Abstract
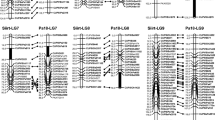
The high versatility of the mode of reproduction and the retention of a pollen recognition system are the factors responsible for the extreme complexity of the genome in Poa pratensis L. Two genetic maps, one of an apomictic and one of a sexual genotype, were constructed using a two-way pseudo-testcross strategy and multiplex PCR-based molecular markers (AFLP and SAMPL). Due to the high ploidy level and the uncertainty of chromosome pairing-behavior at meiosis, only parent-specific single-dose markers (SDMs) that segregated 1:1 in an F1 mapping population (161 out of 299 SAMPLs, and 70 out of 275 AFLPs) were used for linkage analysis. A total of 41 paternal (33 SAMPLs and 8 AFLPs) and 47 maternal (33 SAMPLs and 14 AFLPs) SDMs, tested to be linked in coupling phase, were mapped to 7+7 linkage groups covering 367 and 338.4 cM, respectively. The comparison between the two marker systems revealed that SAMPL markers were statistically more efficient than AFLP ones in detecting parent-specific SDMs (75% vs 32.4%). There were no significant differences in the percentages of distorted marker alleles detected by the two marker systems (27.8% of SAMPLs vs 21.3% of AFLPs). The pairwise comparison of co-segregational groups for linkage detection between marker loci suggested that at least some of the P. pratensis chromosomes pair preferentially at meiosis-I.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 31 August 2000 / Accepted: 12 January 2001
Rights and permissions
About this article
Cite this article
Porceddu, A., Albertini, E., Barcaccia, G. et al. Linkage mapping in apomictic and sexual Kentucky bluegrass (Poa pratensis L.) genotypes using a two way pseudo-testcross strategy based on AFLP and SAMPL markers. Theor Appl Genet 104, 273–280 (2002). https://doi.org/10.1007/s001220100659
Issue Date:
DOI: https://doi.org/10.1007/s001220100659