Abstract
Aims/hypothesis
Absent in melanoma 2 (AIM2) is a cytosolic sensor for double-stranded DNA and a tumour suppressor. Binding of double-stranded DNA to AIM2 forms the AIM2 inflammasome, leading to activation of caspase-1 and production of IL-1β and IL-18. Although inflammasome-independent effects of AIM2 have been reported, its role in energy metabolism is unknown. We aimed to evaluate the effect of AIM2 in energy metabolism and glucose homeostasis.
Methods
Male and female whole body Aim2 knockout (Aim2−/−) mice were used in the current study. Body weight, food intake, body composition, energy expenditure, fasting blood glucose levels, GTT and body temperature were measured at indicated time points. RNA sequencing was carried out on gonadal white adipose tissue (gWAT) in 14-month-old female mice. mRNA and protein levels in tissues were analysed by quantitative real-time PCR and immunoblot. Immune cell infiltration in gWAT was examined by flow cytometry. Stromal vascular fractions isolated from gWAT were used to investigate adipocyte differentiation.
Results
Male and female Aim2−/− mice were obese compared with wild-type controls from 7 weeks of age until 51 weeks of age, with increased adiposity in both subcutaneous and visceral fat depots. While there were no differences in food intake, Aim2−/− mice demonstrated decreased energy expenditure and impaired brown adipose tissue function compared with wild-type controls. Fasting glucose and insulin levels were elevated, and Aim2−/− mice were glucose intolerant on intraperitoneal GTT. RNA sequencing revealed marked upregulation of the IFN-inducible gene Ifi202b, which encodes protein 202 (p202) and elevated inflammatory signalling in gWAT of Aim2−/− mice. Increased infiltration of total and Ly6Clow monocytes was noted at 8 weeks of age in gWAT, before the onset of obesity and insulin resistance. Ifi202b knockdown blocked adipogenesis in stromal vascular fractions and reduced inflammation in bone marrow-derived macrophages, demonstrating a key role of p202 in mediating the increased adipogenesis and inflammation in Aim2−/− mice.
Conclusions/interpretation
These results demonstrate a fundamental role for AIM2 in energy metabolism, inflammation and insulin resistance. Our studies establish a novel link between the innate immunity proteins, AIM2 and p202, and metabolism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction

The prevalence of obesity has markedly increased worldwide in recent decades. Fat mass expansion is mediated through increased adipocyte number (hyperplasia), adipocyte size (hypertrophy) or both [1]. Adipogenesis (cause of hyperplasia) is mainly governed by the transcription factors peroxisome proliferator activated receptor γ (PPARγ) and the CCAAT/enhancer binding protein (C/EBP) α, and downstream signalling pathways [2, 3]. Decreased capacity for de novo adipogenesis coupled with an increased demand for NEFA storage in obesity leads to a switch from adipose tissue hyperplasia to hypertrophy, which contributes to insulin resistance [4].
Substantial evidence links obesity and chronic inflammation [5]. Obesity is associated with the infiltration of immune cells into the white adipose tissue (WAT), including macrophages [6, 7], dendritic cells [8], neutrophils [9, 10], and T and B cells [11,12,13,14]. WAT is a source of proinflammatory cytokines, including IL-6 and TNF-α, and adipokines such as leptin and adiponectin [15]. Adipose tissue immune cells produce cytokines, metalloproteinases and reactive oxygen species, causing adipose tissue dysfunction and insulin resistance [16]. The increase in immune cell infiltration is secondary to an increase in expression of chemokines in the WAT of obese individuals [17]. These chemokines, secreted from mature adipocytes, promote infiltration of monocytes in WAT, which differentiate into adipose tissue macrophages [4]. For instance, expression of fractalkine, also known as CX3CL1, is markedly upregulated in obese human adipose tissue [18] and has been defined as an adipochemokine associated with type 2 diabetes [19].
Inflammasomes are multiprotein oligomers and an integral component of the innate immune system. They serve as receptors and sensors in the pathogen-induced innate immune response and regulate the activation of caspase-1 through interaction with apoptosis-associated speck-like protein containing a caspase activation and recruitment domain (ASC), an inflammasome adaptor protein [20]. Upon inflammasome activation, IL-1β and IL-18 are released through caspase-1 activation and subsequent cleavage of pro-IL-1β and IL-18 [21]. Ablation of key components of the inflammasome, such as NOD-like receptors (NLRs) including NLRP1 and NLRP3, ASC, caspase-1 and IL-1β, protects mice from high-fat diet (HFD)-induced obesity and insulin resistance [22,23,24]. Neutralisation of IL-1β signalling using IL-1β antibody significantly improves glycaemic control and β-cell function in HFD-induced obesity and insulin resistance [25], indicating a role of inflammasomes in glucose homeostasis.
Absent in melanoma 2 (AIM2) belongs to the IFN-inducible HIN200 domain-containing protein family [26,27,28]. It is a tumour suppressor and receptor of cytosolic double-stranded DNA [29, 30]. The AIM2 inflammasome has been reported to play a role in viral infections [30,31,32,33]. In addition, the AIM2 inflammasome is also well characterised in autoimmune and inflammatory diseases such as dermatitis and arthritis and cancers [34, 35]. A role of AIM2 in atherosclerosis through regulation of vascular muscle cell migration and inflammation [36, 37] was reported. To date, a direct role of AIM2 in metabolism has not yet been reported. Here, we aimed to explore the role of AIM2 in energy metabolism and glucose homeostasis using Aim2 knockout (Aim2−/−) mice.
Methods
Mice
C57Bl6/J wild-type (WT), Aim2−/− (B6.129P2-Aim2Gt(CSG445)Byg/J) and New Zealand Obese (NZO) mice were purchased from The Jackson Laboratory (Bar Harbor, ME, USA) and bred in the animal facility at the Children’s Hospital of Pittsburgh of UPMC, University of Pittsburgh School of Medicine. Aim2−/− mice were backcrossed to C57Bl6/J for at least seven generations. All mice (4–5 per cage) were housed under a standard 12 h light/dark cycle (lights on at 07:00) with access to food and water ad libitum unless otherwise stated. For HFD-fed mice, C57Bl6/J mice were fed with a 60% HFD (Research Diets, New Brunswick, NJ, USA) for 4 months. Body weight and food intake were assessed weekly from 7–8 weeks of age (male and female, n = 8). Body composition of mice, including fat mass and lean mass, was determined using the EchoMRI-100 system (Echo Medical Systems, Houston, TX, USA). Gonadal, perirenal and brown adipose tissue (BAT) fat pads were dissected and weighed after completion of the study. All animal studies were approved by the University of Pittsburgh Institutional Animal Care and Use Committee.
CLAMS
Energy expenditure (EE), respiratory exchange ratio (RER), v̇O2, CO2 production and total locomotor activity were assessed by the Comprehensive Lab Animal Monitoring System (CLAMS) at room temperature (22 ± 0.5°C). In brief, mice were pre-housed in CLAMS for 24 h for acclimatisation, and then data were recorded for 2 days with ad libitum access to food and water. EE data were normalised to body weight. Linear regression was performed to correlate EE to lean body mass (LBM) [38].
Fasting glucose levels and IPGTT
Fasting glucose levels were measured after overnight (16 h) fasting. For IPGTT, mice were fasted for 6 h. After estimation of blood glucose, mice were injected with 2 g/kg glucose intraperitoneally. Blood glucose levels were then recorded at 15, 30, 60 and 120 min post glucose injection.
