Abstract
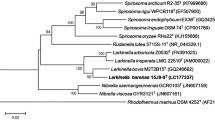
Strain YIM B00363T, a Gram-positive, aerobic, non-motile, rod-shaped, spore-forming bacterium, was isolated from saline soil samples collected from a salt lake in Xinjiang province, north-west China, and was characterized using a polyphasic approach. The optimum growth temperature was 37 °C and the optimum pH was 7.5–8.0. The major menaquinone was MK-7; anteiso-C15:0 (53.52%), iso-C15:0 (15.04%) and C16:0 (12.76%) were the predominant cellular fatty acids. The diagnostic diamino acid of the cell wall peptidoglycan was meso-diaminopimelic acid. The phospholipids were phosphatidylethanolamine, diphosphatidylglycerol, phosphatidylglycerol, unidentified phospholipids, unidentified glycolipids and unknown lipids. The DNA G + C content of the type strain was 50.4 mol%. Phylogenetic analysis based on 16S rRNA gene sequences indicated that the strain YIM B00363T belonged to a cluster comprising species of the genus Paenibacillus. The nearest relatives were P. residui MC-246T and P. senegalensis JC66T, with 93.2% and 92.8% gene sequence similarities, respectively. On the basis of its phenotypic characteristics and phylogenetic distinctivenes, strain YIM B00363T represents a novel species of the genus Paenibacillus, for which the name Paenibacillus turpanensis sp. nov. is proposed. The type strain is YIM B00363T (= CGMCC 1.17507T = KCTC 43184T).

Similar content being viewed by others
Abbreviations
- NFA:
-
Nitrogen Fixing agar
- TSA:
-
Tryptic soy agar
- DPG:
-
Diphosphatidylglycerol
- PG:
-
Phosphatidylglycerol
- PE:
-
Phosphatidylethanolamine
- PL:
-
Unknown phospholipid
- GL:
-
Unidentified glycolipids
- UL:
-
Unknown lipids
References
Ash C, Priest FG, Collins MD (1993) Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Proposal for the creation of a new genus Paenibacillus. Antonie Van Leeuwenhoek 64:253–260
Baron EJ, Finegold SM (1990) Bailey and Scott’s diagnostic microbiology, 8th edn. Mosby, St Louis
Collins MD, Jones D (1980) Lipids in the classification and identification of coryneform bacteria containing peptidoglycans based on 2, 4-diaminobutyric acid. J Appl Bacteriol 48:459–470
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
Dai XL, Shi KX, Wang X, Fan J, Wang R, Zheng SX, Wang GJ (2019) Paenibacillus flagellatus sp. nov., isolated from selenium mineral soil. Int J Syst Evol Microbiol 69:183–188
Dong XZ, Cai MY (2001) Determinative manual for routine bacteriology. Scientific Press, Beijing
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Grau FH, Wilson PW (1962) Physiology of nitrogen fixationby Bacillus polymyxa. J Bacteriol 83:490–496
Hong YY, Ma YC, Zhou YG, Gao F, Liu HC, Chen SF (2009) Paenibacillus sonchi sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Sonchus oleraceus. Int J Syst Evol Microbiol 59:2656–2661
Jin HJ, Lv J, Chen SF (2011) Paenibacillus sophorae sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Sophora japonica. Int J Syst Evol Microbiol 61:767–771
Kampfer P, Rabinovitch L, Salkinoja-Salonen MS, Seldin L, Ventosa A (2009) Proposed minimal standards for describing new taxa of aerobic, endospore-forming bacteria. Int J Syst Evol Microbiol 59:2114–2121
Keswani J, Whitman WB (2001) Relationship of 16S rRNA sequence similarity to DNA hybridization in prokaryotes. Int J Syst Evol Microbiol 51:667–678
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lechevalier MP, Lechevalier HA (1970) Chemical composition as a criterion in the classification of aerobic actinomycetes. Int J Syst Bacteriol 20:435–443
Logan NA, Berge O, Bishop AH, Busse HJ, de Vos P, Fritze D, Heyndrickx M, Kämpfer P, Rabinovitch L, Salkinoja-Salonen MS, Seldin L, Ventosa A (2009) Proposed minimal standards for describing new taxa of aerobic, endospore-forming bacteria. Int J Syst Evol Microbiol 59:2114–2121
Ma YC, Chen SF (2008) Paenibacillus forsythiae sp. nov., a nitrogen-fixing species isolated from rhizosphere soil of Forsythia mira. Int J Syst Evol Microbiol 58:319–323
Ma Y, Xia Z, Liu X, Chen S (2007) Paenibacillus sabinae sp., nov., a nitrogen-fixing species isolated from the rhizosphere soils of shrubs. Int J Syst Evol Microbiol 57:6–11
Minnikin DE, Collins MD, Goodfellow M (1979) Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J Appl Bacteriol 47:87–95
Mishra AK, Lagier JC, Rivet R, Raoult D, Fournier PE (2012) Non-contiguous finished genome sequence and description of Paenibacillus senegalensis sp. nov. Stand Genom Sci 7:70–81
