Abstract
Key message
GWAS identified eight yield-related, peak starch type of waxy and wild-type starch and 21 starch pasting property-related traits (QTLs). Prediction ability of eight GS models resulted in low to high predictability, depending on trait, heritability, and genetic architecture.
Abstract
Cassava is both a food and an industrial crop in Africa, South America, and Asia, but knowledge of the genes that control yield and starch pasting properties remains limited. We carried out a genome-wide association study to clarify the molecular mechanisms underlying these traits and to explore marker-based breeding approaches. We estimated the predictive ability of genomic selection (GS) using parametric, semi-parametric, and nonparametric GS models with a panel of 276 cassava genotypes from Thai Tapioca Development Institute, International Center for Tropical Agriculture, International Institute of Tropical Agriculture, and other breeding programs. The cassava panel was genotyped via genotyping-by-sequencing, and 89,934 single-nucleotide polymorphism (SNP) markers were identified. A total of 31 SNPs associated with yield, starch type, and starch properties traits were detected by the fixed and random model circulating probability unification (FarmCPU), Bayesian-information and linkage-disequilibrium iteratively nested keyway and compressed mixed linear model, respectively. GS models were developed, and forward predictabilities using all the prediction methods resulted in values of − 0.001–0.71 for the four yield-related traits and 0.33–0.82 for the seven starch pasting property traits. This study provides additional insight into the genetic architecture of these important traits for the development of markers that could be used in cassava breeding programs.










Similar content being viewed by others
Data availability
Phenotypic data used in the analyses and the genotypic data are available in Cassavabase (http://cassavabase.org/search/traits). The imputed SNP genotypic data obtained from the 276 genotypes used in this study are available in Cassavabase.
References
Aiemnaka P, Wongkaew A, Chanthaworn J, Nagashima SK, Boonma S, Authapun J et al (2012) Molecular characterization of a spontaneous waxy starch mutation in cassava. Crop Sci 52:2121–2130. https://doi.org/10.2135/cropsci2012.01.0058
de Albuquerque HYG, Carmo CDD, Brito AC, Oliveira EJD (2018) Genetic diversity of manihot esculenta crantz germplasm based on single-nucleotide polymorphism markers. Annals Appl Biol 173(3):271–284. https://doi.org/10.1111/aab.12460
Aldana AS, Quintero AF (2013) Physicochemical characterization of two cassava (Manihot esculenta Crantz) starches and flours. Rev Sci Agroaliment 1:19–25
Andrade LRB, Sousa MBE, Oliveira EJ, Resende MDV, Azevedo CF (2019) Cassava yield traits predicted by genomic selection methods. PLoS One 14(11):e0224920. https://doi.org/10.1371/journal.pone.0224920
Asoro FG, Newell MA, Beavis WD, Scott MP, Jannink J-L (2011) Accuracy and training population design for genomic selection on quantitative traits in elite North American oats. Plant Gen 4:132–144. https://doi.org/10.3835/plantgenome2011.02.0007
Bates D, Mächler M, Bolker B, Walker S (2015) Fitting linear mixed-effects models using lme4. J Stat Soft 67(1):1–48. https://doi.org/10.18637/jss.v067.i01
Begum H, Spindel JE, Lalusin A, Borromeo T, Gregorio G, Hernandez J et al (2015) Genome-wide association mapping for yield and other agronomic traits in an elite breeding population of tropical rice (Oryza sativa). PLoS ONE 10(3):e0119873. https://doi.org/10.1371/journal.pone.0119873
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) Tassel: software for association mapping of complex traits in diverse samples. Bioinformatics 23(19):2633–2635
BredesonJ V, Lyons JB, Prochnik SE, Wu GA, Ha CM, Edsinger-Gonzales E et al (2016) Sequencing wild and cultivated cassava and related species reveals extensive interspecific hybridization and genetic diversity. Nat Biotechnol 34:562–570. https://doi.org/10.1038/nbt.3535
Breiman L (2001) Random forests. Mach Learn 45:5–32. https://doi.org/10.1023/A:1010933404324
Ceballos H, Iglesias CA, Perez JC, Dixon AG (2004) Cassava breeding: opportunities and challenges. Plant Mol Biol 56:503–516. https://doi.org/10.1007/s11103-004-5010-5