Biochemical measurements
Blood glucose was measured by glucometer (FreeStyle, Abbott Park, IL, USA) and fasting serum insulin levels were determined by UltraSensitive Mouse Insulin ELISA Kit (Crystal Chem, Elk Grove Village, IL, USA). Serum triacylglycerols (TGs) were measured using commercial kits from Thermo Fisher Scientific (Waltham, MA, USA).
H&E staining
Gonadal WAT (gWAT) and interscapular BAT were isolated from the mice, fixed in 10% neutral buffered formalin solution overnight and transferred into 70% ethanol solution. Tissues were then embedded in paraffin for sectioning (5 μm sections) and stained with H&E.
Adipocyte size and number quantification
Quantification of adipocytes in gWAT was done using ImageJ (version 1.48 V; https://imagej.nih.gov/ij/download.html). Images were processed as described previously [39]. Adipocytes were manually selected, and diameters were measured by ImageJ. The frequency of each diameter range was calculated as the percentage of cell counts in the range over the total cell counts.
Isolation and culture of primary stromal vascular fractions
gWAT from 3–4-month-old WT and Aim2−/− mice was dissected, minced into small pieces and digested in type I collagenase for 45 min at 37°C with agitation. The digestion was stopped by adding 10 ml growth medium containing 10% FBS and 1% penicillin and streptomycin in DMEM. The cell mixture was passed through a 70 μm cell strainer and then centrifuged for 10 min at 250 g. The pellets were collected, washed three times with growth medium and then seeded on 10 cm dishes. Medium was changed every other day.
Adipocyte differentiation and BODIPY staining
The stromal vascular fractions (SVFs) were plated in 12 or 24 well plates and grown until full confluence. Two days after confluence, adipocyte differentiation was induced by adding differentiation medium containing dexamethasone (2 μg/ml), 3-isobutyl-1-methylxanthine (IBMX, 0.5 mmol/l) and insulin (2 μg/ml) to the cells for 3 days, and changing to the growth medium for another 4 days. For boron dipyrromethene (BODIPY) staining, differentiated adipocytes were washed with PBS twice and fixed by 4% paraformaldehyde. After two rinses with PBS, cells were incubated with PBS containing 1 μg/ml of BODIPY at room temperature for 15 min in the dark. Cells were then rinsed with PBS twice and images taken using EVOS FL Color Imaging System (Life Technologies, Waltham, MA, USA).
Body temperature
Body temperature of mice at basal level and under cold challenge conditions (8°C for 4 h) was assessed by rectal temperature probe and digital thermometer (Physitemps Instruments, Clifton, NJ, USA).
Fatty acid oxidation in BAT
14C-palmitate was conjugated to BSA overnight and used at a final concentration of 125 μmol/l in 200 μl reactions containing 100 mmol/l sucrose, 10 mmol/l Tris-HCl pH 7.4, 5 mmol/l KH2PO4, 0.2 mmol/l EDTA, 80 mmol/l KCl, 1 mmol/l MgCl2, 2 mmol/l l-carnitine, 0.05 mmol/l coenzyme-A, 2 mmol/l ATP, 1 mmol/l dithiothreitol and approximately 15 μg freshly prepared interscapular BAT homogenate. Filter papers soaked in 1 mol/l KOH were embedded into the caps of the reaction tubes. Reaction tubes were gently rotated in a 37°C water bath for 1 h, then reactions were terminated by addition of perchloric acid to 0.5 mol/l. The released 14CO2 was captured over a second hour of incubation at 37°C. 14CO2 captured on the filter papers was subjected to scintillation counting and the data normalised to protein content.
RNA extraction, RNA sequencing and real-time PCR
Tissue or cell RNA was extracted using RNeasy purification kit (Qiagen, Valencia, CA, USA) following the manufacturer’s instructions. First strand cDNA was synthesised from 1 μg RNA using iScript cDNA Synthesis Kit (Bio-Rad, Hercules, CA, USA). RNA sequencing (RNAseq) was performed at the Genomic Core at the University of Pittsburgh. The quality of the RNAseq data was assessed by FastQC (Version 0.11.6; www.bioinformatics.babraham.ac.uk/projects/fastqc/) and aligned to the reference genome using TopHat2 (Version 2.1.1; http://ccb.jhu.edu/software/tophat). RNAseq reads with low sequencing quality or mapping quality were filtered out. Gene expression counts were generated with featureCounts (Version 1.6.1; http://subread.sourceforge.net/) based on the results from TopHat2. Real-time PCR was carried out using TaqMan assays in a 10 μl reaction mixture (Bio-Rad) containing 0.1 μl first strand cDNA and 1X probe and primers mix (Bio-Rad). Probes were purchased from Life Technologies or Bio-Rad (see ESM Table 1 for details). Relative mRNA levels were calculated by \( {2}^{-\varDelta \varDelta {\mathrm{C}}_{\mathrm{t}}} \) and normalised to Gapdh.
Immunoblotting
Cells and tissues were homogenised in RIPA buffer (50 mmol/l Tris [pH 7.4], 150 mmol/l NaCl, 1% Triton X-100, 0.5% SDS) containing proteinase inhibitors (Roche, Indianapolis, IN, USA) and phosphatase inhibitors. Total protein (30 μg) was resolved on SDS-PAGE, then transferred to PVDF membranes. The membranes were blocked with 5% non-fat dry milk in Tris-buffered saline (154 mmol/l NaCl) with Tween 20 (TBS-T) for 1 h at room temperature. The membranes were incubated with primary antibodies (listed in ESM Table 2; diluted in 5% BSA in TBS-T) overnight at 4°C. The membranes were then washed three times with TBS-T and incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies (diluted in 5% non-fat dry milk in TBS-T) for an additional 1 h at room temperature. Images were taken after adding SuperSignal West Dura extended duration substrate (Thermo Fisher Scientific) to the membrane. Antibodies were pre-validated by molecular mass using positive control samples.
Bio-Plex
Serum was used to examine circulating leptin, TNF-α, IFN-γ, IL-6, monocyte chemoattractant protein 1 (MCP-1), IL-1α and IL-1β using Bio-Plex (Bio-Rad) according to the manufacturer’s instructions.
Caspase-1 activity assay
Total protein lysates were extracted from the adipose tissue, and caspase-1 activity was assessed using Caspase-1 Colorimetric Assay Kit (BioVision, Milpitas, CA, USA) according to manufacturer’s instructions.
Human studies
The study was approved by the Institutional Review Board of the University of Pittsburgh. Informed consent was obtained from both parents, and assent was obtained from participating children. Subcutaneous adipose tissue samples were collected from a total of 33 participants aged 6–17 years (14 [42.4%] female, mean age 12.5 years). After total RNA extraction, sequencing libraries were generated with Illumina TruSeq Stranded Total RNA Ribo-Zero H/M/R kits (Illumina, San Diego, CA, USA). Libraries were normalised and run on a NextSeq 500 sequencer (Illumina) at 75 cycles with 40 million reads per sample. After quality control, 3′ adapter trimming and low-quality read filtering, RNAseq data were aligned to the reference human genome using TopHat2. Transcripts per kb per million (TPM) were used as proxy for gene expression level. Analysis was performed using a regression framework in DESeq2 (Version 1.24.0; https://bioconductor.org/packages/release/bioc/html/DESeq2.html) for either BMI percentile or overweight/obesity, adjusting for age and sex.
