Abstract
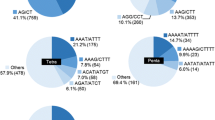
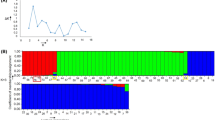
Enrichment methods were optimised in order to isolate large numbers of simple sequence repeat (SSR) markers for perennial ryegrass (Lolium perenne L.), with the aim of developing a comprehensive set of loci for trait mapping and cultivar identification. Two libraries were constructed showing greater than 50% enrichment for a variety of SSR-motif types. Sequence characterisation of 1853 clones identified 859 SSR-containing clones, of which 718 were unique. Truncation of flanking sequences limited potential primer design to 366 clones. One-hundred selected SSR primer pairs were evaluated for amplification and genetic polymorphism across a panel of diverse genotypes. The efficiency of amplification was 81%. A relatively high level of SSR polymorphism was detected (67%), with a range of 2–7 alleles per locus. Mendelian segregation of alleles detected by selected SSR-locus primer pairs was demonstrated in the F1 progeny of a pair cross. Cross-species amplification was detected in a number of related pasture and turfgrass species, with high levels of transfer to other Lolium species and members of the related genus Festuca. The identity of putative SSR ortholoci in these related species was confirmed by DNA sequence analysis. These loci constitute a valuable resource of ideal markers for the molecular breeding of ryegrasses and fescues.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 8 May 2000 / Accepted: 13 June 2000
Rights and permissions
About this article
Cite this article
Jones, E., Dupal, M., Kölliker, R. et al. Development and characterisation of simple sequence repeat (SSR) markers for perennial ryegrass (Lolium perenne L.). Theor Appl Genet 102, 405–415 (2001). https://doi.org/10.1007/s001220051661
Issue Date:
DOI: https://doi.org/10.1007/s001220051661