Abstract
Aims/hypothesis
Overexpression of the gene encoding phosphoprotein enriched in astrocytes 15 (PEA15), also known as phosphoprotein enriched in diabetes (PED), causes insulin resistance and diabetes in transgenic mice and has been observed in type 2 diabetic individuals. The aim of this study was to investigate whether PEA15 overexpression occurs in individuals at high risk of diabetes and whether it is associated with specific type 2 diabetes subphenotypes.
Subjects and methods
We analysed PEA15 expression in euglycaemic first-degree relatives (FDR) of type 2 diabetic subjects.
Results
The expression of PEA15 in peripheral blood leucocytes (PBLs) paralleled that in fat and skeletal muscle tissues. In PBLs from the FDR, PEA15 expression was two-fold higher than in euglycaemic individuals with no family history of diabetes (control subjects), both at the protein and the mRNA level (p < 0.001). The expression of PEA15 was comparable in FDR and type 2 diabetic subjects and in each group close to one-third of the subjects expressed PEA15 levels more than 2 SD higher than the mean of control subjects. Subjects with IFG with at least one type 2 diabetes-affected FDR also overexpressed PEA15 (p < 0.05). In all the groups analysed, PEA15 expression was independent of sex and unrelated to age, BMI, waist circumference, systolic and diastolic BP, and fasting cholesterol, triacylglycerol and glucose levels. However, in euglycaemic FDR of type 2 diabetic subjects, PEA15 expression was inversely correlated with insulin sensitivity (r = −557, p = 0.01).
Conclusions/interpretation
We conclude that PEA15 overexpression represents a common defect in FDR of patients with type 2 diabetes and is correlated with reduced insulin sensitivity in these individuals.
Similar content being viewed by others
Introduction
Phosphoprotein enriched in astrocytes 15 (PEA15), also known as phosphoprotein enriched in diabetes (PED) is a small scaffold protein widely produced in different tissues and highly conserved among mammals [1–3], whose gene (PEA15) maps on human chromosome 1q21–22 [4]. Several studies in cultured cells and in rodent tissues have revealed that Pea15 regulates multiple cellular functions by binding components of major intracellular transduction pathways. PEA15 activates extracellular-regulated kinase (ERK) 1/2 [3, 5]. It binds ERKs in the nucleus, exporting them into the cytoplasm, thus preventing cell cycle entry caused by sustained phospho-ERK nuclear accumulation [5, 6]. PEA15 also interacts with protein kinase B (Akt/PKB) and p90 ribosomal S6 kinase isozyme (RSK2), two key components of the phosphoinositide 3-kinase (PI3-K) and ERK signalling pathways, whose activation is central to control of cell survival [7, 8]. In addition, PEA15 exerts a wide anti-apoptotic action. It binds Fas-associated death domain (FADD) and caspase-8, thereby protecting against cytokine-induced apoptosis [9–11]. PEA15 also binds to Omi/HtrA2 to inhibit apoptosis triggered by stress and physical agents [12]. Importantly, there is evidence indicating that PEA15 anti-apoptotic action has an important role in the development and progression of certain cancers in humans as well as in rodents [13–15].
Recent reports further revealed that PEA15 binds to and enhances phospholipase D stability, resulting in increased intracellular levels of diacylglycerol [16, 17]. This effect deregulates protein kinase C (PKC) signalling and generates resistance to insulin action on glucose transport in muscle and adipose cells and in tissues from transgenic mice overexpressing the Pea15 gene [17, 18]. Inhibited glucose-regulated insulin secretion has also been reported in these animals, contributing to their impaired glucose tolerance. The Pea15 transgenic mice develop diabetes upon weight gain, indicating an important interaction between obesity and the Pea15 gene [18].
Earlier findings in humans evidenced that the PEA15 gene is overexpressed in skeletal muscle and fat tissues as well as in cultured skin fibroblasts from individuals with type 2 diabetes, and this effect occurs independently of obesity, suggesting that it may be a primary component of insulin resistance in these subjects [4]. Whether PEA15 overexpression is also detectable in more easily accessible cells, such as peripheral blood leucocytes, and whether it represents a common abnormality in type 2 diabetes is unknown. Whether the overexpression occurs in individuals at high risk of diabetes, whether it associates with diabetes risk factors other than obesity, and whether it relates to specific type 2 diabetes phenotypes also remains to be determined. The aim of the present work was to investigate these issues in first-degree relatives of type 2 diabetic individuals.