Isolation and culture of bone marrow-derived macrophages
Both femurs were harvested and flushed through both ends on a cell strainer (filter) with a 25-gauge needle attached to a 10 ml syringe. With a syringe plunger, flushed femur was minced on the cell strainer and washed with DMEM until all of the cells passed through the filter. After spinning down the cells, supernatant was carefully discarded, the pellet was washed twice with growth medium and cells were cultured in growth medium containing 20 ng/ml granulocyte-macrophage colony-stimulating factor (GM-CSF) for at least 4 days before experiments.
Flow cytometry
Two-month-old female WT and Aim2−/− mice were euthanised and perfused thoroughly with 30 ml ice-cold PBS through the left ventricle to remove blood from solid organs. gWAT was perfused, harvested and digested in FACS buffer (PBS containing 0.5% BSA) containing 1 mg/ml collagenase IV and 20 μl HEPES at 37°C at 120 g for 45 min. Cells were passed through a 40 μm cell strainer, washed in 20 ml FACS buffer and centrifuged (4°C, 350 g, 7 min). Erythrocytes were lysed using ammonium-chloride-potassium (ACK) lysis buffer for 3 min, and 3 ml FACS buffer was added and centrifuged. Bone marrow harvested as stated above was washed and resuspended in FACS buffer. Total viable cell numbers were obtained from the above aliquots using trypan blue (Cellgro, Corning, NY, USA). Following the harvesting of single-cell suspensions, cells were stained in FACS buffer. All antibodies used in this study were purchased from eBioscience (San Diego, CA, USA), BioLegend (San Diego, CA, USA) and BD Biosciences (San Jose, CA, USA). For mature myeloid cell analysis, monoclonal antibodies including anti-CD11b (M1/70), Ly6G (1A8), CD115 (AFS98) and Ly6C (AL-21) were used. Neutrophils were identified as CD11b+ Ly6G+, Ly6Clow monocytes were identified as CD11b+ CD115+ Ly6Clow and Ly6Chigh monocytes were defined as CD11b+ CD115+ Ly6Chigh. Macrophages were identified as CD45+ CD11b+ CD64+ F4/80+. T cells were identified as CD19− CD3+ and B cells were identified as CD19+. Data acquisition was performed using Fortessa Flow Cytometer (BD Biosciences). Data were analysed using FlowJo software (Tree Star, Ashland, OR, USA).
Ifi202b RNAi
For knockdown of the IFN activated gene Ifi202b, which encodes protein 202 (p202), in primary SVF and bone marrow-derived macrophages (BMMs), small interfering RNA (siRNA) against mouse Ifi202b (SMARTpool) or scrambled siRNA (Dharmacon, Lafayette, CO, USA) was transfected into primary cells using Neon Electroporation (Life Technologies). For SVF, 50 nmol/l siRNA was transfected into 3 × 105 cells (1150 V, 20 ms, two pulses), and cells were seeded on 24 well plates for 3 days before being induced for differentiation using differentiation medium containing insulin, IBMX and dexamethasone. For BMMs, 50 nmol/l siRNA was transfected into 5 × 105 cells, seeded on 24 well plates and incubated in complete medium containing 20 nmol/l GM-CSF before stimulating with lipopolysaccharide (LPS). Cells were harvested 48–72 h after transfection and knockdown of Ifi202b was examined by real-time PCR and western blotting.
Statistical analysis
There was no criteria for exclusion of any data. The in vivo studies were done once with at least 5–6 animals in each group, and in vitro studies were done at least 3 times, with 2–3 replicates. Experimenters were blind to group assignment and outcome assessment in the in vitro studies and endpoint assays, such as real-time PCR, immunoblotting and Bio-Plex assays. All values shown are presented as means ± SEM. Independent two-tailed t test was used when comparing two groups between WT and Aim2−/− at a single time point. Two-way ANOVA was used when dynamic comparisons were made among variables. For each statistically significant F value observed for the main effect or interaction, a two-tailed post hoc test (Tukey’s) was applied to determine individual differences between means. Difference was considered to be statistically significant when p ≤ 0.05.
Results
Aim2 deletion causes spontaneous obesity in mice
With ad libitum access to chow diet and water, both female (Fig. 1a) and male (ESM Fig. 1a) Aim2−/− mice were larger than their WT counterparts from 7 to 51 weeks of age (n = 7–8, Fig. 1b and ESM Fig. 1b) without significant change in food intake (Fig. 1c, d and ESM Fig. 1c). At 9 weeks of age, Aim2−/− mice had similar fat mass to controls in both sexes (Fig. 1e and ESM Fig. 1d), while female Aim2−/− mice had significantly higher LBM (Fig. 1f and ESM Fig. 1e). At 15 weeks, both female and male Aim2−/− mice had significantly higher fat mass than controls (Fig. 1e and ESM Fig. 1d). The increased fat mass persisted throughout the study. LBM showed no significant differences at 15, 22 or 32 weeks of age, indicating that differences in body weight from 15 weeks of age were entirely attributable to increased fat mass.
Aim2 deletion induces spontaneous obesity and impaired glucose homeostasis. (a) Representative images for 14-month-old female WT and Aim2−/− mice. (b) Body weight of female WT and Aim2−/− mice from 7 to 51 weeks of age. The vertical dashed line represents the missing data for week of 34 and 35. (c, d) Food intake (38–51 weeks; (c) and mean food intake (7–51 weeks; (d) of the mice. The key in (b) also applies to (c). (e, f) Fat and lean mass of female WT and Aim2−/− mice at 9, 15, 22 and 32 weeks of age. The key in (e) also applies to (f). (g–j) CLAMS was used to monitor EE, RER and locomotor activity in female WT and Aim2−/− mice over a 48 h period: heat production (EE) over 48 h (mean EE [kJ [g body weight]−1 h−1] in light and dark phases shown in the inset) (g); regression analysis of the relationship between 24 h total EE and LBM (h); RER in female WT and Aim2−/− mice (mean RER in light and dark phases shown in the inset) (i); and total activity of female WT and Aim2−/− mice over 48 h period (j). (k) Overnight fasting blood glucose levels at ages 18 and 26 weeks in WT and Aim2−/− mice. (l, m) GTT (l) and AUC (m) in 8-month-old female WT and Aim2−/− mice. (n, o) Fasting insulin (n) and serum TG (o) levels in female WT and Aim2−/− mice. Data are presented as mean ± SEM; n=8 for WT and n=7 for Aim2−/− mice; *p<0.05, **p<0.01 by two-way ANOVA for longitudinal analysis of body weight (b) and food intake (c), and two-tailed t test for analyses between two groups (where only one parameter was assessed; (d–g) and (i–o)). BW, body weight
EE, RER and total locomotor activity were measured at 36 weeks of age. EE was significantly decreased in Aim2−/− compared with controls when normalised to body weight (Fig. 1g and ESM Fig. 2a). When adjusted for LBM, the difference in EE persisted in males (data not shown). On regression analysis, the positive relationship between daily total EE and LBM found in WT mice was lost in Aim2−/− mice (Fig. 1h and ESM Fig. 2b). We found no significant changes in RER in both female and male Aim2−/− mice (Fig. 1i and ESM Fig. 2c). The total locomotor activity was significantly lower in female Aim2−/− mice compared with WT (Fig. 1j), but no significant difference was found in males (ESM Fig. 2d).
