Abstract
Aims/hypothesis
Beta cell loss in Type 1 and Type 2 diabetes mellitus may result from apoptosis and necrosis induced by inflammatory mediators. The suppressor of cytokine signalling (SOCS)-3 is a natural inhibitor of cytokine signalling and also influences insulin signalling. SOCS3 could therefore be a candidate gene in the development of Type 1 and Type 2 diabetes mellitus.
Methods
Mutation analysis of the SOCS3 gene was performed in 21 patients with Type 1 diabetes mellitus and in seven healthy subjects. An identified promoter variant was examined in (i) 250 families with Type 1 diabetic family members (1097 individuals); (ii) 212 glucose-tolerant first-degree relatives of Type 2 diabetic patients; and (iii) 370 population-based young, healthy subjects who were unrelated.
Results
Three mutations were identified in the promoter region, but none in the coding region or the 3′UTR. Two of the three mutations had allele frequencies below 1% whereas the C −920→A substitution had a minor allele frequency of 8%. In the group of young healthy subjects the insulin sensitivity index was higher among homozygous carriers of the A-allele than among heterozygous and wild-type subjects (p=0.027, uncorrected). The same trend was found in the group of first-degree relatives of Type 2 diabetic patients. No association or linkage was found to Type 1 diabetes mellitus.
Conclusions/interpretation
Homozygosity for the A-allele of the C −920→A promoter polymorphism of the SOCS3 gene may be associated with increased whole-body insulin sensitivity, but deserves further investigation.
Similar content being viewed by others
Introduction
In terms of aetiology, genetics and pathogenesis, Type 1 and Type 2 diabetes mellitus are considered to be two distinct diseases. There is, however, accumulating evidence that inflammatory mediators are important in the pathogenesis of both diseases. Beta cell loss in Type 1 and Type 2 diabetes mellitus may be a consequence of apoptosis and necrosis induced by inflammatory mediators. In Type 1 diabetes mellitus, beta cell destruction results from an immune-mediated reaction towards the beta cell. Proinflammatory cytokines are released during inflammation of the pancreatic islets and in synergy they lead to apoptosis and necrosis of beta cells [1]. In Type 2 diabetes mellitus, beta cell deficit and increased beta cell apoptosis have also been reported [2], as well as elevated circulating levels of proinflammatory cytokines in patients with recent onset of the disease [3, 4, 5]. In addition, hyperglycaemia in Type 2 diabetic patients is accompanied by increased levels of proinflammatory cytokines [6]. Finally, numerous studies have shown that several proinflammatory cytokines induce insulin resistance [7, 8, 9]. Thus, cytokines that are produced locally as part of the autoimmune infiltrate in Type 1 diabetes mellitus or are expressed in beta cells and/or at high levels in circulation in response to hyperglycaemia in Type 2 diabetes may be a common pathogenetic denominator in beta cell failure in the two diseases.
A number of natural inhibitors of cytokine signalling have recently been characterised. The suppressor of cytokine signalling (SOCS) family is a group of proteins originally identified as negative feedback regulators of IFN- γ signalling [10]. We recently reported that SOCS3 also down-regulates IL-1 signalling [11]. Interestingly, SOCS3 up-regulation in response to IL-1 in native rat islets is delayed and may therefore be insufficient to down-regulate IL-1-mediated signalling and thereby increase beta cell sensitivity to cytokines [11]. TNF-α induces insulin resistance both indirectly by stimulating stress hormone production, and directly by sustained induction of SOCS3, which decreases insulin-induced IRS1 tyrosine phosphorylation and its association with the p85 regulatory subunit of phosphatidylinositol-3 kinase [7]. SOCS3 was shown to modulate insulin signalling by targeting IRS1 and IRS2, the two key signalling proteins in insulin action [12]. It has also been proposed that the adipocyte hormone leptin, which induces SOCS3, impedes insulin signalling. Growth hormone (GH) also induces SOCS3, thereby leading to insulin resistance [13].
Insulin resistance, which often precedes the onset of Type 2 diabetes mellitus by several years, affects a large segment of the general population. There is increasing evidence that genetic components code for insulin resistance [14, 15, 16], which is a feature of the offspring of parents with Type 2 diabetes mellitus and, as shown by longitudinal studies of families affected by the disease, a major risk factor for developing Type 2 diabetes mellitus [17].
