Abstract
We have described that formation of basidiocarps by Ustilago maydis requires illumination. In the current research, we have proceeded to analyze what kind of light receptors are involved in this phenomenon. Accordingly, we investigated whether the homologues of the White Collar (WC), and the phytochrome (PHY) genes played a role in this process. Mutants deficient in either one of the three U. maydis WC homologue genes (WCO1a, WCO1b, WCO2), or the phytochrome-encoding the PHY gene were obtained. Phenotypic analysis of the mutants showed that ∆wco1a mutants formed similar numbers of basidiocarps than wild-type strain, whereas ∆wco1b mutants were severely affected in basidiocarp formation when illuminated with white, blue or red light. ∆wco2 and ∆phy1 mutants did not form basidiocarps under any illumination condition. These data indicate that Wco1a is the main blue light receptor, and Wco1b may operate as a secondary blue light receptor; Phy1 is the red light receptor, and Wco2 the transcription factor that controls the photo stimulation of the genes involved in the formation of fruiting bodies. It is suggested that effectiveness of the light receptors depends on the whole structure of the complex, possibly, because their association is necessary to maintain their functional structure.






Similar content being viewed by others
References
Ambra R, Grimaldi B, Zamboni S, Filetici P, Macino G, Ballario P (2004) Photomorphogenesis in the hypogeous fungus Tuber borchii: isolation and characterization of Tbwc-1, the homologue of the blue-light photoreceptor of Neurospora crassa. Fungal Genet Biol 41:688–697. https://doi.org/10.1016/j.fgb.2004.02.004
Arjona D, Aragón C, Aguilera JA, Ramírez L, Pisabarro AG (2009) Reproducible and controllable light induction of in vitro fruiting of the white-rot basidiomycete Pleurotus ostreatus. Mycol Res 113:552–558. https://doi.org/10.1016/j.mycres.2008.12.006
Bahn YS, Xue C, Idnurm A, Rutherford JC, Heitman J, Cardenas ME (2007) Sensing the environment: lessons from fungi. Nat Rev Microbiol 5:57–69. https://doi.org/10.1038/nrmicro1578
Ballario P, Vittorioso P, Magrelli A, Talora C, Cabibbo A, Macino G (1996) White collar-1, a central regulator of blue light responses in Neurospora, is a zinc finger protein. EMBO J 15:1650–1657
Banuett F, Herskowitz I (1989) Different a alleles of Ustilago maydis are necessary for maintenance of filamentous growth but not for meiosis. Proc Natl Acad Sci USA 86:5878–5882. https://doi.org/10.1073/pnas.86.15.5878
Bayram O, Braus GH, Fischer R, Rodriguez-Romero J (2010) Spotlight on Aspergillus nidulans photosensory systems. Fungal Genet Biol 47:900–908. https://doi.org/10.1016/j.fgb.2010.05.008
Blumenstein A, Vienken K, Tasler R, Purschwitz J, Veith D, Frankenberg-Dinkel N, Fisher R (2005) The Aspergillus nidulans phytochrome FphA represses sexual development in red light. Curr Biol 15:1833–1838. https://doi.org/10.1016/j.cub.2005.08.061
Bölker M (2001) Ustilago maydis a valuable model system for the study of fungal dimorphism and virulence. Microbiology 147:1395–1401. https://doi.org/10.1099/00221287-147-6-1395
Brefort T, Doehlemann G, Mendoza-Mendoza A, Reissmann S, Djamei A, Kahmann R (2009) Ustilago maydis as a pathogen. Annu Rev Phytopathol 47:423–445. https://doi.org/10.1146/annurev-phyto.080508-081923
