Abstract
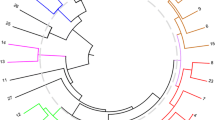
Molecular markers such as RAPDs and microsatellites were used to study genetic diversity in 29 elite Indian chickpea genotypes. In general, microsatellites were more efficient than the RAPD markers in detecting polymorphism in these genotypes. Among the various microsatellites, (AAC)5, (ACT)5, (AAG)5 and (GATA)4 were able to differentiate all 29 chickpea cultivars. The mean value of probability of identical match by chance was 2.32×10-25 using DraI-(ACT)5, TaqI-(AAC)5, TaqI-(AAG)5 and TaqI-(GATA)4 enzyme-probe combinations. The dendrogram, constructed on the basis of similarity index values, grouped the chickpea genotypes into five main clusters with 8 cultivars genetically distant and outgrouped from the main clusters. To investigate if DNA markers are useful in predicting F1 performance and heterosis in chickpea, we crossed 8 genotypes having important agronomic characters in a diallel set. The F1s and their parents in the diallel set were analysed for agronomic traits for better parent and midparent heterosis. Heterosis was found to be much higher for yield than for yield components that fit a multiplicative model. The analysis of genetic divergence using D2 statistics clustered the 8 cultivars into two groups. Although molecular marker-based genetic distance did not linearly correlate to heterosis, two heterotic groups could be identified on the basis of the general marker heterozygosity.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 1 August 1998 / Accepted: 29 September 1998
Rights and permissions
About this article
Cite this article
Sant, V., Patankar, A., Sarode, N. et al. Potential of DNA markers in detecting divergence and in analysing heterosis in Indian elite chickpea cultivars. Theor Appl Genet 98, 1217–1225 (1999). https://doi.org/10.1007/s001220051187
Issue Date:
DOI: https://doi.org/10.1007/s001220051187