For these reasons, we consider SOCS3 to be a candidate gene in the pathogenesis of Type 1 diabetes and insulin resistance. SOCS3 maps to chromosome 17q25 and consists of only one exon spanning 850 nucleotides. The upstream sequence is a G/C rich region and the promoter area is by definition a GpC island [18]. In the present study we performed a mutation scanning of the promoter region, the exon and the 3′UTR of the human SOCS3 gene. We also examined the identified variants for an association with Type 1 diabetes mellitus and insulin sensitivity.
Subjects, materials and methods
Subjects
Mutation scanning was performed in seven control subjects and 21 Type 1 diabetic patients by single-strand conformational polymorphism (SSCP) and direct sequencing. To estimate the frequencies of minor alleles, the single nucleotide polymorphisms (SNPs) identified were tested further in a panel of 100 Type 1 diabetic patients. An identified promoter variant was examined in three separate groups of subjects involving:
-
1.
A collection of families affected by Type 1 diabetes mellitus. This group comprised 250 Danish families (1097 individuals), consisting of 99 simplex and 151 multiplex families. The median age (range) at onset for probands was 11 years (0–29 years) [19, 20]. HLA stratification was performed for all probands and/or affected offspring, who were subsequently divided into high-risk DR3/4 heterozygous and non-high-risk (non-DR3/4 heterozygous) HLA-types.
-
2.
A group of 212 glucose-tolerant Danish first-degree relatives of Type 2 diabetes mellitus patients. This group was examined by a 75-g OGTT and an IVGTT in combination with an intravenous tolbutamide injection .The detailed phenotypic characteristics of this population sample have been reported previously [21].
-
3.
A random population-based sample of young, healthy, unrelated Danish subjects (n=370) from Copenhagen. Aged 18 to 32 years, these subjects underwent an IVGTT in combination with an intravenous tolbutamide injection [22].
Because the penetrance of a single allele on a physiological trait like insulin sensitivity levels is likely to be higher in glucose-tolerant subjects than in patients with a complex metabolic disorder like Type 2 diabetes mellitus, we chose to investigate offspring of Type 2 diabetes mellitus subjects and healthy unrelated subjects. The subjects of the study gave informed consent and the investigations were approved by the responsible ethics committee and carried out in accordance with the Declaration of Helsinki as revised in 2000.
Mutation scanning and genotyping
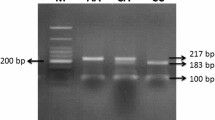
SSCP and sequencing were performed as described previously [23, 24]. Sequence data were analysed using SeqScape version 2.0 (Applied Biosystems, Foster City, Calif., USA). Genotypings of polymorphisms were done by primer extension using the SnapShot Multiplex Kit (Applied Biosystems). In the PCR reaction the following primers were used 5′CTCCGCGCTCAGCCTTTCTCTGC 3′ and 5′TACCTGGTCCCGAATCGAAGTCT 3′. As a template we used 100 ng DNA in a total volume of 25 µl in a thermal cycler (Perkin Elmer, Boston, Mass., USA). In addition the PCR mixture contained 1.75 units Herculase-enhanced DNA polymerase (Stratagene, La Jolla, Calif., USA), 1× Herculase PCR buffer (Stratagene), 1 µmol/l of each primer, 25 µmol/l deoxynucleoside (5′-)triphosphate, DMSO 8% and H2O. Denaturing, annealing and extension temperatures were 95 °C, 64 °C and 72 °C respectively, for 30 seconds each for a total of 35 cycles. In the last 25 cycles 10 seconds were added to each cycle, followed by a final extension for 10 min at 72 °C. Primer extensions were performed according to the manufacturer’s instructions, using the following primer 5′ CGGGAGCTGGGCCGGGCGGGCGGC 3′, terminating one nucleotide before the polymorphism. The assay was performed on an ABI Prism 3100 (Applied Biosystems) and analysed using Genemapper version 2.0 (Applied Biosystems). Evaluations of the SNP database (NCBI [25]) revealed three rare variations of the SOCS gene region. One of these (rs 1061489), which causes a tyrosine to histidine substitution (Y125H), was also evaluated by RFLP-PCR assay in the current study.
