Abstract
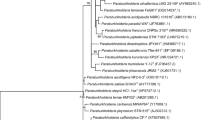
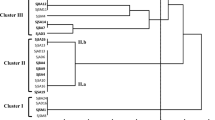
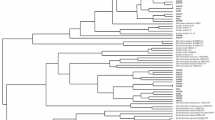
Eleven extra-slow-growing strains were isolated from nodules of the relict legume Vavilovia formosa growing in North Ossetia (Caucasus) and Armenia. All isolates formed a single rrs cluster together with the type strain Tardiphaga robiniae LMG 26467T, while the sequencing of the 16S–23S rDNA intergenic region (ITS) and housekeeping genes glnII, atpD, dnaK, gyrB, recA and rpoB divided them into three groups. North Ossetian isolates (in contrast to the Armenian ones) were clustered separately from the type strain LMG 26467T. However, all isolates were classified as T. robiniae because the DNA–DNA relatedness between them and the type strain LMG 26467T was 69.6 % minimum. Two symbiosis-related genes (nodM and nodT) were amplified in all isolated Tardiphaga strains. It was shown that the nodM gene phylogeny is similar to that of ITS and housekeeping genes. The presence of the other symbiosis-related genes in described Tardiphaga strains, which is recently described genus of rhizobia, as well as their ability to form nodules on any plants are under investigation.




Similar content being viewed by others
References
Bhagwat AA, Mithöfer A, Pfeffer PE, Kraus C, Spickers N, Hotchkiss A, Ebel J, Keister DL (1999) Further studies of the role of cyclic beta-glucans in symbiosis. An NdvC mutant of Bradyrhizobium japonicum synthesizes cyclodecakis-(1–>3)-beta-glucosyl. Plant Physiol 119:1057–1064. doi:10.1104/pp.119.3.1057
Chaintreuil C, Arrighi JF, Giraud E, Miché L, Moulin L, Dreyfus B, Munive-Hernández JA, Villegas-Hernandez Mdel C, Béna G (2013) Evolution of symbiosis in the legume genus Aeschynomene. New Phytol 200:1247–1259. doi:10.1111/nph.12424
De Meyer SE, Willems A (2012) Multilocus sequence analysis of Bosea species and description of Bosea lupini sp. nov., Bosea lathyri sp. nov. and Bosea robiniae sp. nov., isolated from legumes. IJSEM 62:2505–2510. doi:10.1099/ijs.0.035477-0
De Meyer SE, Coorevits A, Willems A (2012) Tardiphaga robiniae gen. nov., sp. nov., a new genus in the family Bradyrhizobiaceae isolated from Robinia pseudoacacia in Flanders (Belgium). Syst App Microbiol 35:205–214. doi:10.1016/j.syapm.2012.02.002
Dekker L, Osborne TH, Santini JM (2014) Isolation and identification of cobalt- and caesium-resistant bacteria from a nuclear fuel storage pond. FEMS Microbiol Lett 359:81–84. doi:10.1111/1574-6968.12562
Ding H, Yip CB, Geddes BA, Oresnik IJ, Hynes MF (2012) Glycerol utilization by Rhizobium leguminosarum requires an ABC transporter and affects competition for nodulation. Microbiology 158:1369–1378. doi:10.1099/mic.0.057281-0
Giraud E, Moulin L, Vallenet D, Barbe V, Cytryn E, Avarre JC, Jaubert M, Simon D, Cartieaux F, Prin Y, Bena G, Hannibal L, Fardoux J, Kojadinovic M, Vuillet L, Lajus A, Cruveiller S, Rouy Z, Mangenot S, Segurens B, Dossat C, Franck WL, Chang WS, Saunders E, Bruce D, Richardson P, Normand P, Dreyfus B, Pignol D, Stacey G, Emerich D, Verméglio A, Médigue C, Sadowsky M (2007) Legumes symbioses: absence of Nod genes in photosynthetic bradyrhizobia. Science 316:1307–1312. doi:10.1126/science.1139548
Goris J, Suzuki K-I, De Vos P, Nakase T, Kersters K (1998) Evaluation of a microplate DNA-DNA hybridisation method compared with the initial renaturation method. Can J Microbiol 44:1148–1153. doi:10.1139/cjm-44-12-1148
Guo HJ, Wang ET, Zhang XX, Li QQ, Zhang YM, Tian CF, Chena WX (2014) Replicon-dependent differentiation of symbiosis-related genes in Sinorhizobium strains nodulating Glycine max chromosome. Appl Environ Microbiol 80:1245–1255. doi:10.1128/AEM.03037-13