Aim2 −/− mice show impaired glucose homeostasis
Fasting blood glucose levels were examined at 18 and 26 weeks of age after overnight fasting. Significantly higher fasting blood glucose level was noted in female Aim2−/− mice at 18 weeks of age, and was sustained at 26 weeks (Fig. 1k). In Aim2−/− males, significantly increased fasting blood glucose levels were noted at 26 weeks of age (ESM Fig. 3a). IPGTT showed glucose intolerance (Fig. 1l and ESM Fig. 3b) in both female and male Aim2−/− mice with increased AUC for glucose (Fig. 1m and ESM Fig. 3c). Fasting insulin levels (Fig. 1n and ESM Fig. 3d) were higher in male and female Aim2−/− mice. Serum TG level was not significantly changed in female Aim2−/− mice at the time of killing (Fig. 1o), while male Aim2−/− mice demonstrated higher serum TG (ESM Fig. 3e).
Aim2 deletion induces adipogenesis in WAT
At age 14 months, individual fat pads were significantly increased in male and female Aim2−/− mice (Fig. 2a). Gonadal, inguinal and perirenal fat pads were all enlarged (Fig. 2b–d), and this was, at least in part, attributable to increased adipocyte size (hypertrophy) in Aim2−/− mice (Fig. 2e, f). Immunoblotting in gWAT demonstrated significantly increased levels of PPARγ, C/EBPα and C/EBPβ with no changes in total and phosphorylated hormone-sensitive lipase (p-HSL; Fig. 2g, h). SVFs isolated from gWAT in 3–4-month-old mice demonstrated higher differentiation potential in Aim2−/− mice (Fig. 2i). Real-time PCR showed increased gene expression levels of Pparg, Fasn, Glut4 (also known as Slc2a4) and Plin2 in differentiated SVF from Aim2−/− mice (Fig. 2j–m).
Aim2 deletion induces adipogenesis. (a) Representative images for gonadal fat in 14-month-old male and female WT and Aim2−/− mice. (b–d) Weight of gonadal (b), inguinal (c) and perirenal (d) fat pads in female WT and Aim2−/− mice. (e, f) Representative images for H&E staining in gWAT (e) and quantification of the frequency of adipocyte size from H&E staining (f). Scale bars, 200 μm. n=8 for WT and n=6 for Aim2−/− mice. (g, h) Immunoblots (g) and quantification (h) of markers for lipolysis and adipogenesis in gonadal fat in 14-month-old female WT and Aim2−/− mice. n=8 for WT and n=6 for Aim2−/− mice. (i–m) BODIPY staining for mature adipocytes (i), and mRNA levels for adipocyte markers Glut4 (j), Pparg (k), Plin2 (l) and Fasn (m) in differentiated SVFs isolated from 3–4-month-old WT and Aim2−/− mice. The key in (j) also applies to (k), (l) and (m). Scale bars, 400 μm. Data are presented as mean ± SEM; *p<0.05, **p<0.01, by two-tailed t test. GAPDH, glyceraldehyde 3-phosphate dehydrogenase
Aim2 deletion induces BAT dysfunction
BATs isolated from Aim2−/− mice were larger compared with WT mice (Fig. 3a, b). H&E staining showed larger lipid droplets in Aim2−/− BAT (Fig. 3c). Fatty acid oxidation (FAO), estimated using 14C-labelled palmitate, was significantly decreased in BAT from Aim2−/− mice (Fig. 3d). Core body temperature was lower in Aim2−/− mice both basally and after 4 h cold exposure (Fig. 3e, f). Consistent with the defect in thermogenesis, both mRNA (Ucp1) and protein levels of uncoupling protein 1 (UCP-1) and mRNA level of Cox7a1 were significantly lower in Aim2−/− mice, whereas phosphorylation of HSL and mRNA levels of Dio2 and Prdm16 were not significantly changed (Fig. 3g–l).
Aim2 deletion induces BAT dysfunction. (a–c) Representative images (a), weight (b) and H&E staining (c) of interscapular BAT isolated from 14-month-old female WT and Aim2−/− mice. Scale bars, 200 μm. (d) Rate of FAO in homogenates of BAT isolated from 5-month-old female WT and Aim2−/− mice. (e, f) Core body temperature in 14-month-old female WT and Aim2−/− mice basally (e) and after 4 h cold challenge conditions (f). (g–k) mRNA levels of Ucp1, Cox7a1, Dio2, Pparg and Prdm16 in BAT; and (l) protein levels of UCP-1 and p-HSL in BAT from 14-month-old female WT and Aim2−/− mice. Data are presented as mean ± SEM; n=8 for WT and n=6 for Aim2−/− mice; *p<0.05, **p<0.01, by two-tailed t test
Aim2 deletion upregulates proinflammatory pathway
Using RNAseq, we identified 143 genes that changed more than twofold (adjusted p < 0.01) in gWAT of 14-month-old female Aim2−/− mice compared with controls (data not shown). IFN-inducible genes such as Ifi202b, chemokines such as Cxcl10 and Ccl8, and macrophage markers such as Cd68 and F4/80 (also known as Adgre1) were significantly upregulated in Aim2−/− mice (Fig. 4a). Quantitative PCR confirmed the upregulation of Ifi202b, CD5 molecule like (Cd5l), Cxcl10 and Ccl8 in Aim2−/− mouse gWAT (Fig. 4b–e). Ingenuity pathway analysis (IPA) revealed that most of the differentially expressed genes are involved in inflammatory and immune response signalling pathways such as IFN signalling, dendritic cell maturation and T cell development (Fig. 4f).
RNAseq in WAT from 14-month-old WT and Aim−/− mice. RNAseq was performed in gWAT in 14-month-old female WT and Aim2−/− mice. (a) Heat map for inflammatory markers. (b–e) Data validation using real-time PCR in gonadal fat in 14-month-old female WT and Aim2−/− mice. (f) IPA analysis. Data are presented as mean ± SEM; n=5; *p<0.05, **p<0.01, by two-tailed t test. Th, t helper
Aim2 −/− mice show inflammation in WAT
Macrophage markers were examined by PCR in 14-month-old female Aim2−/− mice. Expression levels of CD68 and F4/80 were threefold higher in Aim2−/− mice (Fig. 5a, b). In addition, there was a 2.5-fold higher expression level of Inos (also known as Nos2; M1 macrophage marker), but similar level of Arg1 (M2 macrophage marker), in Aim2−/− mice (Fig. 5c, d). The mRNA levels of Tnfa (also known as Tnf) and Il6, but not Il18, were significantly increased in Aim2−/− mice (Fig. 5e–g). Circulating leptin was increased in Aim2−/− mice (Fig. 5h), but inflammatory cytokines TNF-α, IFN-γ, IL-1α and MCP-1 were not significantly changed (Fig. 5i–l), and IL-6 was not detectable (data not shown) in 14-month-old female Aim2−/− mice. Circulating IL-1β was also not altered by Aim2 deletion (Fig. 5m), indicating an unchanged total inflammasomal activity in Aim2−/− mice. Indeed, we found that basal caspase-1 activity in Aim2−/− gWAT was comparable to WT controls (Fig. 5n), suggesting that metabolic changes in Aim2−/− mice are independent of changes in AIM2 inflammasome activity.