Subjects and methods
Subjects
The subjects investigated in the present study were recruited consecutively at the outpatient facilities of the Department of Preventive Medical Sciences (DSMP), the Metabolic Unit (DMCS) at the University of Naples Federico II Medical School and the Metabolic Unit at the University of Catanzaro ‘Magna Graecia’ (offspring from the European Network on Functional Genomics of Type 2 Diabetes [EUGENE2] study). Written informed consent was obtained from all participants. The study protocol was approved by the ethics committees of the participating institutions and conducted in accordance to the principles of the Declaration of Helsinki as revised in 2000. The euglycaemic individuals (n = 150) were healthy employees of the Campania Region who were not receiving pharmacological treatment, and were undergoing a routine health survey at the DSMP. The presence of type 2 diabetes-affected first-degree relatives (FDR) was ascertained through a written questionnaire and further verified through the medical history. The average number of relatives with type 2 diabetes in the subgroup of FDR was 1.1. The type 2 diabetic patients (n = 142) were recruited at the DMCS. Of these individuals, 19% were treated with diet alone, 48% with sulfonylureas and/or metformin, 30% with insulin alone, and 3% with insulin and oral hypoglycaemic agents. The mean diabetes duration was 11 ± 9 years. The mean HbA1c level of the diabetic patients was 7.0 ± 1.0%. Of the diabetic patients, 48% showed no evidence of long-term complications; 24% had microvascular and 12% had macrovascular complications; 16% had both microvascular and macrovascular complications. Anthropometric indexes (BMI was calculated by dividing the weight in kilograms by the square of the height in metres; waist circumference was measured midway between the lowest rib margin and the iliac crest to the nearest 0.5 cm) and detailed medical history (including information on type 2 diabetes-affected relatives and smoking habit) were obtained from all of the participants through identical procedures at each participating institution. Blood pressure values were measured in the left arm of the supine patient, after 5 min of quiet rest, with a mercury sphygmomanometer. For each subject, whole-blood samples were drawn from an antecubital vein in the morning after an overnight fast on two separate occasions. Plasma glucose, total cholesterol, HDL cholesterol, triacylglycerol and serum insulin levels were determined in a centralised facility as described previously [19, 20]. Diagnoses of type 2 diabetes and IFG were established according to criteria listed previously [21]. For studies in matched groups (n = 115) further individuals were recruited at the DSMP. Inclusion criteria in this arm of the study were as follows: age 52–58 years, BMI 26–32 kg/m2, waist circumference 90–100 cm, systolic BP 120–140 mmHg, diastolic BP 70–85 mmHg, total cholesterol 4.9–5.4 mmol/l, HDL cholesterol 1.2–1.4 mmol/l, serum triacylglycerol 1.4–1.6 mmol/l. Fifty-six type 2 diabetic individuals (mean disease duration 11 ± 4 years, mean HbA1c level 7.2 ± 4.0%) and 59 healthy euglycaemic subjects (of which 34 had no family history of type 2 diabetes and 25 had at least one type 2 diabetes-affected FDR) were consecutively enrolled. Studies in euglycaemic offspring of type 2 diabetic patients were performed using the EUGENE2 cohort http://www.eugene2.com, last accessed in August 2006; [20, 22, 23] at the University of Catanzaro Magna Graecia. The clinical characterisation of these individuals has been previously reported [20, 22, 23]. The OGTT and euglycaemic–hyperinsulinaemic clamp procedures have also been previously reported [20]. Briefly, for clamp studies, insulin (Humulin; Eli Lilly, Indianapolis, IN, USA) was given as a prime continuous infusion to produce plasma insulin levels of ∼420 pmol/l. Thereafter, the insulin infusion rate was fixed at 40 mU m−2 min−1. Blood glucose was maintained at a constant level throughout the study by infusing 20% glucose at varying rates according to blood glucose measurements performed at 5-min intervals. Lean body mass was assessed by bioimpedence. Duration of the clamp was 120 min. Glucose disposal is expressed as milligrams per kilogram of lean body per mass per minute. Insulin secretion was estimated by the homeostasis model assessment of pancreatic beta cell function (HOMA-B) index [24] and the insulinogenic index as the difference between the 30- and 0-min OGTT plasma insulin values divided by the difference between the corresponding plasma glucose values, (ΔI 30/ΔG 30), as described previously [25].