Brych A, Mascarenhas J, Jaeger E, Charkiewicz E, Pokorny R, Bölker M, Doehlemann G, Batschauer A (2015) White collar 1-induced photolyase expression contributes to UV-tolerance of Ustilago maydis. MicrobiologyOpen 5:224–243. https://doi.org/10.1002/mbo3.322
Cabrera-Ponce JL, León-Ramírez CG, Verver-Vargas A, Palma-Tirado L, Ruiz-Herrera J (2012) Metamorphosis of the basidiomycota Ustilago maydis: transformation of yeast-like cells into basidiocarps. Fungal Genet Biol 49:765–771. https://doi.org/10.1016/j.fgb.2012.005
Casas-Flores S, Rios-Momberg M, Bibbins M, Ponce-Noyola P, Herrera-Estrella A (2004) BLR-1 and BLR-2 are key regulatory elements for photoconidiation and mycelial growth in Trichoderma atroviride. Microbiology 150:3561–3569. https://doi.org/10.1099/mic.0.27346-0
Chen CH, Dunlap JC, Loros JJ (2010) Neurospora illuminates fungal photoreception. Fungal Genet Biol 47:922–929. https://doi.org/10.1016/fgb.2010.07.005
Chen CH, Ringelberg CS, Gross RH, Dunlap JC, Loros JJ (2009) Genome-wide analysis of light-inducible responses reveals hierarchical light signaling in Neurospora. EMBO J 28:1029–1042. https://doi.org/10.1038/emboj.2009.54
Cheng P, Yang Y, Gardner KH, Liu Y (2002) PAS domain-mediated WC-1/WC-2 interaction is essential for maintaining the steady-state level of WC-1 and the function of both proteins in circadian clock and light responses of Neurospora. Mol Cell Biol 22:517–524. https://doi.org/10.1128/MCM.22.2.517-524.2002
Collett MA, Garceau N, Dunlap JC, Loros JJ (2002) Light and clock expression of the Neurospora clock gene frequency is differentially driven by but dependent on white collar-2. Genetics 160:149–158
Corrochano LM, Kuo A, Marcet-Houben M et al (2016) Expansion of signal transduction pathways in fungi by extensive genome duplication. Curr Biol 26:1577–1584. https://doi.org/10.1016/j.cub.2016.04.038
Crosson S, Rajagopal S, Moffat K (2003) The LOV domain family: photoresponsive signaling modules coupled to diverse output domains. Biochemistry 42:2–10. https://doi.org/10.1021/bi0269781
Crosthwaite SK, Loros JJ, Dunlap JJ (1995) Light-induced resetting of a circadian clock is mediated by a rapid increase in frequency transcript. Cell 81:1003–1012. https://doi.org/10.1016/S0092-8674(05)80005-4
Dasgupta A, Fuller KK, Dunlap JC, Loros JJ (2016) Seeing the world differently: variability in the photosensory mechanisms of two model fungi. Environ Microbiol 18:5–20. https://doi.org/10.1111/1462-2920.13055
Froehlich AC, Liu Y, Loros JJ, Dunlap JC (2002) White Collar-1, a circadian blue light photoreceptor, binding to the frequency promoter. Science 297:815–819. https://doi.org/10.1126/science.1073681
Fuller KK, Loros JJ, Dunlap JC (2015) Fungal photobiology: visible light as a signal for stress, space and time. Curr Genet 61:275–288. https://doi.org/10.1007/s00294-014-0451-0
Fuller KK, Ringelberg CS, Loros JJ, Dunlap JC (2013) The fungal pathogen Aspergillus fumigatus regulates growth, metabolism, and stress resistance in response to light. MBio 4:e00142–e213. https://doi.org/10.1128/mBio.00142-13
Grimaldi B, Coiro P, Filetici P, Berge E, Dobosy JR, Freitag M, Selker EU, Ballario P (2006) The Neurospora crassa white collar-1 dependent blue light response requires acetylation of histone H3 lysine 14 by NGF-1. Mol Biol Cell 17:4576–4583. https://doi.org/10.1091/mbc.e06-03-0232