AIM2 deficiency increases inflammation in WAT without affecting systemic inflammation. (a–g) mRNA levels of general macrophage markers Cd68 (a) and F4/80 (b), markers for M1 (Inos) (c) and M2 macrophages (Arg1) (d), and proinflammatory cytokines Tnfa (e), Il6 (f) and Il18 (g) in gonadal fat in 14-month-old female WT and Aim2−/− mice; n=5. (h–l) Serum levels of leptin (h), TNF-α (i), IFN-γ (j), MCP-1 (k), IL-1α (l) and IL-1β (m) in 14-month-old female WT and Aim2−/− mice; n=8 for WT and n=7 for Aim2−/− mice. (n) Caspase-1 activity in gonadal fat in WT and Aim2−/− mice; n=4. Data are presented as mean ± SEM; *p<0.05, **p<0.01, by two-tailed t test
Ifi202b is upregulated in obese mice and humans
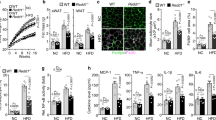
The mRNA and protein levels of Ifi202b, a top hit in the RNAseq study, were examined by both real-time PCR and immunoblot. Both mRNA and protein levels of Ifi202b were low in gWAT of 14-month-old WT, and significantly increased in Aim2−/− mice (Fig. 6a, e). The mRNA levels of Ifi202b were also significantly increased in BAT, bone marrow and liver (Fig. 6b–d). To examine whether high expression of Ifi202b is associated with obesity, we assessed the expression of Ifi202b in gWAT of NZO and HFD mice. We demonstrated that Ifi202b mRNA is markedly increased in both NZO and HFD mice (Fig. 6f, g). The mRNA level of IFI16, the human homologue of Ifi202b, was assessed by RNAseq in the subcutaneous adipose tissue of 33 school-aged children. We found that IFI16 expression positively correlated with BMI percentile (R2 = 0.085, p = 0.037 after adjusting for age and sex, Fig. 6h). Overweight and obese children had significantly higher IFI16 expression than normal weight children (p = 0.003, Fig. 6i).
Ifi202b/IFI16 expression is associated with obesity in rodents and humans. (a–d) mRNA levels of Ifi202b in WAT (a), BAT (b), bone marrow (BM; c) and liver (d) in 2-month-old female WT and Aim2−/− mice; n=4. (e) Protein level of p202 in gWAT in 14-month-old WT and Aim2−/− mice. (f, g) mRNA level of Ifi202b in the gonadal fat of NZO (f) and HFD (g) mice. C57Bl6/J mice (B6) and C57Bl6/J mice fed a regular-chow diet (RC) were used as controls, respectively; n=3 for NZO/B6 and n=4 for HFD/RC mice. (h) Association of IFI16 expression in human subcutaneous adipose tissue with BMI percentile after adjustment for age and sex. The regression line is shown in red and the grey shaded area shows the 95% CI. (i) Expression of IFI16 in human subcutaneous adipose tissue in lean and overweight (Ow) and obese (Ob) children. The thick central line shows the median, the box encases the 25th to 75th percentiles, and the whiskers represent the highest and lowest non-outlier values; the one outlier in the Ow/Ob group is indicated by the small circle. Data are presented as mean ± SEM; **p<0.01, by two-tailed t test. GAPDH, glyceraldehyde 3-phosphate dehydrogenase
Aim2 −/− mice demonstrate increased monocyte infiltration in WAT prior to onset of obesity
To examine whether the increased inflammation in Aim2−/− mice occurs before onset of obesity, we characterised the immune cell infiltration in gWAT using flow cytometry in 2-month-old male WT and Aim2−/− mice. Immune cells were labelled in bone marrow and gWAT (n = 6) using cell surface markers and gating strategies shown in Fig. 7a and ESM Fig. 4a. There was a significant increase in infiltration of total and Ly6Clow monocytes in gWAT of Aim2−/− mice (Fig. 7b–d), while the macrophages, neutrophils, dendritic cells, B cells and Ly6Chigh monocytes were unchanged (Fig. 7e–j). Consistently, we found a significant increase in mRNA levels of monocyte-recruiting chemokines including Cx3cl1, Ccl8 and Ccl3 in gWAT of Aim2−/− mice, but the Cxcl10 level was not significantly different (p = 0.056) (Fig. 7k). In bone marrow, there was a decrease in total monocytes and Ly6Chigh monocytes, and an increase in dendritic cells and macrophages, but no differences in myeloid cells, T cells, B cells, neutrophils and Ly6Clow monocytes in Aim2−/− mice (ESM Fig. 4b–k).
Increased monocyte infiltration in gonadal WAT in Aim2−/− mice before the onset of obesity and insulin resistance. (a) Gating strategy for flow cytometry study using SVFs from 8-week-old WT and Aim2−/− mice. Samples were gated for macrophages (CD64+ and F4/80+), classical DCs (MHC class II+ and CD11c+), neutrophils (Ly6G+ and Ly6Chigh), total monocytes (CD11b+, CD45+ and Ly6G−), and Ly6Chigh (CD11b+, CD115+ and Ly6Chigh) and Ly6Clow (CD11b+, CD115+ and Ly6Clow) monocytes. (b) Representative images for gated macrophages, classical DCs and neutrophils. (c–j) Quantification of the percentages of different cell populations. The key in (c) also applies to (d–j). (k) Expression levels of chemokines in gonadal fat in 8-week-old WT and Aim2−/− mice. Data are presented as mean ± SEM; n=6; *p<0.05, **p<0.01, by two-tailed t test. CDC, classical DC; DC, dendritic cell; MΦ, macrophage; Mono, monocyte; Neut, neutrophils
Ifi202b knockdown blocks adipocyte differentiation in Aim2 −/− SVFs and inflammation in Aim2 −/− BMMs
To evaluate whether p202 is the key mediator for the increased adipogenesis and/or inflammation, Ifi202b was knocked down using siRNA in SVFs and BMMs isolated from 3–4-month-old male or female WT and Aim2−/− mice. The knockdown was confirmed by PCR (>80% knockdown, Fig. 8a, f). SVFs were induced for adipocyte differentiation for 7 days as detailed above. Knockdown of Ifi202b (sip202) caused significant decrease in adipocyte differentiation in SVFs isolated from Aim2−/− mice (Fig. 8b), and reduced TG levels in the media (Fig. 8c). mRNA levels of adipocyte markers Acc1 (also known as Acaca), Acc2 (Acacb), Cebpa and Fasn (Fig. 8d) and protein levels of fatty acid synthase (FAS), acetyl-CoA carboxylase (ACC), PPARγ and C/EBPα were decreased in the Ifi202b knockdown SVFs from Aim2−/− mice (Fig. 8e). In BMMs isolated from WT and Aim2−/− mice, expression levels of IL1b, Il6 and Tnfa were upregulated in Aim2−/− BMMs when challenged with LPS, while knockdown of Ifi202b significantly reduced the levels of proinflammatory cytokines (Fig. 8f).
