019). The bacterial viability as well as their biological activity after freezing and frozen storage depends on: the strain, culture conditions, cryoprotectants and their formulations, freezing conditions and storage time (Fonseca et al. 2001, 2006; Smith and Ryan 2012). In the presented paper, long-term storage of tested LAB isolates was performed according to the protocol with 80% glycerol, the freezing formulation ratio was 1:1 cryoprotectant solution with freshly cultivated 24 h LAB cultures. After 18 months of frozen storage, in most cases LAB isolates maintained antagonistic effect against indicator bacteria, compared to the results obtained in previous antimicrobial tests. However, both reduction and disappearance of LAB biological activity were observed. This was in accordance with results previously reported by Fonseca et al. (2006), where the viability as well as acidification properties decreased after freezing storage.
The selected five LAB isolates were identified at both the genus and species level by MALDI-TOF MS and 16S rRNA gene sequencing. Chosen LAB isolates were identified as Lentilactobacillus buchneri (DG 068), Pediococcus pentosaceus (DG 059) as well as Lacticaseibacillus paracasei (KK 008, KK 033, KK 160). Lacticaseibacillus group (LCG) includes phenotypically as well as genotypically closely related species as L. casei, L. paracasei and L. rhamnosus, therefore, identification and differentiation of these probiotic strains using 16S rRNA gene sequencing is burdened by obtaining ambiguous results, due to inability to discriminate between species of very high sequence homology. Therefore, the Authors also used MALDI-TOF mass spectrometry as tool for identification of isolated LAB (Bizzini and Greub 2010; Dec et al. 2016). As the literature data states, MALDI-TOF MS using proteomics based identification, is able to discriminate between bacterial strains at both the genus and species levels and subspecies. It is a simple, fast, accurate and affordable method of identification (not including initial equipment costs) (Spinali et al. 2015; Tran et al. 2015; Huang et al. 2018). Using, 16S rRNA gene sequencing, in presented studies three selected LAB isolates were identified as L. paracasei species. The results were inconclusive on the subspecies level identification, indicating homology of isolated strains to L. paracasei subsp. tolerans, L. paracasei subsp. paracasei or just to L. paracasei. Identification based on the MALDI-TOF MS gave clear results, identifying these three LAB strains, on species level, as L. paracasei. To increase identification rates, as well as for obtaining identification on subspecies level, through matching the expansion of the BioTyper database is indispensable. (Dušková et al. 2012), also obtained ambiguous results, as two out of 148 Lactobacillus strains characterized by PCR and MALDI-TOF MS were, as the Authors stated, incorrectly assigned to L. paracasei and L. zeae. The Authors indicated that obtained results are related to the presence of species limitated entries in the BioTyper database as well as unclear Lactobacillus spp. taxonomic description.(Dec et al. 2014) also observed that the MALDI-TOF MS identification of the Lactobacillus species resulted with more than one assignment for 11.5% of tested strains, mostly in case of L. johnsonii. Therefore, it is clear that both 16S rRNA gene sequencing as well as MALDI-TOF MS are not infallible as they failed to differentiate between L. casei and L. paracasei. Lacticaseibacillus sp. can be identified with the greatest accuracy using approach which combines several genotypic methods (Huang et al. 2018).
Finally, the prototype of feed additive was prepared based on the selected LAB strains and survival tests during freeze-drying, storage as well as during the passage through the simulated GIT were carried out. Freeze-drying is commonly used as a preservation method; however, the stress factors can cause undesirable loss of viability of probiotics, therefore, different cryoprotectants are applied to increase their survival rate during the procedure (Siaterlis et al. 2009; Tomás et al. 2009). The mostly studied protectants include skim milk, whey proteins, varied sugars, or other bio-polymers (Meng et al. 2008; Han et al. 2018). In the presented study, the milk medium with trehalose and maltodextrin was used for LAB culture subjected to freeze-drying, which allowed to obtain good survival of bacteria during lyophilization and storage; however, both processes affects the cells viability. This is consistent with literature data indicating that some compounds such as trehalose, used in the presented study, as well as maltodextrin, glucose or sucrose may affect the viability during freeze-drying and storage (Strasser et al. 2009; Ren et al. 2019). Also, the effect of temperature during storage was underlined in literature data, which has been stated in the presented work. The stability of freeze-dried samples decreases during storage, and higher survival rates are recorded at lower storage temperatures. For example, (Ren et al. 2019) observed better viability of two Lactobacillus strain (L. salivarius and L. agilis) in the temperature of 4 °C in comparison with room temperature. Also, (Deeseenthum et al. 2007) reported that probiotic Bacillus spp. strains were more stable at the temperature of 4 °C than in room temperature. Moreover, it is worth underlining that the effects of storage may be highly variable and strain dependent (Turuvekere Sadguruprasad and Basavaraj 2018). Similar results were noticed in the presented study.