Phenotypic characterisation
In Groups 2 and 3, comprising glucose-tolerant first-degree relatives of Danish Type 2 diabetic patients and healthy, unrelated, Danish subjects respectively, the insulin sensitivity index (SI) was measured applying Bergman’s minimal model computing data, obtained during an intravenous glucose and tolbutamide tolerance test. The insulinogenic index for insulin was calculated as (insulin t=30 min−insulin t=0 min/glucose t=30 min). The insulinogenic index for C-peptide was calculated as (C-peptide t=30 min−C-peptide t=0 min/glucose t=30 min).
Statistical analysis
The families with Type 1 diabetes mellitus were evaluated by the Sib Transmission Disequilibrium Test (Sib-TDT) [26, 27]. In relatives of Type 2 diabetic patients and healthy glucose-tolerant subjects, differences in continuous variables between carriers of the polymorphism were tested using a general linear model or a mixed model for analysis of variance with age, BMI and sex as covariates. All residuals were tested for normal distribution and proper transformation of the variables was made if necessary (log-transformation). SPSS for Windows version 11.0 (SPSS, Chicago, Ill., USA) or the SAS System for Windows version 8.2 (SAS Institute, Cary, N.C., USA) were used for statistical analysis. We considered p values of less than 0.05 statistically significant. All genotype distributions were tested for Hardy Weinberg equilibrium using a likelihood ratio test.
Results
Mutation scanning
Three novel mutations were identified in the promoter region, but none in the coding or 3′UTR region. Two mutations (C −1044→A and G −202→A) had allele frequencies below 1% in the group of 100 Type 1 diabetic individuals and were not tested further. The third, a C −920→A substitution in the putative promoter had an allele frequency of 8%. Two different software analyses [28, 29] demonstrated that the mutation causes deletion of an activator protein 2 (AP2) transcription factor binding site within the sequence encompassing the polymorphism. AP2 is involved, for example, in angiogenesis, tumour invasion/metastasis, and chronic inflammation [30]. Sib-TDT analysis showed no significant linkage to Type 1 diabetes mellitus in the 250 families with Type 1 diabetes (Table 1). Stratification by high- and low-risk HLA-type also failed to reveal significant differences (data not shown). In addition the dbSNP (rs1061489) in the coding region (Y125H) was genotyped by RFLP in a group of 100 Danish Type 1 diabetic patients. None carried this mutation. Two other dbSNPs (rs 12059, rs 2280148) were not identified in the group screened, preventing construction of haplotypes covering the region.
For offspring of Type 2 diabetic patients and the control subjects, phenotypic data stratified according to genotype are shown in Tables 2 and 3.
In the 370 young healthy Danish Caucasians (Table 3) the SI-values using IVGTT and Bergman minimal modelling were significantly higher among homozygous carriers of the A-allele than among heterozygous and wild-type subjects (AC and CC), despite higher BMI. In the offspring of Type 2 diabetic patients the same trend was found (Table 2). However, due to the low number of homozygous subjects, we were unable to perform any valid statistical analyses in this group of subjects. In the phenotype variables describing beta cell function there was a tendency towards lower acute insulin response (AIR) in the A/A groups with higher SI-values.
Discussion
We performed a mutation scanning of approximately 1000 bp upstream of the 5′UTR region, and of the coding region and approximately 200 bp of the 3′UTR region of the human SOCS3 gene. Three mutations were found in the promoter region of subjects of Danish Caucasoid origin. The most frequent polymorphism was tested in three different groups of individuals. In a population-based family collection no significant linkage to Type 1 diabetes mellitus was observed. Because of the known effects on IRS1 and IRS2 and the fact that prolonged cytokine stimulation leads to insulin resistance [31], this promoter polymorphism was also tested in a group of glucose-tolerant offspring of Type 2 diabetic patients (Group 2) and a group of young healthy subjects (Group 3). Comparing insulin sensitivity values from wild-type, heterozygous and homozygous carriers of the polymorphism in Group 3, the allele dose response suggests a recessive mode of inheritance of this relatively rare insulin-sensitive phenotype. Although the number of homozygous subjects was small, the observation was statistically significant (p=0.027), and a trend towards higher insulin sensitivity in subjects homozygous for the A allele was also found in Group 2. In the phenotype variables describing pancreatic beta cell function, the tendency towards lower AIR in the AA groups with higher SI-values is consistent with the known correlation between these variables. Because the phenotype data are not independent of each other, a conservative Bonferroni correction was not performed.