Knockdown of Ifi202b inhibits adipogenesis and inflammation in Aim2−/− mice. (a) Scrambled siRNA (Scr) and sip202 were transfected into the SVF using electroporation. The mRNA level of Ifi202b was examined by real-time PCR 3 days after transfection. (b) BODIPY staining for the mature adipocytes in Scr- or sip202-transfected SVFs isolated from 3–4-month-old female Aim2−/− mice after differentiation for 7 days. Scale bars, 400 μm. (c–e) TG levels in the growth medium (GM) and differentiation medium (DM) (c), and mRNA levels of the adipocyte markers (d) and protein levels of adipocyte markers (e) in the undifferentiated (GM) and differentiated (DM) SVFs isolated from Aim2−/− transfected with Scr or sip202. (f–i) Expression levels of Ifi202b and proinflammatory markers in BMMs isolated from 3–4-month-old female WT or Aim2−/− mice transfected with Scr or sip202. The key in (f) also applies to (g), (h) and (i). Experiments were repeated at least three times. Data are presented as mean ± SEM; *p<0.05, **p<0.01, by two-tailed t test. ACC, acetyl-CoA carboxylase; FAS, fatty acid synthase
Discussion
Our data highlight a role for AIM2 protein in energy metabolism and glucose homeostasis. We show that deletion of Aim2 results in obesity, increased inflammation in WAT and impaired glucose homeostasis. Our data also demonstrate that the obesity and insulin resistance are potentially secondary to upregulation of Ifi202b and its effects on adipogenesis and inflammation (Fig. 9). These findings are intriguing considering previous observations showing that deficiency of key inflammasome components, such as NLRP1, NLRP3, ASC, caspase-1 and IL-1β, protects mice from HFD-induced obesity and insulin resistance [22,23,24].
One of the striking phenotypes of Aim2 deletion is increased body weight. The difference in body weight between Aim2−/− and WT mice is not attributable to excess energy intake as their food intakes were similar. In fact, Aim2−/− mice ate less food per day if body weight is considered. Data on EE suggest that at least part of the effect on obesity is mediated through a decrease in EE noted in both sexes. A decrease in EE could be secondary to dysfunction of BAT in Aim2−/− mice as evidenced by ‘paleing’ of BAT on morphology and histology, as well as decreased FAO and downregulation of UCP-1. In BAT of Aim2−/−, there is lipid accumulation instead of FAO. This is supported by the increased expression of Pparg but unchanged Prdm16, which encodes the key transcriptional factor for BAT differentiation. Furthermore, BAT dysfunction leads to defects in thermogenesis in Aim2−/− mice and thereby contributes to obesity and insulin resistance. Indeed, defects in BAT development or activity have been shown to cause obesity and the related metabolic syndrome [40], while boosting the development and function of BAT has been found to provide beneficial effects on the metabolic syndrome [41].
Adipose tissue plays a key role in energy balance, systemic metabolism and inflammation [42]. Aim2−/− mice have increased fat mass compared with WT controls, mediated through both hypertrophy and hyperplasia. Increased adipogenesis could be associated with increased insulin sensitivity, as seen with insulin sensitisers such as PPARγ agonists. However, adipogenesis along with inflammation, as seen in Aim2−/− mice, leads to the phenotype of insulin resistance.
Interestingly, our data demonstrate that the immune cell infiltration of WAT precedes the onset of obesity in Aim2−/− mice. The monocyte infiltration in WAT of Aim2−/− mice appears to be secondary to increased expression of monocyte chemokines including Ccl8, Ccl3 and Cx3cl1. While CCL8 and CCL3 are general chemokines for monocytes, CX3CL1 is a chemokine specifically for recruiting Ly6Clow monocytes. Flow cytometer data confirmed the increased Ly6Clow monocyte infiltration in gWAT of Aim2−/− mice. Through adoptive transfer study, Geissmann et al showed that the Ly6Clow monocytes are recruited to noninflamed tissue and can differentiate into dendritic cells [43]. The Ly6Clow monocytes have been shown to secrete anti-inflammatory cytokines and are likely to differentiate into M2 macrophages [44]; however, it is highly possible that monocytes and macrophages can transdifferentiate into different subsets in response to environment changes. This has been shown in models of arthritis where Ly6Clow monocytes differentiate into proinflammatory macrophages [45]. Our data showed an increased expression level of M1 macrophage marker Inos and proinflammatory cytokines TNF-α and IL-6 at 1 year of age, suggesting that the infiltrated Ly6Clow monocytes have differentiated into proinflammatory M1 macrophages, thereby inducing inflammation and insulin resistance. A lineage-tracing study is needed to confirm this speculation. Our observations are consistent with studies that have shown associations of expression level of CX3CL1 in both mature adipocytes and SVF in human subcutaneous adipose tissue with obesity, insulin resistance and type 2 diabetes [19].
There is an upregulation of Ifi202b in many tissues, including WAT of Aim2−/− mice. This is consistent with previous studies that showed increased Ifi202b in splenocytes isolated from Aim2−/− mice [46]. While Panchanathan et al showed that deletion of Aim2 induces Ifi202b through the increase in type 1 IFNs and downstream activation of STAT1 in splenocytes [46], neither the circulating type 1 IFNs nor the expression level of local type 1 IFNs in gWAT were detectable in our study (data not shown), suggesting that alternative signalling pathway(s) regulate Ifi202b expression in the metabolic tissues in Aim2−/− mice. Previous studies have established the connection between Ifi202b and obesity [47,48,49,50]. It has been reported that p202 regulates adipogenesis in mouse adipose tissue-derived stem cells [51]. In addition, Ifi202b overexpression induces obesity with increased hypertrophy and decreased expression of thermogenic genes in mice [50]. These reports are consistent with our findings in Aim2−/− mice, indicating a role of p202 in mediating Aim2 deletion-induced fat mass expansion. Interestingly, the expression level of IFI16 is higher in subcutaneous fat in overweight and obese children compared with control participants, and the expression level of IFI16 increases with BMI percentile in a linear regression analysis adjusted for age and sex. While increased expression of IFI16 in obese adult individuals has been reported in the visceral adipose tissue [47], our data demonstrate increased expression of IFI16 in the subcutaneous tissue of both overweight and obese school-aged children. Knockdown of Ifi202b in BMMs isolated from 2-month-old Aim2−/− mice reduced the expression of proinflammatory cytokines such as Il1β, Il6 and Tnfa, indicating a key role for p202 in inflammation.