The feed additive prototype composed of five selected strains in two forms, as freeze-dried cultures and freeze-dried cultures mixed with feed, has been passaged in the in vitro digestive tract, where similar tendency was observed for both of them. The viability of LAB decreased in simulated solution of gastric juice and increased in simulated mixture of intestinal juice, which complies with other studies in literature. (Guerra et al. 2007) tested in vitro the survival of four strains including Pediococcus acidilactici, Enterococcus faecium, L. casei and L. lactis in the GIT and observed a decrease in the viable cells population during the passage through the stomach, while after 180 min of passage including the intestinal part the number of living cells was at a level of 106 CFU/mL. Similarly, Simões da Silva et al. (2020) observed significant decrease in the viability of different LAB strains in fermented products made of buffalo and cow milk after the gastric phase. The survival rate of bacteria may depend on the form in which they are administered, taking into account both the form of preservation and the type of matrix in which they are applied (Klingberg and Budde 2006). It has been found that the survival of LAB in the stomach can increase in the presence of food products that affect the pH value and can protect cells from the effects of pepsin and acid in the stomach (Bergamini et al. 2005). Ranadheera et al. (2012) have stated higher survival rate of L. acidophilus in ice cream and goat milk yogurt with 10% fat, when they were transferred through GIT. Also, Simões da Silva et al. (2020) reported that viability of LAB strains during the gastrointestinal simulation test remained more viable in dairy products made with buffalo milk compared to cow milk. In this study, however, a significant effect of the matrix was not observed, although in some sections of the simulated GIT, greater stability of the LAB population in the presence of feed could be observed.
Conclusions
LAB populations are specific as well as unique for different animals, therefore, bacterial isolation from suckling and weaned pigs was carried out to compose a multi-species probiotic feed additive prototype for swine. Functional properties of the obtained isolates varied significantly, therefore, selection of LAB isolates characterized by complementary features took place. Finally chosen 5 LAB isolates presenting satisfactory probiotic properties may be exploited as host specific probiotics for swine. The biggest advantage lies in the selection of functional microorganisms to maintain the relationship of bacterial origin with the group of monogastric animals for which the additive would be dedicated. All the research was conducted in vitro, therefore, further in vivo trials should proceed to assess the effect of the feed additive prototype on the performance and health of pigs. Preliminary studies for multistrain feed additive as well as animal feeding tests should be designed and performed before the field feeding use.
References
Adetoye A, Pinloche E, Adeniyi BA, Ayeni FA (2018) Characterization and anti-salmonella activities of lactic acid bacteria isolated from cattle faeces. BMC Microbiol 18:1–11. https://doi.org/10.1186/S12866-018-1248-Y
Angmo K, Kumari A, Savitri BTC (2016) Probiotic characterization of lactic acid bacteria isolated from fermented foods and beverage of Ladakh. LWT Food Sci Technol 66:428–435. https://doi.org/10.1016/j.lwt.2015.10.057
Arena MP, Silvain A, Normanno G et al (2016) Use of Lactobacillus plantarum strains as a bio-control strategy against food-borne pathogenic microorganisms. Front Microbiol 7:1–10. https://doi.org/10.3389/fmicb.2016.00464
Aujoulat F, Lebreton F, Romano S et al (2011) Comparative diffusion assay to assess efficacy of topical antimicrobial agents against Pseudomonas aeruginosa in burns care. Ann Clin Microbiol Antimicrob 10:1–10. https://doi.org/10.1186/1476-0711-10-27
Avantaggiato G, Havenaar R, Visconti A (2003) Assessing the zearalenone-binding activity of adsorbent materials during passage through a dynamic in vitro gastrointestinal model. Food Chem Toxicol 41:1283–1290. https://doi.org/10.1016/S0278-6915(03)00113-3
