Abstract
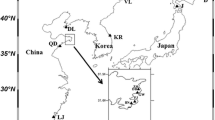
We have investigated the genetic diversity of 11 natural populations of C. japonica using 13 polymorphic STS markers. The average unbiased heterozygosities (H e ), the average number of alleles per locus (N a ) and the proportion of polymorphic loci (Pl) were 0.281, 1.93 and 76.92%, respectively. Coefficients of linkage disequilibrium were calculated, and no significant deviation was found except in four combinations – which might have occurred by chance alone. The fixation index (F IS ) for 3 loci showed statistically significant values at the 1% level. The genetic differentiation between populations was only 0.047, and there were no clear geographical tendencies in the allele frequencies or the heterozygosities among populations. Consequently, the results from STS-based co-dominant DNA marker analysis were very similar to those from a previous allozyme study. However, the resolution of the technique is greater than allozyme analysis because many loci with high heterozygosities can be evaluated, and it is very simple. Therefore, the STS-based marker approach is very useful and convenient for population genetics and genome mapping of C. japonica.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 18 July 1998 / Accepted: 13 August 1998
Rights and permissions
About this article
Cite this article
Tsumura, Y., Tomaru, N. Genetic diversity of Cryptomeria japonica using co-dominant DNA markers based on sequenced-tagged sites. Theor Appl Genet 98, 396–404 (1999). https://doi.org/10.1007/s001220051085
Issue Date:
DOI: https://doi.org/10.1007/s001220051085