Cach NT, Perez JC, Lenis JI, Calle F, Morante N et al (2005) Epistasis in the expression of relevant traits in cassava (Manihot esculenta Crantz) for subhumid conditions. J Hered 96:586–592
Calle F, Perez JC, Gaitán W, Morante N, Ceballos H, Llano G et al (2005) Diallel inheritance of relevant traits in cassava (Manihot esculenta Crantz) adapted to acid-soil savannas. Euphytica 144:177–186
Ceballos H, Pérez JC, Barandica OJ, Lenis JI, Morante N, Calle F, Pino L, Hershey CH (2016) Cassava breeding I: the value of breeding value. Front Plant Sci 7:1227
Ceballos H, Kawuki RS, Gracen VE, Yencho GC, Hershey CH (2015) Conventional breeding, marker-assisted selection, genomic selection and inbreeding in clonally propagated crops: a case study for cassava. Theor Appl Genet 128:1647–1667
Ceballos H, Rojanaridpiched C, Phumichai C et al (2020) Excellence in cassava breeding: perspectives for the future. Crop Breed Gene Genom 2:e200008
Chaengsee P, Kongsil P, Siriwong N, Kittipadakul P, Piyachomkwan K, Petchpoung K (2020) Potential yield and cyanogenic glucoside content of cassava root and pasting properties of starch and flour from cassava Hanatee var and breeding lines grown under rain-fed condition. Agri Natural Resour 54(3):237–244
Combs E, Bernardo R (2013) Accuracy of genomewide selection for different traits with constant population size, heritability, and number of markers. Plant Genome 6:1–7. https://doi.org/10.3835/plantgenome2012.11.0030
Crossa J, Beyene Y, Kassa S, Pérez P, Hickey JM, Chen C, de los Campos G, Burgueño J, Windhausen VS, Buckler E, Jannink J-L, Lopez Cruz MA, Babu R (2013) Genomic prediction in maize breeding populations with genotyping-by-sequencing. G3 Genes|Genomes|Genetics 3(11):1903–1926. https://doi.org/10.1534/g3.113.008227
Crossa J et al (2014) Genomic prediction in CIMMYT maize and wheat breeding programs. Heredity 112:48–60
Crossa J et al (2017) Genomic selection in plant breeding: methods, models, and perspectives. Trends Plant Sci 22:961–975
Daetwyler HD, Pong-Wong R, Villanueva B, Woolliams JA (2010) The impact of genetic architecture on genome-wide evaluation methods. Genetics 185:1021–1031
de losCampos G, Naya H, Gianola D, Crossa J, Legarra A, Manfredi E (2009) Predicting quantitative traits with regression models for dense molecular markers and pedigree. Genetics 182:375–385. https://doi.org/10.1534/genetics.109.101501
de losCampos G, Gianola D, Rosa GJM, Weigel KA, Cross J (2010) Semi-parametric genomic-enabled prediction of genetic values using reproducing kernel Hilbert spaces methods. Genet Res 92:295–308. https://doi.org/10.1017/S0016672310000285
de losCampos, G., Pérez, P. 2013. BGLR: Bayesian generalized linear regression. R package version 1.0.4. https://cran.r-project.org/web/packages/BGLR
Desta ZA, Ortiz R (2014) Genomic selection: genome-wide prediction in plant improvement. Trends Plant Sci 19:592–601
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (gbs) approach for high diversity species. PloS one 6(5):e19379
Ehret A, Hochstuhl D, Gianola D, Thaller G (2015) Application of neural networks with back-propagation to genome-enabled prediction of complex traits in Holstein-Friesian and German Fleckvieh cattle. Genet Sel Evol 47:22
Elango D, Xue W, Chopra S (2020) Genome wide association mapping of epi-cuticular wax genes in Sorghum bicolor. Physio Mol Biol Plants 26(8):1727–1737. https://doi.org/10.1007/s12298-020-00848-5
Endelman JB (2011) Ridge regression and other kernels for genomic selection with R package rrBLUP. The Plant Genome 4:250–255
Endelman JB, Jannink JL (2012) Shrinkage estimation of the realized relationship matrix. G3 Genes|Genomes|Genetics 2:1405–1413
Evans ID, Lips A (1992) Viscoelasticity of gelatinized starch dispersions. J Texture Stud 23:69–86. https://doi.org/10.1111/j.1745-4603.1992.tb00512.x
Ezenwaka L, Del Carpio DP, Jannink J-L, Rabbi I, Danquah E, Asante I, Danquah A, Blay E, Egesi C (2018) Genome-wide association study of resistance to cassava green mite pest and related traits in cassava. Crop Sci 58:1907–1918
FAO. 2018 Food Outlook - biannual report on global food markets – November 2018. Rome. 104 pp.