Separation of white blood cells and tissue sampling
For peripheral blood leucocyte (PBL) separation, EDTA-treated whole-blood samples were first centrifuged at 300 ×g for 10 min and the plasma removed. PBLs were separated using a 6% dextran gradient in filtered PBS, pH 7.4, as previously described [26], washed in PBS three times, counted and resuspended in 1 ml of PBS for subsequent use. Neutrophils and other major leucocyte subpopulations were further separated as previously described [26, 27]. The purity of the different cell populations was confirmed by microscopic examination. Subcutaneous adipose tissue from the anterior abdominal wall and skeletal muscle (rectus abdominis) biopsies were obtained simultaneously from patients undergoing elective abdominal surgery for gall bladder disease. For this part of the study, the diabetic patients and the euglycaemic subjects (with and without type 2 diabetic FDR in their pedigree) were consecutively recruited to achieve a 1:1:1 ratio between the three groups (total n = 21). Tissues were rinsed and dissected free of erythrocytes and connective tissues, as previously described [28]. PBLs were collected at the same time as fat and muscle specimens.
Cell and tissue harvesting and western blotting
PBL and tissue samples were solubilised at 4°C in TAT buffer (50 mmol/l HEPES, pH 7.5, 150 mmol/l NaCl, 10 mmol/l EDTA, 10 mmol/l Na4P2O7, 2 mmol/l Na3VO4, 100 mmol/l NaF, 10% glycerol, 1% Triton X-100) supplemented with 1 mmol/l phenylmethanesulfonylfluoride and 10 μg/ml aprotinin, as described previously [28]. Lysates were centrifuged at 500 ×g for 20 min and supernatant fractions were frozen at −20°C until used. For Western blotting, 50 μg of lysate proteins were heated at 100°C in Laemmli buffer [29]. Proteins were separated by 15% SDS-PAGE and then transferred to 0.45-mm Immobilon-P membranes (Millipore, Bedford, MA, USA). Filters were probed with phospholipase D1 (PLD1) (BioSource International, Camarillo, CA, USA) or PEA15 antibodies [4], revealed by enhanced chemiluminescence and autoradiography, and PEA15 bands were quantitated by laser densitometry. Inter-assay variation was <15%.
RNA samples and real-time RT-PCR
RNA from white blood cells were immediately extracted following cell isolation using the QIAamp RNA Blood Mini Kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions. PEA15 gene expression was measured by real-time RT-PCR (SYBRGreen with iCycler IQ Multicolor Real-Time Detection System, Biorad, Richmond, VA, USA). The following primer pairs were designed using the Beacon Designer Software (Biorad) based on published gene sequences (GeneBank Accession nos. AH008227 and M10277, respectively). Human hped1 (forward) 5′-GAGCGCTCAGCTCCAGAGG-3′; human hped3 (reverse) 5′-CAGGACGGCGGGAGATCT-3′; human β-actin (forward) 5′-TGCGTGACATTAAGGAGAAG-3′; human β-actin (reverse) 5′-GCTCGTAGCTCTTCTCCA-3′. Real-time RT-PCR was performed with a cDNA input of 50 ng total RNA with all combinations of primers. Data acquisition and analysis were automatically performed by the iCycler IQ Multicolour Real-Time Detection System Optical System Software, version 3.1 (Biorad). Primers for real-time detection of PLD1 were as follows. Human PLD1 (forward) 5′-TGGGACCAAAGGGCATAGAAGG-3′ human PLD1 (reverse) 5′-TACCAGCAGGACGAAGGCAATG-3′.
Statistical analysis
Data are means ± SEM. Comparisons between groups were tested by the unpaired Student’s t test or the Mann–Whitney U test, and among groups by ANOVA. Mean values, after adjusting for covariates, were evaluated by ANCOVA. A p value of < 0.05 was considered significant. Analyses were performed using the statistical package SPSS, version 12 (SPSS, Chicago, IL, USA).
Results
To identify tissues for which PEA15 expression can be non-invasively quantified in a large number of subjects, we first analysed lysates from PBLs by immunoblotting with specific PEA15 antibodies [4]. In samples from 21 subjects (seven with type 2 diabetes, seven euglycaemic subjects with diabetic FDR [EuF+], and seven euglycaemic subjects with no family history of diabetes [EuF−]), these assays revealed the expression of PEA15, with no significant differences between the granulocyte, lymphocyte and monocyte cells (Fig. 1a). On a per-mg-protein basis, PEA15 levels in these cells were 30% higher than those in human skeletal muscle and adipose tissues (p < 0.001). Importantly, in these same subjects, there were significant correlations between PEA15 levels in the PBLs and those in fat and in skeletal muscle tissues (p < 0.001; Fig. 1a,b).