Nam JH, Bae W, Lee DH (2008) Oceanobacillus caeni sp. nov., isolated from a Bacillus-dominated wastewater treatment system in Korea. Int J Syst Evol Microbiol 58:1109–1113
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids, MIDI Technical Note, vol 10 1. MIDI Inc, Newark
Seldin L (2011) Paenibacillus, nitrogen fixation and soil fertility. In: Logan NA, De Vos P (eds) Endospore-forming soil bacteria, soil biology, vol 27. Springer, Berlin, Heidelberg, pp 287–307
Seldin L, Van Elsas JD, Penido EGC (1984) Bacillus azotofixans sp. nov., a nitrogen-fixing species from Brazilian soils and grass roots. Int J Syst Bacteriol 34:451–456
Shida O, Takagi H, Kadowaki K, Nakamura LK, Komagata K (1997) Transfer of Bacillus alginolyticus, Bacillus chondroitinus, Bacillus curdlanolyticus, Bacillus glucanolyticus, Bacillus kobensis, and Bacillus thiaminolyticus to the genus Paenibacillus and emended description of the genus Paenibacillus. Int J Syst Bacteriol 47:289–298
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Takeda M, Suzuki I, Koizumi J (2005) Paenibacillus hodogayensis sp. nov., capable of degrading the polysaccharide produced by Sphaerotilus natans. Int J Syst Evol Microbiol 55:737–741
Tang SL, Nuttall S, Ngui K, Fisher C, Lopez P, Dyall-Smith M (2002) HF2: a double-stranded DNA tailed haloarchaeal virus with a mosaic genome. Mol Microbiol 44:283–296
Tang SK, Wang Y, Chen Y, Lou K, Cao LL, Xu LH, Li WJ (2009) Zhihengliuella alba sp. nov., and emended description of the genus Zhihengliuella. Int J Syst Evol Microbiol 59:2025–2032
Tang SK, WangY ZH, Lee JC, Lou K, Kim CJ, Li WJ (2010) Haloechinothrix alba gen. nov., sp. nov., a halophilic, filamentous actinomycete of the suborder Pseudonocardineae. Int J Syst Evol Microbiol 60:2154–2158
Tarrand JJ, Gröschel DH (1982) Rapid, modified oxidase test for oxidase-variable bacterial isolates. J Clin Microbiol 16:772–774
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tindall BJ, Rosselló-Móra R, Busse HJ, Ludwig W, Kämpfer P (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60:249–266
Ueda J, Yamamoto SC, Kurosawa N (2013) Paenibacillus thermoaerophilus sp. nov., a moderately thermophilic bacterium isolated from compost. Int J Syst Evol Microbiol 63:3330–3335
Vaz-Moreira I, Figueira V, Lopes AR, Pukall R, Sproer C, Schumann P, Nunes OC, Manaia CM (2010) Paenibacillus residui sp. nov. isolated from urban waste compost. Int J Syst Evol Microbiol 60:2415–2419
Xie CH, Yokota A (2003) Phylogenetic analyses of Lampropedia hyalina based on the 16S rRNA gene sequence. J Gen Appl Microbiol 49:345–349
Xie JB, Zhang LH, Zhou YG, Liu HC, Chen SF (2012) Paenibacillus taohuashanense sp., nov. a nitrogen-fixing species isolated from rhizosphere soil of the root of Caragana kansuensis Pojark. Antonie Van Leeuwenhoek 102:735–741
Yoon MH, Ten LN, Im WT (2007) Paenibacillus ginsengarvi sp. nov., isolated from soil from ginseng cultivation. Int J Syst Evol Microbiol 57:1810–1814
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: A taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Acknowledgements
This study was performed with the support of the National Natural Science Foundation of China (31760003, 31500011 and 31270055), the Natural Science Foundation of Yunnan Province (2017FB039) and the South and Southeast Asia Cooperation Base on Microbiological Resource Prevention and Utilization (2018IA100)
Author information
Authors and Affiliations
Contributions
LY and H-WH: carried out the data analysis, part of chemical classification and wrote the manuscript. YW, Y-RK and MY: performed the polyphasic taxonomy except chemical classification. YL, X-QW and G-FZ: prepared the experiments and isolated the novel strain. W-YZ and S-KT: designed the separation medium and directed the classification and takes full responsibility for the final submission. All the authors reviewed and approved the final version of the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The NCBI GenBank accession number for the 16S rRNA gene sequence of strain YIM B00363T is MT032315. The draft whole-genome sequence for YIM B00363T has been deposited at DDBJ/ENA/GenBank under accession number WTLI00000000.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, L., Huang, HW., Wang, Y. et al. Paenibacillus turpanensis sp. nov., isolated from a salt lake of Turpan city in Xinjiang province, north-west China. Arch Microbiol 203, 77–83 (2021). https://doi.org/10.1007/s00203-020-02003-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02003-w