Harding RW, Turner RV (1981) Photoregulation of the carotenoid biosynthetic pathway in albino and white collar mutants of Neurospora crassa. Plant Physiol 68:745–749. https://doi.org/10.1104/pp.68.3.745
He Q, Cheng P, Yang Y, Wang L, Gardner KH, Liu Y (2002) White Collar-1, a DNA binding transcription factor and a light sensor. Science 297:840–843. https://doi.org/10.1126/science.1072795
Hedtke M, Rausher S, Röring J, Rodríguez-Romero J, Yu Z, Fisher R (2015) Light-dependent gene activation in Aspergillus nidulans is strictly dependent on phytochrome and involves the interplay of phytochrome and White Collar-regulated histone H3 acetylation. Mol Microbiol 97:733–745. https://doi.org/10.1111/mmi.13062
Hoffman CS, Wriston F (1987) A ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli. Gene 57:267–272. https://doi.org/10.1016/0378-1119(87)90131-4
Holliday R (1974) Ustilago maydis. In: King RC (ed) Handbook of genetics, vol 1. Plenum, New York, pp 575–659
Idnurm A, Heitman J (2005) Light controls growth and development via a conserved pathway in the fungal kingdom. PLoS Biol 3:e95. https://doi.org/10.1371/journal.pbio.0030095
Idnurm A, Rodríguez-Romero J, Corrochano LM, Sanz C, Iturriaga EA, Eslava AP, Heitman J (2006) The Phycomyces madA gene encodes a blue-light photoreceptor for phototropism and other light responses. Proc Natl Acad Sci USA 103:4546–4551. https://doi.org/10.1073/pnas.0600633103
Idnurm A, Verma S, Corrochano LM (2010) A glimpse into the basis of vision in the kingdom Mycota. Fungal Genet Biol 47:881–892. https://doi.org/10.1016/j.fgb.2010.04.009
Igbalajobi O, Yu Z, Fisher R (2019) Red- and blue-light sensing in the plant pathogen Alternaria alternata depends on Phytochrome and the White-Collar protein LreA. MBio 10:e00371–e419. https://doi.org/10.1128/mBio.00371-19
Innocenti FD, Pohl U, Russo VE (1983) Photoinduction of protoperithecia in Neurospora crassa by blue light. Photochem Photobiol 37:49–51. https://doi.org/10.1111/j.1751.1097.1983.tb04432.x
Kämper J, Kahmann R, Bölker M et al (2006) Insights from the genome of the biotrophic fungal plant pathogen Ustilago maydis. Nature 444:97–101. https://doi.org/10.1038/nature05248
Kuratani M, Tanaka K, Terashima K, Muraguchi H, Nakazawa T, Nakahori K, Kamada T (2010) The dst2 gene essential for photomorphogenesis of Coprinopsis cinerea encoded a protein with a putative FAD-binding-4 domain. Fungal Genet Biol 47:152–158. https://doi.org/10.1016/j.fgb.2009.10.006
Lee K, Dunlap JC, Loros JJ (2003) Roles for White Collar-1 in circadian and general photoperception in Neurospora crassa. Genetics 163:103–114
León-Ramírez C, Cabrera-Ponce JL, Martínez-Espinoza A, Herrera-Estrella L, Méndez-Morán L, Reynaga-Peña G, Ruiz-Herrera J (2004) Infection of alternative host plant species by Ustilago maydis. New Phytol 164:337–346. https://doi.org/10.1111/j.1469-8137.2004.01171.x
León-Ramírez C, Cabrera-Ponce JL, Martínez-Soto D, Sánchez-Arreguin A, Aréchiga-Carvajal ET, Ruiz-Herrera J (2017) Transcriptomic analysis of basidiocarp development in Ustilago maydis (DC) Cda. Fungal Genet Biol 101:34–45. https://doi.org/10.1016/j.fgb.2017.02.007
Linden H, Macino G (1997) White collar 2, a partner in blue-light signal transduction, controlling expression of light-regulated genes in Neurospora crassa. EMBO J 16(1):98–109. https://doi.org/10.1093/emboj/16.1.98
Liu Y, He Q, Cheng P (2003) Photoreception in Neurospora: a tale of two White Collar proteins. Cell Mol Life Sci 60:2131–2138. https://doi.org/10.1007/s00018-003-3109-5