Expression of PEA15 protein in human white blood cells. a PBLs and the relative granulocyte, lymphocyte and monocyte fractions were prepared from samples from 21 individuals (type 2 diabetic, EuF+ and Eu− subjects at a 1:1:1 ratio). Biopsy specimens of skeletal muscle and adipose tissues from these same patients were collected at the time they underwent elective abdominal surgery, as described in the Subjects and methods section. Cells and tissues were solubilised and analysed by western blotting PEA15 antibodies [4]. Blots were revealed by enhanced chemiluminescence and autoradiography, and PEA15 levels quantified by laser densitometry. Bars represent the means ± SEM of four independent experiments. Asterisks indicate statistical significance between bracketed columns (p < 0.001). Correlation between PEA15 levels in PBLs and muscle (r = 0.881, p < 0.001) (b) or fat (r = 0.845, p < 0.001) (c) was tested by linear regression analysis, as described in Subjects and methods
We then compared the levels of PEA15 in PBL lysates from 30 EuF+ individuals (these subjects are known to be at increased risk of diabetes [30, 31]) with those in PBL lysates from 120 EuF− subjects. Their clinical characteristics are shown in Table 1. A two-fold increase in PEA15 levels was found in the EuF+ subjects (p < 0.001; Fig. 2a). This increase was of a similar magnitude to that observed in an additional group of 142 type 2 diabetic patients, whose clinical features are also shown in Table 1. As in the case of the type 2 diabetic patients, close to one-third of the EuF+ subjects expressed amounts of PEA15 protein that were more than two SDs higher than the mean for the EuF− subjects. As previously demonstrated in isolated fibroblasts, skeletal muscle and adipose tissues from type 2 diabetic individuals [4], the increased levels of PEA15 protein detected in PBLs from the EuF+ and the type 2 diabetic subjects occurred in parallel with an increase in PEA15 mRNA (Fig. 2b). Also, the increased PEA15 gene expression in the EuF+ and the type 2 diabetic individuals was paralleled by a three-fold increase in levels of the PEA15 downstream target PLD1 (p = 0.001; Fig. 2c), with no change in PLD1 mRNA (data not shown).
Expression of PEA15 in PBLs from euglycaemic subjects. a PBLs from 30 euglycaemic individuals with at least one type 2 diabetic FDR (EuF+) and 120 euglycaemic subjects lacking a family history of diabetes (EuF−) were consecutively collected. For comparison, PBLs from 142 type 2 diabetic patients (T2D) were also analysed. Cells were solubilised and PEA15 (a) and PLD protein levels (c) were quantified in the lysates as outlined in the legend to Fig. 1. Data points represent the mean of three independent determinations in each individual subject. RNA was extracted from PBLs and PEA15 mRNA (b) quantified by real-time PCR as described in the Subjects and methods section. Data points are the means of three independent determinations in each individual subject
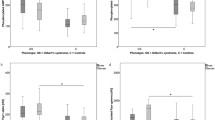
As in our previous report [4], in the present study we found no difference in PEA15 expression between the obese and lean individuals, irrespective of whether they were euglycaemic or diabetic (data not shown). To address the effect of other major risk factors for type 2 diabetes, we analysed the levels of PEA15 in 55 further individuals diagnosed with IFG according to a previous report [21]. Ten of these subjects had at least one type 2 diabetes-affected FDR (IFG-F+), while the others had no family history of disease (IFG-F−). PBL levels of PEA15 in IFG-F− subjects were found to be very comparable to those detected in EuF− subjects (Fig. 3a). As in the case of the euglycaemic individuals, however, IFG-F+ subjects exhibited PEA15 levels that were significantly greater than those detected in the IFG-F− subjects (p < 0.05), further pointing to an important effect of family history on PEA15 levels in euglycaemic and IFG humans.