Kenicer G, Smýkal P, Vishyakova M, Mikić A (2009) Vavilovia formosa, an intriguing Pisum relative. Grain Legumes 51:8–12. http://www.ias.csic.es/grainlegumesmagazine/Grain_Legumes_issue_51.pdf
Lohrke SM, Day B, Kolli VS, Hancock R, Yuen JP, de Souza ML, Stacey G, Carlson R, Tong Z, Hur H-G, Orf JH, Sadowsky MJ (1998) The Bradyrhizobium japonicum noeD gene: a negatively acting, genotype-specific nodulation gene for soybean. Mol Plant-Microbe Interact 11:476–488. doi:10.1094/MPMI.1998.11.6.476
Menna P, Barcellos FG, Hungria M (2009) Phylogeny and taxonomy of a diverse collection of Bradyrhizobium strains based on multilocus sequence analysis of the 16S rRNA gene, ITS region and glnII, recA, atpD and dnaK genes. Int J Syst Evol Microbiol 59:2934–2950. doi:10.1099/ijs.0.009779-0
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G + C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167. doi:10.1099/00207713-39-2-159
Mikić A, Smýkal P, Kenicer G Vishnyakova M, Akopian J, Sarukhanyan N, Gabrielyan I, Vanyan A, Toker C, Ćupina B, Ambrose M, Mihailović V, Ellis N (2009) A revival of the research on beautiful vavilovia (Vavilovia formosa syn. Pisum formosum). Pisum Genet 41:34–39. http://pisum.narod.ru/pg/41/34.pdf
Mikić A, Smýkal P, Kenicer G, Vishnyakova M, Sarukhanyan N, Akopian JA, Vanyan A, Gabrielyan I, Smýkalová I, Sherbakova E, Zorić L, Atlagić J, Zeremski-Škorić T, Cupina B, Krstić D, Jajić I, Antanasović S, Dorđević V, Mihailović V, Ivanov A, Ochatt S, Toker C, Zlatković B, Ambrose M (2014) Beauty will save the world, but will the world save beauty? The case of the highly endangered Vavilovia formosa (Stev.) Fed. Planta 240:1139–1146. doi:10.1007/s00425-014-2136-9
Mornico D, Miché L, Béna G, Nouwen N, Verméglio A, Vallenet D, Smith AA, Giraud E, Médigue C, Moulin L (2012) Comparative genomics of aeschynomene symbionts: insights into the ecological lifestyle of nod-independent photosynthetic bradyrhizobia. Genes 3:35–61. doi:10.3390/genes3010035
Novikova N, Safronova V (1992) Transconjugants of Agrobacterium radiobacter harbouring sym genes of Rhizobium galegae can form an effective symbiosis with Medicago sativa. FEMS Microbiol Lett 93:261–268. doi:10.1111/j.1574-6968.1992.tb05107.x
Okubo T, Fukushima S, Itakura M, Oshima K, Longtonglang A, Teaumroong N, Mitsui H, Hattori M, Hattori R, Hattori T, Minamisawa K (2013) Genome analysis suggests that the soil oligotrophic bacterium Agromonas oligotrophica (Bradyrhizobium oligotrophicum) is a nitrogen-fixing symbiont of Aeschynomene indica. Appl Environ Microbiol 79:2542–2551. doi:10.1128/AEM.00009-13
Pinto FG, Chueire LM, Vasconcelos AT, Nicolás MF, Almeida LG, Souza RC, Menna P, Barcellos FG, Megías M, Hungria M (2009) Novel genes related to nodulation, secretion systems, and surface structures revealed by a genome draft of Rhizobium tropici strain PRF 81. Funct Integr Genomics 9:263–270. doi:10.1007/s10142-009-0109-z
Rivilla R, Sutton JM, Downie JA (1995) Rhizobium leguminosarum NodT is related to a family of outer-membrane transport proteins that includes TolC, PrtF, CyaE and AprF. Gene 161:27–31. doi:10.1016/0378-1119(95)00235-X
Rosselló-Móra R (2006) DNA–DNA reassociation methods applied to microbial taxonomy and their critical evaluation. In: Stackebrandt E (ed) Molecular identification, systematics, and population structure of prokaryotes. Springer, New York, pp 23–50