In summary, our findings demonstrate an inflammasome-independent role of AIM2 in obesity and insulin resistance that is mediated through an upregulation of Ifi202b. These observations highlight a novel role for AIM2, a protein that has so far been recognised for its role in innate immunity and as a tumour suppressor, in whole body energy homeostasis. AIM2 and p202 may serve as potential drug targets for treatment of human obesity and the related metabolic syndrome.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- AIM2:
-
Absent in melanoma 2
- ASC:
-
Apoptosis-associated speck-like protein containing a caspase activation and recruitment domain
- BAT:
-
Brown adipose tissue
- BMM:
-
Bone marrow-derived macrophage
- BODIPY :
-
Boron dipyrromethene
- C/EBP :
-
CCAAT/enhancer binding protein
- CLAMS:
-
Comprehensive Lab Animal Monitoring System
- EE:
-
Energy expenditure
- FAO:
-
Fatty acid oxidation
- gWAT:
-
Gonadal white adipose tissue
- GM-CSF:
-
Granulocyte-macrophage colony-stimulating factor
- HFD:
-
High-fat diet
- HSL:
-
Hormone-sensitive lipase
- IBMX:
-
3-Isobutyl-1-methylxanthine
- IPA:
-
Ingenuity pathway analysis
- LBM:
-
Lean body mass
- LPS:
-
Lipopolysaccharide
- MCP-1:
-
Monocyte chemoattractant protein 1
- NLR:
-
NOD-like receptor
- NZO:
-
New Zealand Obese
- p202:
-
Protein 202
- PPARγ:
-
Peroxisome proliferator activated receptor γ
- RER:
-
Respiratory exchange ratio
- RNAseq:
-
RNA sequencing
- siRNA:
-
Small interfering RNA
- SVF:
-
Stromal vascular fraction
- TBS-T:
-
Tris-buffered saline with Tween 20
- TG:
-
Triacylglycerol
- UCP-1:
-
Uncoupling protein 1
- WAT:
-
White adipose tissue
- WT:
-
Wild-type
References
Rutkowski JM, Stern JH, Scherer PE (2015) The cell biology of fat expansion. J Cell Biol 208(5):501–512. https://doi.org/10.1083/jcb.201409063
Spiegelman BM, Hu E, Kim JB, Brun R (1997) PPAR gamma and the control of adipogenesis. Biochimie 79(2-3):111–112. https://doi.org/10.1016/S0300-9084(97)81500-3
Rosen ED, Hsu CH, Wang X et al (2002) C/EBPalpha induces adipogenesis through PPARgamma: a unified pathway. Genes Dev 16(1):22–26. https://doi.org/10.1101/gad.948702
McArdle MA, Finucane OM, Connaughton RM, McMorrow AM, Roche HM (2013) Mechanisms of obesity-induced inflammation and insulin resistance: insights into the emerging role of nutritional strategies. Front Endocrinol 4:52. https://doi.org/10.3389/fendo.2013.00052
Hotamisligil GS (2017) Inflammation, metaflammation and immunometabolic disorders. Nature 542(7640):177–185. https://doi.org/10.1038/nature21363
Xu H, Barnes GT, Yang Q, Tan G et al (2003) Chronic inflammation in fat plays a crucial role in the development of obesity-related insulin resistance. J Clin Invest 112(12):1821–1830. https://doi.org/10.1172/JCI19451
Weisberg SP, McCann D, Desai M, Rosenbaum M, Leibel RL, Ferrante AW Jr (2003) Obesity is associated with macrophage accumulation in adipose tissue. J Clin Invest 112(12):1796–1808. https://doi.org/10.1172/JCI19246
Wu H, Perrard XD, Wang Q et al (2010) CD11c expression in adipose tissue and blood and its role in diet-induced obesity. Arterioscler Thromb Vasc Biol 30(2):186–192. https://doi.org/10.1161/ATVBAHA.109.198044
Elgazar-Carmon V, Rudich A, Hadad N, Levy R (2008) Neutrophils transiently infiltrate intra-abdominal fat early in the course of high-fat feeding. J Lipid Res 49(9):1894–1903. https://doi.org/10.1194/jlr.M800132-JLR200
Talukdar S, Oh DY, Bandyopadhyay G et al (2012) Neutrophils mediate insulin resistance in mice fed a high-fat diet through secreted elastase. Nat Med 18(9):1407–1412. https://doi.org/10.1038/nm.2885
Nishimura S, Manabe I, Nagasaki M et al (2009) CD8+ effector T cells contribute to macrophage recruitment and adipose tissue inflammation in obesity. Nat Med 15(8):914–920. https://doi.org/10.1038/nm.1964
Lumeng CN, Maillard I, Saltiel AR (2009) T-ing up inflammation in fat. Nat Med 15(8):846–847. https://doi.org/10.1038/nm0809-846
Feuerer M, Herrero L, Cipolletta D et al (2009) Lean, but not obese, fat is enriched for a unique population of regulatory T cells that affect metabolic parameters. Nat Med 15(8):930–939. https://doi.org/10.1038/nm.2002
Winer DA, Winer S, Shen L et al (2011) B cells promote insulin resistance through modulation of T cells and production of pathogenic IgG antibodies. Nat Med 17(5):610–617. https://doi.org/10.1038/nm.2353
Kershaw EE, Flier JS (2004) Adipose tissue as an endocrine organ. J Clin Endocrinol Metab 89(6):2548–2556. https://doi.org/10.1210/jc.2004-0395
Guzik TJ, Skiba DS, Touyz RM, Harrison DG (2017) The role of infiltrating immune cells in dysfunctional adipose tissue. Cardiovasc Res 113(9):1009–1023. https://doi.org/10.1093/cvr/cvx108
Tourniaire F, Romier-Crouzet B, Lee JH et al (2013) Chemokine expression in inflamed adipose tissue is mainly mediated by NF-κB. PLoS One 8(6):e66515. https://doi.org/10.1371/journal.pone.0066515
Shah R, Lu Y, Hinkle CC et al (2009) Gene profiling of human adipose tissue during evoked inflammation in vivo. Diabetes 58(10):2211–2219. https://doi.org/10.2337/db09-0256
Shah R, Hinkle CC, Ferguson JF et al (2011) Fractalkine is a novel human adipochemokine associated with type 2 diabetes. Diabetes 60(5):1512–1518. https://doi.org/10.2337/db10-0956
Prochnicki T, Latz E (2017) Inflammasomes on the crossroads of innate immune recognition and metabolic control. Cell Metab 26(1):71–93. https://doi.org/10.1016/j.cmet.2017.06.018
Man SM, Karki R, Kanneganti TD (2016) AIM2 inflammasome in infection, cancer, and autoimmunity: role in DNA sensing, inflammation, and innate immunity. Eur J Immunol 46(2):269–280. https://doi.org/10.1002/eji.201545839
Murphy AJ, Kraakman MJ, Kammoun HL et al (2016) IL-18 Production from the NLRP1 inflammasome prevents obesity and metabolic syndrome. Cell Metab 23(1):155–164. https://doi.org/10.1016/j.cmet.2015.09.024
Stienstra R, van Diepen JA, Tack CJ et al (2011) Inflammasome is a central player in the induction of obesity and insulin resistance. Proc Natl Acad Sci U S A 108(37):15324–15329. https://doi.org/10.1073/pnas.1100255108
Vandanmagsar B, Youm YH, Ravussin A et al (2011) The NLRP3 inflammasome instigates obesity-induced inflammation and insulin resistance. Nat Med 17(2):179–188. https://doi.org/10.1038/nm.2279
Osborn O, Brownell SE, Sanchez-Alavez M, Salomon D, Gram H, Bartfai T (2008) Treatment with an Interleukin 1 beta antibody improves glycemic control in diet-induced obesity. Cytokine 44(1):141–148. https://doi.org/10.1016/j.cyto.2008.07.004
DeYoung KL, Ray ME, Su YA et al (1997) Cloning a novel member of the human interferon-inducible gene family associated with control of tumorigenicity in a model of human melanoma. Oncogene 15(4):453–457. https://doi.org/10.1038/sj.onc.1201206