Type 2 diabetes-associated phenotypes and risk-factors related to PEA15 expression. a PBL were obtained from 55 consecutive individuals with IFG diagnosed as described previously [21]. Ten of these subjects had at least one type 2 diabetes-affected FDR (IFG-F+), while the others had no family history of diabetes (n = 45; IFG-F−). Cells were solubilised and PEA15 protein levels were quantified in the lysates as outlined in the legend to Fig. 1. For comparison, data obtained from PBL from the euglycaemic subjects lacking a family history of diabetes and shown in Fig. 2 (EuF−) are also presented. b Two further groups of euglycaemic subjects with (n = 25; EuF+) or without (n = 36; EuF−) a family history of type 2 diabetes and of type 2 diabetic subjects (n = 58; T2D) were matched for age, BMI, waist circumference, systolic BP and diastolic BP and fasting cholesterol and triacylglycerol levels as described in the Subjects and methods section. PBLs from these subjects were obtained and solubilised, and PEA15 protein levels quantitated in the lysates as outlined in the legend to Fig. 1. Bars represent the mean ± SEM of three (a) or four (b) independent experiments, each in duplicate. Asterisks indicate statistical significance between bracketed columns (p < 0.001). c PEA15 protein levels were determined in PBLs from 25 euglycaemic offspring of couples, one of which was affected by type 2 diabetes. Glucose disposal was assessed by the euglycaemic–hyperinsulinaemic clamp as described in the Subjects and methods section and corrected for fat-free mass. Linear regression analysis performed as described in the Subjects and methods section revealed a correlation between PEA15 levels in PBLs and glucose disposal (r = −0.557, p = 0.01)
In the euglycaemic subjects (both EuF+ and EuF−) and in the type 2 diabetic patients described in Table 1, there were no sex-related differences in PEA15 level. Also, no significant correlation was found between the production of PED15 and the age, BMI, waist circumference, systolic BP or diastolic BP, fasting cholesterol, triacylglycerol or glucose levels of the subjects (data not shown), indicating that PEA15 levels are independent of these variables in humans. To further examine this hypothesis, we compared PEA15 levels in three additional groups of consecutively recruited euglycaemic (EuF+ and EuF−) and type 2 diabetic individuals matched for age, BMI, waist circumference, systolic BP and diastolic BP, and fasting HDL cholesterol and triacylglycerol levels (Table 2). A two-fold increase in PEA15 levels was again observed in both the diabetic subjects and the EuF+ subjects compared with the EuF− subjects (p < 0.01; Fig. 3b), providing corroborating evidence that PEA15 is independent these variables in humans.
To address the functional significance of PEA15 overexpression in type 2 diabetic FDR, we searched for associations with diabetes-related phenotypes. We analysed an additional group of 25 euglycaemic offspring of type 2 diabetes-affected couples from the EUGENE2 cohort http://www.eugene2.com, last accessed in August 2006; [20, 22, 23]. PBL expression levels of PEA15 in these offspring were comparable to those of the other type 2 diabetic FDR investigated in this study (Table 3). The other clinical characteristics of these subjects are also shown in Table 3. A negative correlation was evidenced between the individual levels of PEA15 in PBLs and insulin-stimulated glucose disposal according to fat-free mass, as determined by the euglycaemic–hyperinsulinaemic clamp (r = −557, p = 0.01; Fig. 3c). Insulin-stimulated glucose disposal also negatively correlated with BMI in these individuals (r = −0.64, p = 0.01). However, the correlation with PEA15 levels remained significant after adjustment for age, sex and BMI (r = −491, p = 0.02). As in the case of the other FDR investigated in this study, PEA15 levels did not correlate with age, BMI, waist circumference, SBP or DBP, or fasting cholesterol, triacylglycerol or glucose levels in these offspring (data not shown). Thus, the overexpression of PEA15 may induce insulin-resistance in euglycaemic offspring of type 2 diabetes-affected couples.
Discussion
Previous studies have reported that PEA15 is a multifunctional protein that controls a number of cellular functions, including proliferation, apoptosis and insulin-regulated glucose transport [4–12, 17, 18, 32]. In transgenic mice fed high-fat diets, the overexpression of Pea15 leads to diabetes [18]. In humans, no genetic variability accounting for the differential expression of the PEA15 gene has been identified to date. However, PEA15 overexpression was shown to occur in skeletal muscle and adipose tissues from type 2 diabetic patients, independently of drug treatment and obesity [4]. Whether this abnormality is associated with other risk factors for type 2 diabetes is unknown, as is whether it contributes to specific phenotype(s) associated with diabetes in humans. In the present work we have addressed these questions in FDR of type 2 diabetic subjects. These individuals have a very high risk of type 2 diabetes [30, 31] and develop different diabetes-related phenotypes years before diabetes onset, independently of the metabolic abnormalities associated with this disorder [33–35]. We have shown that the PEA15 gene is overexpressed in PBLs from EuF+ subjects. As previously shown in cells from type 2 diabetic subjects [4], this overexpression occurs at both the mRNA and protein level indicating that, as previously demonstrated in cells from type 2 diabetic subjects [4], it is caused, at least in part, by a transcriptional abnormality. Earlier studies in isolated cells and in vivo showed that PEA15 binds to and increases the cellular stability of PLD1, deregulating PKC signalling and impairing insulin-dependent glucose disposal [17, 18]. We now show that enhanced PLD1 stability also appears to occur in PBLs from type 2 diabetic subjects and FDR, suggesting that it occurs in individuals who overexpress PEA15. Indeed, in parallel with PEA15, cells from both type 2 diabetic and EuF+ individuals exhibit PLD protein levels that are three times higher than those seen in EuF− individuals, with no change in mRNA. PEA15 levels in the PBLs are closely correlated with those in fat and skeletal muscle tissues. Thus, although PBLs do not represent a classical target for insulin, they do allow the non-invasive investigation of the role of PEA15 overexpression during progression towards type 2 diabetes. In this study the overexpression of PEA15 relative to the expression seen in EuF− individuals was demonstrated in almost one-third of the EuF+ subjects and in a similar proportion of the type 2 diabetic subjects. It appears therefore that increased PEA15 levels represent a common abnormality in both type 2 diabetic subjects and individuals at increased risk of this disease, suggesting that it might precede diabetes onset in the latter.