Barba-Vidal E, Martín-Orúe SM, Castillejos L (2019) Practical aspects of the use of probiotics in pig production: A review. Livest Sci 223:84–96. https://doi.org/10.1016/j.livsci.2019.02.017
Bautista-Gallego J, De CG, Reyes-Gavilan L et al (2017) Lactic acid bacteria and bifidobacteria with potential to design natural biofunctional health-promoting dairy foods. Biofunctional health-promoting dairy foods. Front Microbiol 8:846. https://doi.org/10.3389/fmicb.2017.00846
Bergamini CV, Hynes ER, Quiberoni A et al (2005) Probiotic bacteria as adjunct starters: influence of the addition methodology on their survival in a semi-hard Argentinean cheese. Food Res Int 38:597–604. https://doi.org/10.1016/j.foodres.2004.11.013
Birri DJ, Brede DA, Nes IF (2012) Salivaricin D, a novel intrinsically trypsin-resistant lantibiotic from Streptococcus salivarius 5M6c isolated from a healthy infant. Appl Environ Microbiol 78:402–410. https://doi.org/10.1128/AEM.06588-11
Bizzini A, Greub G (2010) Matrix-assisted laser desorption ionization time-of-flight mass spectrometry, a revolution in clinical microbial identification. Clin Microbiol Infect 16:1614–1619. https://doi.org/10.1111/j.1469-0691.2010.03311.x
Boisen S, Fernández JA (1997) Prediction of the total tract digestibility of energy in feedstuffs and pig diets by in vitro analyses. Anim Feed Sci Technol 68:277–286. https://doi.org/10.1016/S0377-8401(97)00058-8
Braune A, Blaut M (2016) Bacterial species involved in the conversion of dietary flavonoids in the human gut. Gut Microbes 7:216–234
Cao GT, Zeng XF, Chen AG et al (2013) Effects of a probiotic, Enterococcus faecium, on growth performance, intestinal morphology, immune response, and cecal microflora in broiler chickens challenged with Escherichia coli K88. Poult Sci 92:2949–2955. https://doi.org/10.3382/ps.2013-03366
Casarotti SN, Carneiro BM, Todorov SD et al (2017) In vitro assessment of safety and probiotic potential characteristics of Lactobacillus strains isolated from water buffalo mozzarella cheese. Ann Microbiol 67:289–301. https://doi.org/10.1007/s13213-017-1258-2
Charteris WP, Kelly PM, Morelli L, Collins JK (1998) Antibiotic susceptibility of potentially probiotic Lactobacillus species. J Food Prot 61:1636–1643. https://doi.org/10.4315/0362-028X-61.12.1636
Charteris WP, Kelly PM, Morelli L, Collins JK (1999) Development of an agar overlay disc diffusion method for the antibiotic susceptibility testing of potentially probiotic Lactobacillus and Bifidobacterium species. Egypt J Dairy Sci 27:71–82
Chen T, Wu Q, Li S et al (2014) Microbiological quality and characteristics of probiotic products in china. J Sci Food Agric 94:131–138. https://doi.org/10.1002/jsfa.6221
Cheng G, Hao H, Xie S et al (2014) Antibiotic alternatives: the substitution of antibiotics in animal husbandry? Front Microbiol. https://doi.org/10.3389/fmicb.2014.00217
Chiang ML, Chen HC, Chen KN et al (2015) Optimizing production of two potential probiotic lactobacilli strains isolated from piglet feces as feed additives for weaned piglets. Asian Austr J Anim Sci 28:1163–1170. https://doi.org/10.5713/ajas.14.0780
Clinical and Laboratory Standards Institute (CLSI) (2019) Performance standards for antimicrobial susceptibility testing. In: 29th ed. CLSI supplement M100. Wane, PA
Commane DM, Shortt CT, Silvi S et al (2005) Effects of fermentation products of pro- and prebiotics on trans-epithelial electrical resistance in an in vitro model of the colon. Nutr Cancer 51:102–109. https://doi.org/10.1207/s15327914nc5101_14
Daşkiran M, Öno AG, Cengiz Ö et al (2012) Influence of dietary probiotic inclusion on growth performance, blood parameters, and intestinal microflora of male broiler chickens exposed to posthatch holding time. J Appl Poultry Res 21:612–622. https://doi.org/10.3382/japr.2011-00512
de Souza BMS, Borgonovi TF, Casarotti SN et al (2019) Lactobacillus casei and Lactobacillus fermentum strains isolated from mozzarella cheese: probiotic potential, safety, acidifying kinetic parameters and viability under gastrointestinal tract conditions. Probiot Antimicrob Proteins 11:382–396. https://doi.org/10.1007/s12602-018-9406-y
Dec M, Urban-Chmiel R, Gnat S et al (2014) Identification of Lactobacillus strains of goose origin using MALDI-TOF mass spectrometry and 16S–23S rDNA intergenic spacer PCR analysis. Res Microbiol 165:190–201. https://doi.org/10.1016/j.resmic.2014.02.003