Cresswell KS, Clarke CJP, Jackson JT, Darcy PK, Trapani JA, Johnstone RW (2005) Biochemical and growth regulatory activities of the HIN-200 family member and putative tumor suppressor protein, AIM2. Biochem Biophys Res Commun 326(2):417–424. https://doi.org/10.1016/j.bbrc.2004.11.048
Chen IF, Ou-Yang F, Hung JY et al (2006) AIM2 suppresses human breast cancer cell proliferation in vitro and mammary tumor growth in a mouse model. Mol Cancer Ther 5(1):1–7. https://doi.org/10.1158/1535-7163.MCT-05-0310
Roberts TL, Idris A, Dunn JA et al (2009) HIN-200 proteins regulate caspase activation in response to foreign cytoplasmic DNA. Science 323(5917):1057–1060. https://doi.org/10.1126/science.1169841
Hornung V, Ablasser A, Charrel-Dennis M et al (2009) AIM2 recognizes cytosolic dsDNA and forms a caspase-1-activating inflammasome with ASC. Nature 458(7237):514–518. https://doi.org/10.1038/nature07725
Rathinam VA et al (2010) The AIM2 inflammasome is essential for host defense against cytosolic bacteria and DNA viruses. Nat Immunol 11(5):395–402. https://doi.org/10.1038/ni.1864
Strittmatter GE, Sand J, Sauter M et al (2016) IFN-γ primes keratinocytes for HSV-1-induced inflammasome activation. J Invest Dermatol 136(3):610–620. https://doi.org/10.1016/j.jid.2015.12.022
Zhang H, Luo J, Alcorn JF et al (2017) AIM2 inflammasome is critical for influenza-induced lung injury and mortality. J Immunol 198(11):4383–4393. https://doi.org/10.4049/jimmunol.1600714
Man SM, Zhu Q, Zhu L et al (2015) Critical role for the DNA sensor AIM2 in stem cell proliferation and cancer. Cell 162(1):45–58. https://doi.org/10.1016/j.cell.2015.06.001
Wilson JE, Petrucelli AS, Chen L et al (2015) Inflammasome-independent role of AIM2 in suppressing colon tumorigenesis via DNA-PK and Akt. Nat Med 21(8):906–913. https://doi.org/10.1038/nm.3908
Pan J, Han L, Guo J et al (2018) AIM2 accelerates the atherosclerotic plaque progressions in ApoE−/− mice. Biochem Biophys Res Commun 498(3):487–494. https://doi.org/10.1016/j.bbrc.2018.03.005
Pan J, Lu L, Wang X et al (2018) AIM2 regulates vascular smooth muscle cell migration in atherosclerosis. Biochem Biophys Res Commun 497(1):401–409. https://doi.org/10.1016/j.bbrc.2018.02.094
Ravussin E, Lillioja S, Anderson TE, Christin L, Bogardus C (1986) Determinants of 24-hour energy expenditure in man. Methods and results using a respiratory chamber. J Clin Invest 78(6):1568–1578. https://doi.org/10.1172/JCI112749
Parlee SD, Lentz SI, Mori H, MacDougald OA (2014) Quantifying size and number of adipocytes in adipose tissue. Methods Enzymol 537:93–122. https://doi.org/10.1016/B978-0-12-411619-1.00006-9
Schulz C, Fork C, Bauer T et al (2014) SLC22A13 catalyses unidirectional efflux of aspartate and glutamate at the basolateral membrane of type A intercalated cells in the renal collecting duct. Biochem J 457(2):243–251. https://doi.org/10.1042/BJ20130654
Cypess AM, Kahn CR (2010) Brown fat as a therapy for obesity and diabetes. Curr Opin Endocrinol Diabetes Obes 17(2):143–149. https://doi.org/10.1097/MED.0b013e328337a81f
Cristancho AG, Lazar MA (2011) Forming functional fat: a growing understanding of adipocyte differentiation. Nat Rev Mol Cell Biol 12(11):722–734. https://doi.org/10.1038/nrm3198
Geissmann F, Jung S, Littman DR (2003) Blood monocytes consist of two principal subsets with distinct migratory properties. Immunity 19(1):71–82. https://doi.org/10.1016/S1074-7613(03)00174-2
Auffray C, Fogg D, Garfa M et al (2007) Monitoring of blood vessels and tissues by a population of monocytes with patrolling behavior. Science 317(5838):666–670. https://doi.org/10.1126/science.1142883
Misharin AV, Cuda CM, Saber R et al (2014) Nonclassical Ly6C− monocytes drive the development of inflammatory arthritis in mice. Cell Rep 9(2):591–604. https://doi.org/10.1016/j.celrep.2014.09.032
Panchanathan R, Duan X, Shen H et al (2010) Aim2 deficiency stimulates the expression of IFN-inducible Ifi202, a lupus susceptibility murine gene within the Nba2 autoimmune susceptibility locus. J Immunol 185(12):7385–7393. https://doi.org/10.4049/jimmunol.1002468
Vogel H, Scherneck S, Kanzleiter T et al (2012) Loss of function of Ifi202b by a microdeletion on chromosome 1 of C57BL/6J mice suppresses 11β-hydroxysteroid dehydrogenase type 1 expression and development of obesity. Hum Mol Genet 21(17):3845–3857. https://doi.org/10.1093/hmg/dds213
Vogel H, Montag D, Kanzleiter T et al (2013) An interval of the obesity QTL Nob3.38 within a QTL hotspot on chromosome 1 modulates behavioral phenotypes. PLoS One 8(1):e53025. https://doi.org/10.1371/journal.pone.0053025
Vogel H, Jähnert M, Stadion M, Matzke D, Scherneck S, Schürmann A (2017) A vast genomic deletion in the C56BL/6 genome affects different genes within the Ifi200 cluster on chromosome 1 and mediates obesity and insulin resistance. BMC Genomics 18(1):172. https://doi.org/10.1186/s12864-017-3552-6
Stadion M, Schwerbel K, Graja A et al (2018) Increased Ifi202b/IFI16 expression stimulates adipogenesis in mice and humans. Diabetologia 61(5):1167–1179. https://doi.org/10.1007/s00125-018-4571-9
Li H, Liu F, Guo H, Zhu Z, Jiao Y (2014) Role of interferon-inducible protein 202 (p202) in the regulation of adipogenesis in mouse adipose-derived stem cells. Mol Cell Endocrinol 382(2):814–824. https://doi.org/10.1016/j.mce.2013.11.006
Funding
This work was supported in part by grants from Children’s Hospital of Pittsburgh of the University of Pittsburgh Medical Center Health System (RM), CHP Foundation grants (to ZG and RM) and National Institutes of Health R00HL12076, 1R01HL143967 (to PD), 1K08HL125666 (to EF), R01HL113655 (to JW) and R01DK090242 (to ESG).
Author information
Authors and Affiliations
Contributions
ZG and RM designed the experiments, performed statistical analysis and composed the manuscript. ZG, XZ, KS, RJ, EF, ESG, JW, HHD and PD conducted experiments, acquired and analysed data, and edited the manuscript. ZS and WC analysed the RNAseq data, conducted the bioinformatics analysis and edited the manuscript. All listed authors approved the final version of the manuscript. ZG and RM are the guarantors of this work.
Corresponding authors
Ethics declarations
The authors declare that there is no duality of interest associated with this manuscript.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM
(PDF 393 kb)
Rights and permissions
About this article
Cite this article
Gong, Z., Zhang, X., Su, K. et al. Deficiency in AIM2 induces inflammation and adipogenesis in white adipose tissue leading to obesity and insulin resistance. Diabetologia 62, 2325–2339 (2019). https://doi.org/10.1007/s00125-019-04983-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-019-04983-x