The presence of the AA genotype is compatible with a protective mechanism against the development of diabetes and even though this group had a higher BMI, insulin sensitivity was above normal. Whether the mutation in the promoter region influences the level of SOCS3 expression in beta cells when exposed to cytokines is still under investigation, but a possible molecular mechanism of the A-allele could be decreased SOCS3 expression leading to less degradation of IRS1 and IRS2 in insulin-sensitive tissues. Functional studies are being carried out to address this possibility. De novo methylation of the GpC island and the following silencing of neighbouring genes could be part of the reason for the effect of SOCS3 overexpression in beta cell lines and this also needs to be evaluated experimentally. Taking the data together, it could be hypothesised that homozygosity for the A-allele is a marker for insulin sensitivity, independent of genetic risk of getting Type 2 diabetes mellitus.
In conclusion, a mutation scanning of the human SOCS3 gene identified no variations associated with Type 1 diabetes mellitus. Homozygosity for a C −920→A promoter polymorphism with a minor allele frequency of 8% was shown to be associated with increased whole-body insulin sensitivity in a population of young healthy unrelated individuals. This finding should be interpreted with caution, and large-scale genetic epidemiology and functional studies are needed to clarify the potential role of this variant in the regulation of whole-body insulin sensitivity.
Abbreviations
- AIR:
-
acute insulin response
- AP2:
-
activator protein 2
- GH:
-
growth hormone
- SI:
-
Insulin sensitivity index
- Sib-TDT:
-
Sib Transmission Disequilibrium Test
- SNP:
-
single nucleotide polymorphism
- SOCS:
-
suppressor of cytokine signalling
- SSCP:
-
single-strand conformational polymorphism
References
Nerup J, Mandrup-Poulsen T, Helqvist S et al. (1994) On the pathogenesis of IDDM. Diabetologia 37:S82–S89
Butler AE, Janson J, Bonner-Weir S, Ritzel R, Rizza RA, Butler PC (2003) Beta-cell deficit and increased beta-cell apoptosis in humans with type 2 diabetes. Diabetes 52:102–110
Esposito K, Nappo F, Giugliano F et al. (2003) Cytokine milieu tends toward inflammation in type 2 diabetes. Diabetes Care 26:1647
Kubaszek A, Pihlajamäki J, Komarovski V et al. (2003) Promoter polymorphisms of the TNF-alpha (G-308A) and IL-6 (C-174G) genes predict the conversion from impaired glucose tolerance to type 2 diabetes: the Finnish Diabetes Prevention Study. Diabetes 52:1872–1876
Spranger J, Kroke A, Möhlig M et al. (2003) Inflammatory cytokines and the risk to develop type 2 diabetes: results of the prospective population-based European Prospective Investigation into Cancer and Nutrition (EPIC)-Potsdam Study. Diabetes 52:812–817
Esposito K, Nappo F, Marfella R et al. (2002) Inflammatory cytokine concentrations are acutely increased by hyperglycemia in humans: role of oxidative stress. Circulation 106:2067–2072
Grimble RF (2002) Inflammatory status and insulin resistance. Curr Opin Clin Nutr Metab Care 5:551–559
Kim F, Gallis B, Corson MA (2001) TNF-alpha inhibits flow and insulin signaling leading to NO production in aortic endothelial cells. Am J Physiol Cell Physiol 280:C1057–C1065
Starr R, Willson TA, Viney EM et al. (1997) A family of cytokine-inducible inhibitors of signalling. Nature 387:917–921
Krebs DL, Hilton DJ (2001) SOCS proteins: negative regulators of cytokine signaling. Stem Cells 19:378–387
Karlsen AE, Rønn SG, Lindberg K et al. (2001) Suppressor of cytokine signaling 3 (SOCS-3) protects beta cells against interleukin-1beta- and interferon-gamma-mediated toxicity. Proc Natl Acad Sci USA 98:12191–12196
Rui L, Yuan M, Frantz D, Shoelson S, White MF (2002) SOCS-1 and SOCS-3 block insulin signaling by ubiquitin-mediated degradation of IRS1 and IRS2. J Biol Chem 277:42394–42398