Dec M, Puchalski A, Urban-Chmiel R, Wernicki A (2016) 16S-ARDRA and MALDI-TOF mass spectrometry as tools for identification of Lactobacillus bacteria isolated from poultry. BMC Microbiol. https://doi.org/10.1186/s12866-016-0732-5
Deeseenthum S, Leelavatcharamas V, Brooks JD (2007) Effect of feeding Bacillus sp. as probiotic bacteria on growth of giant freshwater prawn (Macrobrachium rosenbergii de Man). Pak J Biol Sci 10:1481–1485. https://doi.org/10.3923/pjbs.2007.1481.1485
Divya JB, Varsha KK, Nampoothiri KM (2012) Newly isolated lactic acid bacteria with probiotic features for potential application in food industry. Appl Biochem Biotechnol 167:1314–1324. https://doi.org/10.1007/s12010-012-9561-7
Dowarah R, Verma AK, Agarwal N (2017) The use of Lactobacillus as an alternative of antibiotic growth promoters in pigs: a review. Anim Nutr 3:1–6. https://doi.org/10.1016/j.aninu.2016.11.002
Dowarah R, Verma AK, Agarwal N et al (2018a) Selection and characterization of probiotic lactic acid bacteria and its impact on growth, nutrient digestibility, health and antioxidant status in weaned piglets. PLoS ONE 13:e0192978. https://doi.org/10.1371/journal.pone.0192978
Dowarah R, Verma AK, Agarwal N, Singh P (2018b) Efficacy of species-specific probiotic Pediococcus acidilactici FT28 on blood biochemical profile, carcass traits and physicochemical properties of meat in fattening pigs. Res Vet Sci 117:60–64. https://doi.org/10.1016/j.rvsc.2017.11.011
Dušková M, Šedo O, Kšicová K et al (2012) Identification of lactobacilli isolated from food by genotypic methods and MALDI-TOF MS. Int J Food Microbiol 159:107–114. https://doi.org/10.1016/j.ijfoodmicro.2012.07.029
European Food Safety Authority Panel on Biological Hazards (BIOHAZ) (2013) Scientific opinion on the maintenance of the list of QPS biological agents intentionally added to food and feed (2013 update). EFSA J 11:1–107. https://doi.org/10.2903/j.efsa.2013.3449
European Commission (2020) Farm to Fork Strategy: for a fair, healthy and environmentally-friendly food system, European Commission. In: DG SANTE/Unit ‘Food information and composition, food waste’
European Food Safety Authority (EFSA) (2012) Guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA J 10:1–10. https://doi.org/10.2903/j.efsa.2012.2740
European Parliament and the Council of the European Union (2003) Regulation (EC) No 1831/2003 on additives for use in animal nutrition. Off J Eur Union 4:29–43
FAO/WHO (2002) Joint FAO/WHO working group report on drafting guidelines for the evaluation of probiotics in food. London, Ontario, Canada
Fayol-Messaoudi D, Berger CN, Coconnier-Polter MH et al (2005) pH-, lactic acid-, and non-lactic acid-dependent activities of probiotic lactobacilli against Salmonella enterica serovar typhimurium. Appl Environ Microbiol 71:6008–6013. https://doi.org/10.1128/AEM.71.10.6008-6013.2005
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Fonseca F, Béal C, Corrieu G (2001) Operating conditions that affect the resistance of lactic acid bacteria to freezing and frozen storage. Cryobiology 43:189–198. https://doi.org/10.1006/CRYO.2001.2343
Fonseca F, Marin M, Morris GJ (2006) Stabilization of frozen Lactobacillus delbrueckii subsp. bulgaricus in glycerol suspensions: freezing kinetics and storage temperature effects. Appl Environ Microbiol 72:6474–6482. https://doi.org/10.1128/AEM.00998-06
Fonseca F, Cenard S, Passot S (2015a) Freeze-drying of lactic acid bacteria. Cryopreservation and freeze-drying protocols. Springer, New York, pp 477–488
Fonseca F, Passot S, Meneghel J et al (2015b) Towards a better understanding of cryotolerance mechanisms in bacteria. Cryobiology 71:548. https://doi.org/10.1016/J.CRYOBIOL.2015.10.048
Gandomi H, Farhangfar A, Basti AA et al (2019) Auto and co-aggregation, hydrophobicity and adhesion properties of Lactobacillus plantarum strains isolated from Siahmazgi traditional cheese. Food Health 2:1–5
García-Hernández Y, Pérez-Sánchez T, Boucourt R et al (2016) Isolation, characterization and evaluation of probiotic lactic acid bacteria for potential use in animal production. Res Vet Sci 108:125–132. https://doi.org/10.1016/j.rvsc.2016.08.009