In both type 2 diabetic subjects and their FDR, PEA15 levels were independent of BMI, waist circumference, systolic and diastolic blood pressure, HDL cholesterol, triacylglycerol and glucose levels, indicating that expression of the PEA15 gene is independent of the main variables associated with the metabolic syndrome in humans. PEA15 levels were also not related to sex or age, or to reduced physical activity or smoke habit, two recognised risk factors for type 2 diabetes and insulin resistance. Interestingly, we report that, similar to EuF+ individuals, individuals with IFG who are FDR of type 2 diabetic individuals (IFG-F+) also exhibit high levels of PEA15. In contrast, IFG subjects lacking a family history of type 2 diabetes (IFG-F−) do not, further indicating familiar clustering of this trait. The overexpression of the PEA15 gene in the type 2 diabetic subjects and their FDR may be genetically or environmentally determined, or determined by both factors. Indeed, previous studies [4] have shown that the overexpression persists in human skin fibroblasts after several generations in culture, suggesting that the occurrence of similarly high PEA15 levels in the EuF+, IFG+ and type 2 diabetic individuals is at least partly a genetically determined effect. At variance with these euglycaemic and IFG individuals, the presence of a FDR affected by type 2 diabetes had no effect on PEA15 levels in patients with an established diagnosis of type 2 diabetes (data not shown). Once deranged glucose tolerance enables overt diabetes to occur, factors secondary to the disease might modify PEA15 gene expression and/or PEA15 protein levels, thereby masking the primary overexpression. Consistent with this possibility, recent studies by our own as well as other laboratories have shown that PEA15 is highly regulated at the post-translational level [1, 2, 7, 12].
Previous studies in cultured cells and in transgenic mice have shown that overexpression of the gene encoding PEA15 determines resistance to insulin action and impairs glucose-induced insulin secretion. In this study, we investigated whether PEA15 overexpression is associated with impaired insulin action and/or secretion in euglycaemic offspring of type 2 diabetes-affected couples. In these individuals, who are at increased risk of diabetes, insulin resistance and reduced non-oxidative glucose metabolism appear years before the onset of hyperglycaemia [33, 34]. Using results from euglycaemic–hyperinsulinaemic clamp studies, a negative correlation was established between PEA15 levels and the insulin-stimulated glucose disposal by the fat-free mass of the offspring, suggesting that high levels of PEA15 protein contribute to development of skeletal muscle resistance to insulin action in these individuals. PEA15 levels were also weakly correlated with fasting plasma insulin in these FDR, but not in euglycaemic individuals lacking a family history of type 2 diabetes (data not shown). PEA15 might induce insulin resistance only in conjunction with other diabetes-related genes enriched in at-risk individuals. In this case, as previously demonstrated by Mootha et al. [36], gene enrichment analysis [37] may help to further unravel the mechanistic significance of increased levels of PEA15 in human diabetes. However, whether the PEA15 effect is weaker in individuals lacking a particular, as-yet-undefined, genetic background cannot be concluded from these studies. Indeed, unlike the hyperglycaemic–euglycaemic clamp, fasting plasma insulin is a very crude index of insulin sensitivity [38]. In addition to insulin resistance, offspring of type 2 diabetes-affected couples may also have beta cell dysfunction, as evidenced by decreases in insulin and amylin secretion in response to glucose stimulation [35]. However, PEA15 levels were not correlated with any index of beta cell function in the offspring. Overexpression of the gene may impair glucose-triggered insulin secretion in transgenic mice but not in humans. This is unlikely, however, as transfection of a PEA15 cDNA in human beta cell lines grossly impairs the insulin response to glucose (data not shown). Alternatively, in the type 2 diabetic FDR, the consequence of the high PEA15 levels on insulin action might occur earlier during life than the PEA15 effect on beta cell function.
In conclusion, in the present study we have shown that PEA15 overexpression represents a common abnormality in both type 2 diabetic patients and their FDR and can be conveniently detected in PBLs. In at-risk individuals, this defect is associated with the presence of a type 2 diabetes-affected FDR and not with other major type 2 diabetes risk factors. In euglycaemic FDR of type 2 diabetic patients, high PEA15 levels are strongly correlated with resistance to insulin action in the lean mass, suggesting that PEA15 contributes to the early appearance of insulin resistance in these individuals. Follow-up studies are in progress to further elucidate the effect of PEA15 levels in type 2 diabetes.