Lu YK, Sun KH, Shen WC (2005) Blue light negatively regulates the sexual filamentation via the Cwc1 and Cwc2 proteins in Cryptococcus neoformans. Mol Microbiol 56:480–491. https://doi.org/10.1111/j.1365-2958.2005.04549.x
Martínez-Espinoza DD, García-Pedrajas MD, Gold SE (2002) The Ustilaginales as plant pest and model systems. Fungal Genet Biol 35:1–20. https://doi.org/10.1006/fgbi.2001.1301
Mayorga ME, Gold SE (1998) Characterization and molecular genetic complementation of mutants affecting dimorphism in the fungus Ustilago maydis. Fungal Genet Biol 124:364–376. https://doi.org/10.1006/fgbi.1998.1078
Méndez-Morán L, Reynaga-Peña GC, Springer PS, Ruiz-Herrera J (2005) Ustilago maydis infection of the nonnatural host Arabidopsis thaliana. Phytopathology 95:480–488. https://doi.org/10.1094/PHYTO-95-0480
Möglich A, Yang X, Ayers RA, Moffat K (2010) Structure and function of plant photoreceptors. Annu Rev Plant Biol 61:21–47. https://doi.org/10.1146/annurev-arplant-042809-112259
Morimoto N, Oda Y (1973) Effects of light on fruit-body formation in a basidiomycete Coprinus macrorhizus. Plant Cell Physiol 14:217–225. https://doi.org/10.1093/oxfordjournals.pcp.a074854
Murashige T, Skoog FA (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Ohm RA, Aerts D, Wösten HA, Lugones LG (2013) The blue light receptor complex WC-1/2 of Schizophyllum commune is involved in mushroom formation and protection against phototoxicity. Environ Microbiol 15:943–955. https://doi.org/10.1111/j.1462.2920.2012.02878.x
Purschwitz J, Müller S, Fischer R (2009) Mapping the interactions sites of Aspergillus nidulans phytochrome FphA with the global regulator VeA and the White collar protein LreB. Mol Genet Genomics 281:35–42. https://doi.org/10.1007/s00438-008-0390-x
Purschwitz J, Müller S, Kastner C, Fischer R (2006) Seeing the rainbow: light sensing in fungi. Curr Opin Microbiol 9:566–571. https://doi.org/10.1016/j.mib.2006.10.011
Purschwitz J, Müller S, Kastner C, Schöser M, Haas H, Espeso EA, Atoui A, Calvo AM, Fischer R (2008) Functional and physical interaction of blue and red-light sensors in Aspergillus nidulans. Curr Biol 18:255–259. https://doi.org/10.1016/j.cub.2008.01.061
Rodríguez-Romero J, Hedtke M, Kastner C, Müller S, Fisher R (2010) Fungi, hidden in soil or up in the air: light makes a difference. Annu Rev Microbiol 64:585–610. https://doi.org/10.1146/annurev.micro.112408.13400
Ruiz-Herrera J (2012) Dimorphic fungi: their importance as models for differentiation and fungal pathogenesis. Bentham eBooks, Sharjah (ISBN:978-1-60805-364-3)
Sanz C, Rodríguez-Romero J, Idnurm A, Christie JM, Heitman J, Corrochano LM, Eslava AP (2009) Phycomyces MADB interacts with MADA to form the primary photoreceptor complex for fungal phototropism. Proc Natl Acad Sci USA 106:7095–7100. https://doi.org/10.1073/pnas.0900879106
Silva F, Torres-Martínez S, Garre V (2006) Distinct white collar-1 genes control specific light responses in Mucor circinelloides. Mol Microbiol 6:1023–1037. https://doi.org/10.1111/j.1365-2958.2006.05291.x
Taylor BL, Zhulin IB (1998) In search of higher energy: metabolism-dependent behavior in bacteria. Mol Microbiol 28:683–690. https://doi.org/10.1046/j.1365-2958.1998.00835.x
Taylor BL, Zhulin IB (1999) PAS domains: internal sensors of oxygen, redox potential, and light. Microbiol Mol Biol Rev 63:479–506