Gueimonde M, Sánchez B, de los Reyes-Gavilán CG, Margolles A (2013) Antibiotic resistance in probiotic bacteria. Front Microbiol 4:1–6. https://doi.org/10.3389/fmicb.2013.00202
Guerra NP, Bernárdez PF, Méndez J et al (2007) Production of four potentially probiotic lactic acid bacteria and their evaluation as feed additives for weaned piglets. Anim Feed Sci Technol 134:89–107. https://doi.org/10.1016/j.anifeedsci.2006.05.010
Guo XH, Kim JM, Nam HM et al (2010) Screening lactic acid bacteria from swine origins for multistrain probiotics based on in vitro functional properties. Anaerobe 16:321–326. https://doi.org/10.1016/j.anaerobe.2010.03.006
Gwiazdowski R, Kluczyńska K, Gwiazdowska D et al (2015) Impact of the selected lactic acid bacteria on the growth of pathogens occurring in oilseed rape cropsWpływ wybranych bakterii fermentacji mlekowej na wzrost patogenów występujących w uprawie rzepaku. Progress Plant Protect 55:446–451. https://doi.org/10.14199/PPP-2015-073
Han J, Chen D, Li S et al (2015) Antibiotic susceptibility of potentially probiotic Lactobacillus strains. Ital J Food Sci 27:282–289. https://doi.org/10.14674/1120-1770/IJFS.V270
Han L, Pu T, Wang X et al (2018) Optimization of a protective medium for enhancing the viability of freeze-dried Bacillus amyloliquefaciens B1408 based on response surface methodology. Cryobiology 81:101–106. https://doi.org/10.1016/j.cryobiol.2018.02.003
He T, Zhu YH, Yu J et al (2019) Lactobacillus johnsonii L531 reduces pathogen load and helps maintain short-chain fatty acid levels in the intestines of pigs challenged with Salmonella enterica infantis. Vet Microbiol 230:187–194. https://doi.org/10.1016/j.vetmic.2019.02.003
Hidalgo-Cantabrana C, Delgado S, Ruiz L et al (2018) Bifidobacteria and their health-promoting effects. In: Bugs as drugs. ASM Press, Washington, DC, USA, pp 73–98
Hill C, Guarner F, Reid G et al (2014) Expert consensus document: the international scientific association for probiotics and prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol 11:506–514. https://doi.org/10.1038/nrgastro.2014.66
Huang CH, Li SW, Huang L, Watanabe K (2018) Identification and classification for the Lactobacillus casei group. Front Microbiol 9:1–13. https://doi.org/10.3389/fmicb.2018.01974
Hung AT, Lin S-Y, Yang T-Y et al (2012) Effects of Bacillus coagulans ATCC 7050 on growth performance, intestinal morphology, and microflora composition in broiler chickens. Anim Prod Sci 52:874. https://doi.org/10.1071/AN11332
Huys G, D’Haene K, Swings J (2002) Influence of the culture medium on antibiotic susceptibility testing of food-associated lactic acid bacteria with the agar overlay disc diffusion method. Lett Appl Microbiol 34:402–406. https://doi.org/10.1046/J.1472-765X.2002.01109.X
International Organization for Standardization (ISO) (2010) ISO 10932|IDF 223:2010 Milk and milk products—determination of the minimal inhibitory concentration (MIC) of antibiotics applicable to bifidobacteria and non-enterococcal lactic acid bacteria (LAB). Geneva
Islam R, Hossain MdN, Alam MdK et al (2020) Antibacterial activity of lactic acid bacteria and extraction of bacteriocin protein. Adv Biosci Biotechnol 11:49–59. https://doi.org/10.4236/abb.2020.112004
Jeronymo-Ceneviva AB, de Paula AT, Silva LF et al (2014) Probiotic properties of lactic acid bacteria isolated from water-buffalo mozzarella cheese. Probiot Antimicrob Proteins 6:141–156. https://doi.org/10.1007/s12602-014-9166-2
Kim E-Y, Kim Y-H, Rhee M-H et al (2007) Selection of Lactobacillus sp. PSC101 that produces active dietary enzymes such as amylase, lipase, phytase and protease in pigs. J Gen Appl Microbiol 53:111–117. https://doi.org/10.2323/jgam.53.111
Klare I, Konstabel C, Müller-Bertling S et al (2005) Evaluation of new broth media for microdilution antibiotic susceptibility testing of lactobacilli, pediococci, lactococci, and bifidobacteria. Appl Environ Microbiol 71:8982–8986. https://doi.org/10.1128/AEM.71.12.8982-8986.2005
Klingberg TD, Budde BB (2006) The survival and persistence in the human gastrointestinal tract of five potential probiotic lactobacilli consumed as freeze-dried cultures or as probiotic sausage. Int J Food Microbiol 109:157–159. https://doi.org/10.1016/j.ijfoodmicro.2006.01.014