Safronova V, Tikhonovich I (2012) Automated cryobank of microorganisms: unique possibilities for long-term authorized depositing of commercial microbial strains. In: Mendez-Vilas A (ed) Microbes in applied research: current advances and challenges, World Scientific Publishing Co, pp 331–334. doi:10.1142/9789814405041_0066
Safronova VI, Piluzza G, Belimov AA, Bullitta S (2004) Phenotypic and genotypic analysis of rhizobia isolated from pasture legumes native of Sardinia and Asinara Island. Antonie Van Leeuwenhoek 85:115–127. doi:10.1023/B:ANTO.0000020278.58236.77
Safronova VI, Kimeklis AK, Chizhevskaya EP, Belimov AA, Andronov EE, Pinaev AG, Pukhaev AR, Popov KP, Tikhonovich IA (2014) Genetic diversity of rhizobia isolated from nodules of the relic species Vavilovia formosa (Stev.) Fed. Antonie Van Leeuwenhoek 105:389–399. doi:10.1007/s10482-013-0089-9
Safronova VI, Kuznetsova IG, Sazanova AL, Kimeklis AK, Belimov AA, Andronov EE, Pinaev AG, Chizhevskaya EP, Pukhaev AR, Popov KP, Willems A, Tikhonovich IA (2015) Bosea vaviloviae sp. nov. a new species of slow-growing rhizobia isolated from nodules of the relict species Vavilovia formosa (Stev.) Fed. Antonie Van Leeuwenhoek 107:911–920. doi:10.1007/s10482-015-0383-9
Sinjushin AA, Demidenko NV, Gostimskii SA (2009) Preliminary report on taxonomical position of Vavilovia formosa (Stev.) Fed. evidenced from morphological and molecular data. Pisum Genetics 41:15–20. http://pisum.narod.ru/pg/41/15.pdf
Stepkowski T, Moulin L, Krzyzanska A, McInnes A, Law IJ, Howieson J (2005) European origin of Bradyrhizobium populations infecting lupins and serradella in soils of Western Australia and South Africa. Appl Environ Microbiol 71:7041–7052. doi:10.1128/AEM.71.11.7041-7052.2005
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parismony methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121
Tikhonovich IA, Provorov NA (2007) Cooperation of plants and microorganisms: getting closer to the genetic construction of sustainable agro-systems. Biotechnol J 2:833–848. doi:10.1002/biot.200700014
van Workum WA, Canter Cremers HC, Wijfjes AH, van der Kolk C, Wijffelman CA, Kijne JW (1997) Cloning and characterization of four genes of Rhizobium leguminosarum bv. trifolii involved in exopolysaccharide production and nodulation. Mol Plant Microbe Interact 10:290–301. doi:10.1094/MPMI.1997.10.2.290
Vincent JM (1970) A manual for the practical study of root nodule bacteria. IBP Handbook. Blackwell Scientific Publications, Oxford and Edinburgh, pp 73–97
Zakhia F, Jeder H, Willems A, Gillis M, Dreyfus B, de Lajudie P (2006) Diverse bacteria associated with root nodules of spontaneous legumes in Tunisia and first report for nifH-like gene within the genera Microbacterium and Starkeya. Microb Ecol 51:375–393. doi:10.1007/s00248-006-9025-0
Acknowledgments
We thank Pia Clercx for performing DNA–DNA hybridizations and excellent technical assistance. This work was partially supported by the Ministry of Education and Sciences of the Russian Federation (Agreement No. 14.604.21.0024, RFMEFI60414X0024) and by the European Union’s Seventh Framework Programme (BRIO project, grant KBBE 2010-4-266106). The nodM and nodT sequencing analysis was supported by the Russian Science Foundation (Grant No. 14-26-00094).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Safronova, V.I., Kuznetsova, I.G., Sazanova, A.L. et al. Extra-slow-growing Tardiphaga strains isolated from nodules of Vavilovia formosa (Stev.) Fed.. Arch Microbiol 197, 889–898 (2015). https://doi.org/10.1007/s00203-015-1122-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-015-1122-3