Abbreviations
- ERK:
-
extracellular-regulated kinase
- EUGENE:
-
European Network on Functional Genomics of Type 2 Diabetes
- FDR:
-
first-degree relatives
- PBL:
-
peripheral blood leucocyte
- PEA15:
-
phosphoprotein enriched in diabetes/phosphoprotein enriched in astrocytes
- PKC:
-
protein kinase C
- PLD1:
-
phospholipase D1
References
Danziger N, Yokoyama M, Jay T, Cordier J, Glowinski J, Chneiweiss H (1995) Cellular expression, developmental regulation, and phylogenic conservation of PEA-15, the astrocytic major phosphoprotein and protein kinase C substrate. J Neurochem 64:1016–1025
Estelles A, Yokoyama M, Nothias F et al (1996) The major astrocytic phosphoprotein PEA-15 is encoded by two mRNAs conserved on their full length in mouse and human. J Biol Chem 271:14800–14806
Ramos JW, Hughes PE, Renshaw MW et al (2000) Death effector domain protein PEA-15 potentiates Ras activation of extracellular signal receptor-activated kinase by an adhesion-independent mechanism. Mol Biol Cell 11:2863–2872
Condorelli G, Vigliotta G, Iavarone C et al (1998) PED/PEA-15 gene controls glucose transport and is overexpressed in type 2 diabetes mellitus. EMBO J 17:3858–3866
Formstecher E, Ramos JW, Fauquet M et al (2001) PEA-15 mediates cytoplasmic sequestration of ERK MAP kinase. Dev Cell 1:239–250
Brunet A, Roux D, Lenormand P, Dowd S, Keyse S, Pouyssegur J (1999) Nuclear translocation of p42/p44 mitogen-activated protein kinase is required for growth factor-induced gene expression and cell cycle entry. EMBO J 18:664–674
Trencia A, Perfetti A, Cassese A et al (2003) Protein kinase B/Akt binds and phosphorylates PED/PEA-15, stabilizing its antiapoptotic action. Mol Cell Biol 23:4511–4521
Vaidyanathan H, Ramos JW (2003) RSK2 activity is regulated by its interaction with PEA-15. J Biol Chem 278:32367–32372
Estelles A, Charlton CA, Blau HM (1999) The phosphoprotein protein PEA-15 inhibits Fas- but increases TNF-R1-mediated caspase-8 activity and apoptosis. Dev Biol 1999 216:16–28
Kitsberg D, Formstecher E, Fauquet M et al (1999) Knock-out of the neural death effector domain protein PEA-15 demonstrates that its expression protects astrocytes from TNFalpha-induced apoptosis. J Neurosci 19:8244–8251
Condorelli G, Vigliotta G, Cafieri A et al (1999) PED/PEA-15: an anti-apoptotic molecule that regulates FAS/TNFR1-induced apoptosis. Oncogene 18:4409–4415
Trencia A, Fiory F, Maitan MA et al (2004) Omi/HtrA2 promotes cell death by binding and degrading the anti-apoptotic protein ped/pea-15. J Biol Chem 279:46566–46572
Xiao C, Yang BF, Asadi N, Beguinot F, Hao C (2002) Tumor necrosis factor-related apoptosis-inducing ligand-induced death-inducing signaling complex and its modulation by c-FLIP and PED/PEA-15 in glioma cells. J Biol Chem 277:25020–25025
Hao C, Beguinot F, Condorelli G et al (2001) Induction and intracellular regulation of tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) mediated apoptosis in human malignant glioma cells. Cancer Res 61:1162–1170
Formisano P, Perruolo G, Libertini S et al (2005) Raised expression of the antiapoptotic protein ped/pea-15 increases susceptibility to chemically induced skin tumor development. Oncogene 24:7012–7021
Zhang Y, Redina O, Altshuller YM et al (2000) Regulation of expression of phospholipase D1 and D2 by PEA-15, a novel protein that interacts with them. J Biol Chem 275:35224–35232
Condorelli G, Vigliotta G, Trencia A et al (2001) Protein kinase C (PKC)-alpha activation inhibits PKC-zeta and mediates the action of PED/PEA-15 on glucose transport in the L6 skeletal muscle cells. Diabetes 50:1244–1252
Vigliotta G, Miele C, Santopietro S et al (2004) Overexpression of the PED/PEA-15 gene causes diabetes by impairing glucose-stimulated insulin secretion in addition to insulin action. Mol Cell Biol 24:5005–5015
Di Paola R, Frittitta L, Miscio G et al (2002) A variation in 3′ UTR of hPTP1B increases specific gene expression and associates with insulin resistance. Am J Hum Genet 70:806–812
Marini MA, Frontoni S, Mineo D et al (2003) The Arg972 variant in insulin receptor substrate-1 is associated with an atherogenic profile in offspring of type 2 diabetic patients. J Clin Endocrinol Metab 88:3368–3371