Kos B, Šušković J, Vuković S et al (2003) Adhesion and aggregation ability of probiotic strain Lactobacillus acidophilus M92. J Appl Microbiol 94:981–987. https://doi.org/10.1046/j.1365-2672.2003.01915.x
Kumar S, Stecher G, Li M et al (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Landy N, Kavyani A (2013) A multi-strain probiotic on performance, immune responses and cecal microflora composition in broiler chickens reared under cyclic heat stress condition. Iran J Appl Anim Sci 3:703–708
Lee H, Gilliland S, Carter S (2001) Amylolytic Cultures of Lactobacillus acidophilus: potential probiotics to improve dietary starch utilization. J Food Sci 66:338–344. https://doi.org/10.1111/j.1365-2621.2001.tb11343.x
Leser TD, Amenuvor JZ, Jensen TK et al (2002) Culture-independent analysis of gut bacteria: the pig gastrointestinal tract microbiota revisited. Appl Environ Microbiol 68:673–690. https://doi.org/10.1128/AEM.68.2.673-690.2002
Lim WS, Lee JW (2013) Influence of pretreatment condition on the fermentable sugar production and enzymatic hydrolysis of dilute acid-pretreated mixed softwood. Biores Technol 140:306–311. https://doi.org/10.1016/J.BIORTECH.2013.04.103
Lin WH, Yu B, Jang SH, Tsen HY (2007) Different probiotic properties for Lactobacillus fermentum strains isolated from swine and poultry. Anaerobe 13:107–113. https://doi.org/10.1016/j.anaerobe.2007.04.006
Lo Verso L, Lessard M, Talbot G et al (2018) Isolation and selection of potential probiotic bacteria from the pig gastrointestinal tract. Probiot Antimicrob Proteins 10:299–312. https://doi.org/10.1007/s12602-017-9309-3
Long SF, Xu YT, Pan L et al (2018) Mixed organic acids as antibiotic substitutes improve performance, serum immunity, intestinal morphology and microbiota for weaned piglets. Anim Feed Sci Technol 235:23–32. https://doi.org/10.1016/j.anifeedsci.2017.08.018
Medellin-Peña MJ, Wang H, Johnson R et al (2007) Probiotics affect virulence-related gene expression in Escherichia coli O157:H7. Appl Environ Microbiol 73:4259–4267. https://doi.org/10.1128/AEM.00159-07
Meng XC, Stanton C, Fitzgerald GF et al (2008) Anhydrobiotics: the challenges of drying probiotic cultures. Food Chem 106:1406–1416. https://doi.org/10.1016/j.foodchem.2007.04.076
Mishra V, Prasad DN (2005) Application of in vitro methods for selection of Lactobacillus casei strains as potential probiotics. Int J Food Microbiol 103:109–115. https://doi.org/10.1016/J.IJFOODMICRO.2004.10.047
Mookiah S, Sieo CC, Ramasamy K et al (2014) Effects of dietary prebiotics, probiotic and synbiotics on performance, caecal bacterial populations and caecal fermentation concentrations of broiler chickens. J Sci Food Agric 94:341–348. https://doi.org/10.1002/jsfa.6365
Moslehishad M, Mirdamadi S, Ehsani MR et al (2013) The proteolytic activity of selected lactic acid bacteria in fermenting cow’s and camel’s milk and the resultant sensory characteristics of the products. Int J Dairy Technol 66:279–285. https://doi.org/10.1111/1471-0307.12017
Mountzouris KC, Balaskas C, Xanthakos I et al (2009) Effects of a multi-species probiotic on biomarkers of competitive exclusion efficacy in broilers challenged with Salmonella enteritidis. Br Poult Sci 50:467–478. https://doi.org/10.1080/00071660903110935
Muñoz-Atienza E, Gómez-Sala B, Araújo C et al (2013) Antimicrobial activity, antibiotic susceptibility and virulence factors of lactic acid bacteria of aquatic origin intended for use as probiotics in aquaculture. BMC Microbiol 13:1–22. https://doi.org/10.1186/1471-2180-13-15/TABLES/6
Niba AT, Beal JD, Kudi AC, Brooks PH (2009) Bacterial fermentation in the gastrointestinal tract of non-ruminants: influence of fermented feeds and fermentable carbohydrates. Trop Anim Health Prod 41:1393–1407. https://doi.org/10.1007/s11250-009-9327-6
Niku-Paavola ML, Laitila A, Mattila-Sandholm T, Haikara A (1999) New types of antimicrobial compounds produced by Lactobacillus plantarum. J Appl Microbiol 86:29–35. https://doi.org/10.1046/j.1365-2672.1999.00632.x
Pang H, Qin G, Tan Z et al (2011) Natural populations of lactic acid bacteria associated with silage fermentation as determined by phenotype, 16S ribosomal RNA and recA gene analysis. Syst Appl Microbiol 34:235–241. https://doi.org/10.1016/j.syapm.2010.10.003
