Abstract
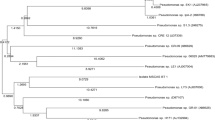
In this study pendimethalin degrading indigenous soil bacterium was isolated from rice field (supplemented with pendimethalin) and identified as, Pseudomonas strain PD1 on the basis of 16S rRNA phylogenetic analysis. Biodegradation of pendimethalin by this strain was evaluated by spectrophotometric scanning and FTIR analysis of degraded compounds in minimal salt media. Decrease in concentration of pendimethalin at λmax (430 nm) under spectrophotometric scanning is a measurement of time taken by bacterium strain PD1 to degrade pendimethalin. Degraded products were further analyzed by comparing stretching and bending pattern of chemical groups attached to compounds using FTIR spectroscopy. FTIR profile represented disappearance of nitrate group in degraded product by bacterium strain PD1 in minimal salt medium. Molecular docking of pendimethalin on nitro-reductase was done to suggest first enzyme of pathway used by bacterium strain PD1 to degrade pendimethalin. Analysis on degradation by strain PD1 shows that newly isolated strain PD1 can degrade 77.05% of pendimethalin at 50 mgL−1 concentration in 30 h incubation under room temperature. Thus, the study here shed a light on degradation potential of Pseudomonas.






Similar content being viewed by others
References
Barbash JE, Resek EA (1996) Pesticides in ground water: distribution, trends, and governing factors. Ann Arbor Press, Ann Arbor
Barriuso E, Houot S, Serra–Wittling C (1997) Influence of compost addition to soil on the behaviour of herbicides. Pestic Sci 49:65–75
Barua AS et al (2010) Degradation of pendimethalin by soil fungi. Pest Manage Sci 29(4):419–425
Belal BE, El-Nady MF (2013) Bioremediation of pendimethalin-contaminated soil. Afr J Microbiol Res 7(21):2574–2588
Capel PD, Lin M, Wotzka PJ (1998) Wet atmospheric deposition of pesticides in Minnesota, 1989–94. In: Capel PD (ed) Water-Resources Investigations Report. US Geological Survey, Mounds View
Chen Y, Zou C, Mastalerz M, Hu S, Gasaway C, Tao X (2015) Applications of micro-fourier transform infrared spectroscopy (FTIR) in the geological sciences-a review. Int J Mol Sci 16(12):30223–30250
Copley SD (2000) Evolution of a metabolic pathway for degradation of a toxic xenobiotic, the patchwork approach. Trends Biochem Sci 25:261–265
Diez MC (2010) Biological aspects involved in the degradation of organic pollutants. J Soil Sci Plant Nutr 10:244–267
Han Y, Tang Z, Bao H, Wu D, Deng X, Guo G, Dai B (2019) Degradation of pendimethalin by the yeast YC2 and determination of its two main metabolites. RSC Adv 9(1):491–497
Hou L et al (2004) Pendimethalin exposure and cancer risk among pesticide applicators: a report from the US-based agricultural health study. Ann Epidemiol 14:608
Hu K et al (2020) Characterization of carbaryl-degrading strain Bacillus licheniformis B-1 and its hydrolase identification. Biodegradation. https://doi.org/10.1007/s10532-020-09899-7
Kamrin MA (1997) Pesticide profiles: toxicity, environmental impact, and fate. CRC Press, Boca Raton
Köck-Schulmeyer M, Ginebreda A, González S, Cortina JL, López de Alda M, Barceló D (2012) Analysis of the occurrence and risk assessment of polar pesticides in the Llobregat River Basin (NE Spain). Chemosphere 86:8–16
Kole RK et al (1994) Bacterial degradation of the herbicide pendimethalin and activity evaluation of its metabolites. Bull Environ Contam Toxicol 52(5):779
Lee YD, Kim HJ, Chung JB, Jeong BR (2000) Loss of pendimethalin in runoff and leaching from turfgrass land under simulated rainfall. J Agric Food Chem 48(11):5376–5382