Terashima K, Yuki K, Muraguchi H, Akiyama M, Kamada T (2005) The dst1 gene involved in mushroom photomorphogenesis of Coprinus cinereus encodes a putative photoreceptor for blue light. Genetics 171:101–108. https://doi.org/10.1534/genetics.104.040048
Tsukuda T, Carleton S, Fotheringham S, Holloman WK (1988) Isolation and characterization of an autonomously replicating sequence from Ustilago maydis. Mol Cell Biol 8:3703–3709. https://doi.org/10.1128/MCB.8.9.3703
Verma S, Idnurm A (2013) The Uve1 endonuclease is regulated by the white collar complex to protect Cryptococcus neoformans from UV damage. PLoS Genet 9:e1003769. https://doi.org/10.1371/journal.pgen.1003769
Vollmeister E, Schipper K, Baumann S, Haag C, Pohlmann T, Stock J, Feldbrügge M (2012) Fungal development of the plant pathogen Ustilago maydis. FEMS Microbiol Rev 36:59–77. https://doi.org/10.1111/j.1574-6976.2011.00296.x
Wang Z, Li N, Dunlap JC, Trail F, Townsend JP (2016) The fast-evolving phy-2 gene modulates sexual development in response to light in the model fungus Neurospora crassa. mBio 7:e02148. https://doi.org/10.1128/mbio.02148-15
Yu JH, Hamari Z, Han KH, Seo JA, Reyes-Dominguez Y, Scazzocchio C (2004) Double-joint PCR: a PCR-based molecular tool for gene manipulations in filamentous fungi. Fungal Genet Biol 41:973–981. https://doi.org/10.1016/j.fgb.2004.08.001
Zhu P, Idnurm A (2018) The contribution of the White Collar complex to Cryptococcus neoformans virulence is independent of its light-sensing capabilities. Fungal Genet Biol 121:56–64. https://doi.org/10.1016/j.fgb.2018.09.008
Acknowledgements
This work was partially supported by Consejo Nacional de Ciencia y Tecnología (CONACYT), México. MOCE was a Ms.C. student with a fellowship from CONACYT. Thanks are given to Miss M. F. Salazar-Chávez for her technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare no conflict of interest regarding this manuscript.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
203_2019_1725_MOESM2_ESM.pptx
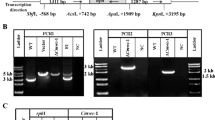
Fig. S1 Strategy and results of deletion of WCO1a, WCO1b, and WCO2 genes in U. maydis. Genes were deleted from the U. maydis wild-type strain FB1 and FB2 by double homologous gene recombination. Only data for FB1 strain are shown; similar results were obtained for FB2 strain. A) Schematic representations of gene deletion strategy. HPH, hygromycin phosphotransferase genes, WCO1a, WCO1b and WCO2, WC gene homologues. Numbers 1-6, primers used (see Table S1). Notice that primers 1, 2, 3, and 6 are outside of the deletion cassettes. B) Results of the confirmation of gene deletion. I, II, and III. Gel electropherograms of the products of the amplicons obtained with the oligonucleotides shown above that confirm the deletion at the loci of WCO1a, WCO1b, and WCO2, respectively. I, lanes 1–5 (a and b), wco1a mutants; lane 6a, and 6b, wt. II, lane 1, wt; lanes 2, 3, wco1b mutants. III, lanes 1–6 (a and b), wco2 mutants; lane 7a and 7b, wt (PPTX 385 kb)
Rights and permissions
About this article
Cite this article
Sánchez-Arreguin, J.A., Cabrera-Ponce, J.L., León-Ramírez, C.G. et al. Analysis of the photoreceptors involved in the light-depending basidiocarp formation in Ustilago maydis. Arch Microbiol 202, 93–103 (2020). https://doi.org/10.1007/s00203-019-01725-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-019-01725-w