Patterson JA, Burkholder KM (2003) Application of prebiotics and probiotics in poultry production. Poult Sci 82:627–631. https://doi.org/10.1093/ps/82.4.627
Pérez-Sánchez T, Balcázar JL, García Y et al (2011) Identification and characterization of lactic acid bacteria isolated from rainbow trout, Oncorhynchus mykiss (Walbaum), with inhibitory activity against Lactococcus garvieae. J Fish Dis 34:499–507. https://doi.org/10.1111/j.1365-2761.2011.01260.x
Petsuriyawong B, Khunajakr N (2011) Screening of probiotic lactic acid bacteria from piglet feces. Kasetsart J Natural Sci 45:245–253
Ranadheera CS, Evans CA, Adams MC, Baines SK (2012) In vitro analysis of gastrointestinal tolerance and intestinal cell adhesion of probiotics in goat’s milk ice cream and yogurt. Food Res Int 49:619–625. https://doi.org/10.1016/j.foodres.2012.09.007
Rautray AK, Patra RC, Sardar KK, Sahoo G (2011) Potential of probiotics in livestock production. Explor Anim Med Res 1:20–28
Ren H, Zentek J, Vahjen W (2019) Optimization of production parameters for probiotic Lactobacillus strains as feed additive. Molecules. https://doi.org/10.3390/molecules24183286
Ripamonti B, Agazzi A, Bersani C et al (2011) Screening of species-specific lactic acid bacteria for veal calves multi-strain probiotic adjuncts. Anaerobe 17:97–105. https://doi.org/10.1016/j.anaerobe.2011.05.001
Saitou N, Imanishi T (1989) Relative efficiencies of the Fitch-Margoliash, maximum-parsimony, maximum-likelihood, minimum-evolution, and neighbor-joining methods of phylogenetic tree construction in obtaining the correct tree’. Mol Biol Evol 6:514–525
Sánchez-Ortiz AC, Luna-González A, Campa-Córdova ÁI et al (2015) Isolation and characterization of potential probiotic bacteria from pustulose ark (Anadara tuberculosa) suitable for shrimp farming. Lat Am J Aquat Res 43:123–136. https://doi.org/10.3856/vol43-issue1-fulltext-11
Sanni AI, Morlon-Guyot J, Guyot JP (2002) New efficient amylase-producing strains of Lactobacillus plantarum and L. fermentum isolated from different Nigerian traditional fermented foods. Int J Food Microbiol 72:53–62. https://doi.org/10.1016/S0168-1605(01)00607-9
Schrezenmeir J, De Vrese M (2001) Probiotics, prebiotics, and synbiotics—approaching a definition. Am J Clin Nutr 2001:361–364
Shim YH, Ingale SL, Kim JS et al (2012) A multi-microbe probiotic formulation processed at low and high drying temperatures: effects on growth performance, nutrient retention and caecal microbiology of broilers. Br Poult Sci 53:482–490. https://doi.org/10.1080/00071668.2012.690508
Siaterlis A, Deepika G, Charalampopoulos D (2009) Effect of culture medium and cryoprotectants on the growth and survival of probiotic lactobacilli during freeze drying. Lett Appl Microbiol 48:295–301. https://doi.org/10.1111/j.1472-765X.2008.02529.x
Simões da Silva TM, Piazentin ACM, Mendonça CMN et al (2020) Buffalo milk increases viability and resistance of probiotic bacteria in dairy beverages under in vitro simulated gastrointestinal conditions. J Dairy Sci 103:7890–7897. https://doi.org/10.3168/jds.2019-18078
Sirichokchatchawan W, Pupa P, Praechansri P et al (2018) Autochthonous lactic acid bacteria isolated from pig faeces in Thailand show probiotic properties and antibacterial activity against enteric pathogenic bacteria. Microb Pathog 119:208–215. https://doi.org/10.1016/j.micpath.2018.04.031
Siva Kumar K, Sastry N, Polaki H, Mishra V (2015) Colon cancer prevention through probiotics: an overview. J Cancer Sci Therapy. https://doi.org/10.4172/1948-5956.1000329
Smirnova DV, Zalomova LV, Zagainova AV et al (2019) Cryopreservation of the human gut microbiota: current state and perspectives. Int J Med Microbiol 309:259–269. https://doi.org/10.1016/J.IJMM.2019.06.001
Smith D, Ryan M (2012) Implementing best practices and validation of cryopreservation techniques for microorganisms. Sci World J. https://doi.org/10.1100/2012/805659
Spaiser SJ, Culpepper T, Nieves C et al (2015) Lactobacillus gasseri KS-13, Bifidobacterium bifidum G9–1, and Bifidobacterium longum MM-2 ingestion induces a less inflammatory cytokine profile and a potentially beneficial shift in gut microbiota in older adults: a randomized, double-blind, placebo-con. J Am Coll Nutr 34:459–469. https://doi.org/10.1080/07315724.2014.983249