Lu J, Wu L, Newman J, Faber B, Gan J (2006) Degradation of pesticides in nursery recycling pond waters. J Agric Food Chem 54(7):2658–2663
Mao K, Chen Y, Wu Z, Zhou X, Shen A, Hu J (2014) Catalytic strategy for efficient degradation of nitroaromatic pesticides by using gold nanoflower. J Agric Food Chem 62(44):10638–10645
Meng XY, Zhang HX, Mezei M, Cui M (2011) Molecular docking: a powerful approach for structure-based drug discovery. Curr Comput Aid Drug 7(2):146–157
Mohan SV, Krishna MR, Muralikrishna P, Shailaja S, Sarma PN (2007) Solid phase bioremediation of pendimethalin in contaminated soil and evaluation of leaching potential. Bioresour Technol 98(15):2905–2910
Ni H, Li N, Qiu J, Chen Q, He J (2018) Biodegradation of pendimethalin by Paracoccus sp. P13. Curr Microbiol 75(8):1077–1083
Oh ST, Kim JR, Premier GC, Lee TH, Kim C, Sloan WT (2010) Sustainable wastewater treatment: how might microbial fuel cells contribute. Biotechnol Adv 28:871–881
Pino N, Peñuela G (2011) Simultaneous degradation of the pesticides methyl parathion and chlorpyrifos by an isolated bacterial consortium from a contaminated site. Int Biodeterior Biodegrad 65:827–883
Racke KD (2000) Pesticides for turfgrass pest management: uses and environmental issues. In: Racke KD (ed) Fate and management of turfgrass chemicals, vol 743. American Chemical Society, Washington, pp 45–64
Roca E, D’Errico E, Izzo A, Strumia S, Esposito A, Fiorentino A (2009) In vitro saprotrophic basidiomycetes tolerance to pendimethalin. Int Biodeterior Biodegrad 63:182–186
Singh BK, Walker A, Morgan JAW, Wright DJ (2004) Biodegradation of chlorpyrifos by Enterobacter strain B-14 and its use in bioremediation of contaminated soils. Appl Environ Microbiol 70(8):4855–4863
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thomsen R, Chistensen MH (2006) MolDock: a new technique for high-accuracy molecular docking. J Med Chem 49(11):3315–3321
Trivedi VD et al (2016) Insights into functional and evolutionary analysis of carbaryl metabolic pathway from Pseudomonas sp. strain C5pp. Sci Rep 6:38430. https://doi.org/10.1038/srep38430
Trivedi N, Tandon S, Dubey A (2017) Fourier transform infrared spectroscopy (FTIR) profiling of red pigment produced by Bacillus subtilis PD5. Afr J Biotech 16:1507–1512
Udatha DG, Sugaya N, Olsson L, Panagiotou G (2012) How well do the substrates KISS the enzyme? Molecular docking program selection for feruloyl esterases. Sci Rep 2(1):1–8
Wang Y, Zhang G, Yan J, Gong D (2014) Inhibitory effect of morin on tyrosinase: insights from spectroscopic and molecular docking studies. Food Chem 163:226–233
Yang L, Zhao YH, Zhang BX, Zhang YCH, X, (2005) Isolation and characterization of a chlorpyrifos and 3, 5, 6-trichloro-2-pyridinol degrading bacterium. FEMS Microbiol Lett 251(1):67–73
Zhang C, Hughes JB, Nishino SF, Spain JC (2000) Slurry-phase biological treatment of 2, 4-dinitrotoluene and 2, 6-dinitrotoluene: role of bioaugmentation and effects of high dinitrotoluene concentrations. Environ Sci Technol 34(13):2810–2816
Acknowledgements
The laboratory assistance of Mr. Farid, Mr. Deepak, scientific discussion with Dr. A. K. Verma (Department of Biochemistry CBSH, GBPUAT, India) are gratefully acknowledged.
Funding
The work was partly supported by DST-FIST India.
Author information
Authors and Affiliations
Contributions
Both authors have equal contributions.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Trivedi, N., Dubey, A. Degradation studies of pendimethalin by indigenous soil bacterium Pseudomonas strain PD1 using spectrophotometric scanning and FTIR. Arch Microbiol 203, 4499–4507 (2021). https://doi.org/10.1007/s00203-021-02439-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02439-8