Genuth S, Alberti KG, Bennett P et al (2003) Follow-up report on the diagnosis of diabetes mellitus. Diabetes Care 26:3160–3167
Prudente S, Hribal ML, Flex E et al (2005) The functional Q84R polymorphism of mammalian Tribbles homolog TRB3 is associated with insulin resistance and related cardiovascular risk in Caucasians from Italy. Diabetes 54:2807–2811
Cardellini M, Perego L, D’Adamo M et al (2005) C-174G polymorphism in the promoter of the interleukin-6 gene is associated with insulin resistance. Diabetes Care 28:2007–2012
Matthews DR, Hosker JP, Rudenski AS, Naylor BA, Treacher DF, Turner RC (1985) Homeostasis model assessment: insulin resistance nad beta-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia 28:412–419
Hanson RL, Pratley RE, Bogardus C et al (2000) Evaluation of simple indices of insulin sensitivity and insulin secretion for use in epidemiologic studies. Am J Epidemiol 151:190–198
Metcalfe P, Waters AH (1992) Location of the granulocyte-specific antigen LAN on the Fc-receptor III. Transfus Med 2:283–287
Seager Danciger J, Lutz M, Hama S et al (2004) Method for large scale isolation, culture and cryopreservation of human monocytes suitable for chemotaxis, cellular adhesion assays, macrophage and dendritic cell differentiation. J Immunol Methods 288:123–134
Miele C, Formisano P, Condorelli G et al (1997) Abnormal glucose transport and GLUT1 cell-surface content in fibroblasts and skeletal muscle from NIDDM and obese subjects. Diabetologia 40:421–429
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Klein BE, Klein R, Moss SE, Cruickshanks KJ (1996) Parental history of diabetes in a population-based study. Diabetes Care 19:827–830
Bennet PH (1990) Epidemiology of diabetes mellitus. In: Rifkin H, Porte D Jr (eds) Hellenberg and Rifkin’s diabetes mellitus. Elsevier, New York, pp 363–377
Condorelli G, Trencia A, Vigliotta G et al (2002) Multiple members of the mitogen-activated protein kinase family are necessary for PED/PEA-15 anti-apoptotic function. J Biol Chem 277:11013–11018
Beck-Nielsen H, Groop LC (1994) Metabolic and genetic characterization of prediabetic states. Sequence of events leading to non-insulin-dependent diabetes mellitus J Clin Invest 94:1714–1721
Eriksson J, Franssila-Kallunki A, Ekstrand A et al (1989) Early metabolic defects in persons at increased risk for non-insulin-dependent diabetes mellitus. N Engl J Med 321:337–343
Knowles NG, Landchild MA, Fujimoto WY, Kahn SE (2002) Insulin and amylin release are both diminished in first-degree relatives of subjects with type 2 diabetes. Diabetes Care 25:292–297
Mootha VK, Lingren CM, Eriksson KF et al (2003) PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34:267–273
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550
Castillo MJ (2002) Simple assessment of insulin sensitivity and secretion. Int Diabetes Monitor 14:1–7
Acknowledgements
This work was supported in part by the European Community’s FP6 EUGENE2 (LSHM-CT-2004-512013), grants from the EFSD to F. Beguinot, the Associazione Italiana per la Ricerca sul Cancro (AIRC) (Italian Association for Cancer Research) to F. Beguinot and P. Formisano, and the Ministero dell’Università e della Ricerca Scientifica (Ministry of Education) (PRIN to F. Beguinot and P. Formisano and FIRB RBNE0155LB to F. Beguinot). The financial support of Telethon – Italy is gratefully acknowledged. The authors also warmly thank R. Napoli (University of Naples ‘Federico II’) for continuous advice during the course of this work and for critical reading of the manuscript, and G. Fratellanza for technical help.
Duality of interest
The authors declare that there are no competing financial interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Valentino, R., Lupoli, G.A., Raciti, G.A. et al. The PEA15 gene is overexpressed and related to insulin resistance in healthy first-degree relatives of patients with type 2 diabetes. Diabetologia 49, 3058–3066 (2006). https://doi.org/10.1007/s00125-006-0455-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-006-0455-5