Spinali S, Van Belkum A, Goering RV et al (2015) Microbial typing by matrix-assisted laser desorption ionization-time of flight mass spectrometry: do we need guidance for data interpretation? J Clin Microbiol 53:760–765. https://doi.org/10.1128/JCM.01635-14
Strasser S, Neureiter M, Geppl M et al (2009) Influence of lyophilization, fluidized bed drying, addition of protectants, and storage on the viability of lactic acid bacteria. J Appl Microbiol 107:167–177. https://doi.org/10.1111/j.1365-2672.2009.04192.x
Taheri H, Tabandeh F, Moravej H et al (2009) Potential probiotic of Lactobacillus johnsonii LT171 for chicken nutrition. Afr J Biotech 8:5833–5837. https://doi.org/10.5897/ajb09.1062
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA 101:11030–11035. https://doi.org/10.1073/pnas.0404206101
Thomas CM, Versalovic J (2010) Probiotics-host communication. Gut Microbes 1:148–163. https://doi.org/10.4161/gmic.1.3.11712
Todorov SD, Furtado DN, Saad SMI et al (2011) Potential beneficial properties of bacteriocin-producing lactic acid bacteria isolated from smoked salmon. J Appl Microbiol 110:971–986. https://doi.org/10.1111/j.1365-2672.2011.04950.x
Tomás MSJ, Bru E, Martos G, Nader-Macías ME (2009) Stability of freeze-dried vaginal Lactobacillus strains in the presence of different lyoprotectors. Can J Microbiol 55:544–552. https://doi.org/10.1139/W08-159
Torshizi MAK, Rahimi S, Mojgani N et al (2008) Screening of indigenous strains of lactic acid bacteria for development of a probiotic for poultry. Asian Austr J Anim Sci 21:1495–1500. https://doi.org/10.5713/ajas.2008.80081
Tran A, Alby K, Kerr A et al (2015) Cost savings realized by implementation of routine microbiological identification by matrix-assisted laser desorption ionization-time of flight mass spectrometry. J Clin Microbiol 53:2473–2479. https://doi.org/10.1128/JCM.00833-15
Turuvekere Sadguruprasad L, Basavaraj M (2018) Statistical modelling for optimized lyophilization of Lactobacillus acidophilus strains for improved viability and stability using response surface methodology. AMB Express 8:10. https://doi.org/10.1186/s13568-018-0659-3
Versantvoort CHM, Oomen AG, Van De Kamp E et al (2005) Applicability of an in vitro digestion model in assessing the bioaccessibility of mycotoxins from food. Food Chem Toxicol 43:31–40. https://doi.org/10.1016/j.fct.2004.08.007
Vu H, Mccoy L, Carino E et al (2002) Burn wound infection susceptibilities to topical agents: the nathan’s agar well diffusion technique. P T 27:390–397
Xin J, Zeng D, Wang H et al (2020) Probiotic Lactobacillus johnsonii BS15 promotes growth performance, intestinal immunity, and gut microbiota in piglets. Probiot Antimicrob Proteins 12:184–193. https://doi.org/10.1007/s12602-018-9511-y
Yang F, Hou C, Zeng X, Qiao S (2015) The use of lactic acid bacteria as a probiotic in swine diets. Pathogens 4:34–45. https://doi.org/10.3390/pathogens4010034
Yousefi B, Eslami M, Ghasemian A et al (2019) Probiotics importance and their immunomodulatory properties. J Cell Physiol 234:8008–8018. https://doi.org/10.1002/jcp.27559
Zhang ZF, Kim IH (2014) Effects of multistrain probiotics on growth performance, apparent ileal nutrient digestibility, blood characteristics, cecal microbial shedding, and excreta odor contents in broilers. Poult Sci 93:364–370. https://doi.org/10.3382/ps.2013-03314
Author information
Authors and Affiliations
Contributions
Conceptualization, KM and DG; methodology, KM and DG; software, KM and DG; validation, KM and DG; formal analysis, KM and DG; investigation, KM and DG; resources, KM and DG; data curation, KM; writing—original draft preparation, KM and DG; writing—review and editing, KM and DG; visualization, KM and DG; supervision, KM and DG; project administration, KM; funding acquisition, KM and DG. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this paper.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marchwińska, K., Gwiazdowska, D. Isolation and probiotic potential of lactic acid bacteria from swine feces for feed additive composition. Arch Microbiol 204, 61 (2022). https://doi.org/10.1007/s00203-021-02700-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-021-02700-0