Abstract
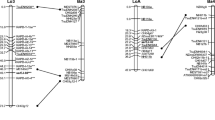
A framework consensus map for rapeseed (Brassica napus L.) was constructed from the integration of three DH mapping populations derived from crosses between or within spring- and winter-type parents. Several sources of genetic markers were used: isozymes, RFLPs, RAPDs, and AFLPs. A total of 992 different markers were mapped to at least one population, of which 540 were included in the consensus map and 253 were common to at least two populations. Markers were distributed over 19 linkage groups, thus reflecting the basic chromosome number of rapeseed and covered 2,429 cM, which was in the mean confidence-interval estimates of genome length (2,127–2,480) cM. Markers were evenly spaced on the entire genome even if, for several linkage groups, both RAPD and AFLP markers were not uniformly distributed. In the population resulting from a cross between two spring lines, a higher recombination rate was observed and a translocation was identified. The consensus approach allowed to map a larger number of markers, to obtain a near-complete coverage of the rapeseed genome, to fill the number of gaps, and to consolidate the linkage groups of the individual maps.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 19 July 2000 / Accepted: 31 October 2000
Rights and permissions
About this article
Cite this article
Lombard, V., Delourme, R. A consensus linkage map for rapeseed (Brassica napus L.): construction and integration of three individual maps from DH populations. Theor Appl Genet 103, 491–507 (2001). https://doi.org/10.1007/s001220100560
Issue Date:
DOI: https://doi.org/10.1007/s001220100560