Luger A, Prager R, Gaube S, Graf H, Klauser R, Schernthaner G (1990) Decreased peripheral insulin sensitivity in acromegalic patients. Exp Clin Endocrinol 95:339–343
Almind K, Doria A, Kahn CR (2001) Putting the genes for type II diabetes on the map. Nat Med 7:277–279
Froguel P, Velho G (2001) Genetic determinants of type 2 diabetes. Recent Prog Horm Res 56:91–105
Kahn CR, Vicent D, Doria A (1996) Genetics of non-insulin-dependent (type-II) diabetes mellitus. Annu Rev Med 47:509–531
Evans JL, Goldfine ID, Maddux BA, Grodsky GM (2002) Oxidative stress and stress-activated signaling pathways: a unifying hypothesis of type 2 diabetes. Endocr Rev 23:599–622
Ohlsson R, Kanduri C (2002) New twists on the epigenetics of CpG islands. Genome Res 12:525–526
Pociot F, Nørgaard K, Hobolth N, Andersen O, Nerup J (1993) A nationwide population-based study of the familial aggregation of type-1 (insulin-dependent) diabetes-mellitus in Denmark. Diabetologia 36:870–875
Lorenzen T, Pociot F, Stilgren L et al. (1998) Predictors of IDDM recurrence risk in offspring of Danish IDDM patients. Diabetologia 41:666–673
Hansen T, Ambye L, Grarup N et al. (2001) Genetic variability of the SUR1 promoter in relation to beta-cell function and Type II diabetes mellitus. Diabetologia 44:1330–1334
Clausen JO, Borch-Johnsen K, Ibsen H et al. (1996) Insulin sensitivity index, acute insulin response, and glucose effectiveness in a population-based sample of 380 young healthy Caucasians. Analysis of the impact of gender, body fat, physical fitness, and life-style factors. J Clin Invest 98:1195–1209
Johannesen J, Pie A, Pociot F, Kristiansen OP, Karlsen AE, Nerup J (2001) Linkage of the human inducible nitric oxide synthase gene to type 1 diabetes. J Clin Endocrinol Metab 86:2792–2796
Larsen ZM, Johannesen J, Kristiansen OP, Nerup J, Pociot F; Danish IDDM Epidemiology and Genetics Group; Danish Study Group of IDDM in Childhood (2004) Evidence for linkage on chromosome 4p16.1 in Type 1 diabetes Danish families and complete mutation scanning of the WFS1 (Wolframin) gene. Diabetic Med 21:218–222
The National Center for Biotechnology Information (NCBI), http://www.ncbi.nlm.nih.gov (latest access 15/12/2003)
Spielman RS, Ewens WJ (1998) A sibship test for linkage in the presence of association: the sib transmission/disequilibrium test. Am J Hum Genet 62:450–458
Spielman RS, Mcginnis RE, Ewens WJ (1993) Transmission test for linkage disequilibrium - the insulin gene region and insulin-dependent diabetes-mellitus (IDDM). Am J Hum Gen 52:506–516
AliBaba version 2.1, http://www.alibaba2.com (latest access 17/12/2003)
MatInspector version 7.0,http://www.genomatix.de/cgi-bin/matinspector_prof/mat_fam (latest access 17/12/2003)
Mertens PR, Alfonso-Jaume MA, Steinmann K, Lovett DH (1998) A synergistic interaction of transcription factors AP2 and YB-1 regulates gelatinase A enhancer-dependent transcription. J Biol Chem 273:32957–32965
Krebs DL, Hilton DJ (2003) A new role for SOCS in insulin action. Sci STKE 2003 (169):PE6
Acknowledgements
This study was supported by the Danish Medical Research Council, the Danish Diabetes Association, Juvenile Diabetes Research Foundation, The Medical Society of Copenhagen, and EU grants (BMH4-CT98-3084 and QLK-CT-2000-01038). We thank the Danish Society for Diabetes in Childhood (DSBD) and the Danish IDDM Epidemiology and Genetics Group for helping identify Type 1 diabetic families. The skilful technical assistance of Bodil Bosmann Jørgensen is highly appreciated.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gylvin, T., Nolsøe, R., Hansen, T. et al. Mutation analysis of suppressor of cytokine signalling 3, a candidate gene in Type 1 diabetes and insulin sensitivity. Diabetologia 47, 1273–1277 (2004). https://doi.org/10.1007/s00125-004-1440-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-004-1440-5