Flint-Garcia SA, Thornsberry JM, Buckler ESIV (2003) Structure of linkage disequilibrium in plants. Annu Rev Plant Biol 54:357–374. https://doi.org/10.1146/annurev.arplant.54.031902.134907
Gelandi P, Kowalski BR (1986) Partial least-squares regression: a tutorial. Anal Chim Acta 185:1–17
Glaubitz JC, Casstevens TM, Lu F, Harriman J, Elshire RJ, Sun Q, Buckler ES (2014) Tassel-gbs: a high capacity genotyping by sequencing analysis pipeline. PLoS One 9(2):e90346
González-Camacho JM, Ornella L, Pérez-Rodríguez P, Gianola D, Dreisigacker S, Crossa J (2018) Applications of machine learning methods to genomic selection in breeding wheat for rust resistance. Plant Genome. https://doi.org/10.3835/plantgenome2017.11.0104
Gouy M, Rousselle Y, Bastianelli D et al (2013) Experimental assessment of the accuracy of genomic selection in sugarcane. Theor Appl Genet 126:2575–2586. https://doi.org/10.1007/s00122-013-2156-z
Gracen VE, Kogsil P, Napasintuwong O, Duangjit J, Phumichai C. The story of Kasetsart 50. The most important cassava variety in the world. Bangkok-Kamphaeng Saen (Thailand): Center for Agricultural Biotechnology, Kasetsart University; 2018.
Guo G, Zhao F, Wang Y, Zhang Y, Du L et al (2014) Comparison of single-trait and multiple-trait genomic prediction models. BMC Genet 15:30. https://doi.org/10.1186/1471-2156-15-30
Hayes BJ, Pryce J, Chamberlain AJ, Bowman PJ, Goddard ME (2010) Genetic Architecture of complex traits and accuracy of genomic prediction: coat colour, milk-fat percentage, and type in holstein cattle as contrasting model traits. PLoS Genet 6(9):e1001139. https://doi.org/10.1371/journal.pgen.1001139
Habier D, Fernando R, Kizilkaya K, Garrick D (2011) Extension of the Bayesian alphabet for genomic selection. BMC Bioinformatics 12:186
Heffner EL, Sorrells ME, Jannink J-L (2009) Genomic selection for crop improvement. Crop Sci 49:1–12
Heffner EL, Lorenz AJ, Jannink J-L, Sorrells ME (2010) Plant breeding with genomic selection: gain per unit time and cost. Crop Sci 50:1681–1690
Heslot N, Yang H, Sorrells M, Jannink J (2012) Genomic selection in plant breeding: a comparison of models. Crop Sci 52:146–160. https://doi.org/10.2135/cropsci2011.09.0297
Hickey JM, Chiurugwi T, Mackay I, Powell W (2017) Implementing genomic selection in CBPWP genomic prediction unifies animal and plant breeding programs to form platforms for biological discovery. Nat Genet 49:1297–1303
Howard R, Carriquiry AL, Beavis WD (2014) Parametric and nonparametric statistical methods for genomic selection of traits with additive and epistatic genetic architectures. G3 Genes|Genomes|Genetics 4:1027–1046
Ikeogu UN, Akdemir D, Wolfe MD, Okeke UG, Amaefula C, Jannink J-L, Egesi CN (2019) Genetic correlation, genome-wide association and genomic prediction of portable NIRS predicted carotenoids in cassava roots. Front Plant Sci 10:1570–1570
Isidro J, Jannink JL, Akdemir D, Poland J, Heslot N, Sorrells ME (2015) Training set optimization under population structure in genomic selection. Theor Appl Genet 128(1):145–158. https://doi.org/10.1007/s00122-014-2418-4
Jaramillo G, Morante N, Pérez JC, Calle F, Ceballos H et al (2005) Diallel analysis in cassava adapted to the midaltitude valleys environment. Crop Sci 45:1058–1063
Jannink JL, Lorenz AJ, Iwata H (2010) Genomic selection in plant breeding: from theory to practice. Brief Funct Genomics 9:166–177
Juliana P, Singh RP, Singh PK, Poland JA, Bergstrom GC, Huerta-Espino J, Bhavani S, Crossa J, Sorrells ME (2018) Genome-wide association mapping for resistance to leaf rust, stripe rust and tan spot in wheat reveals potential candidate genes. Theor Appl Genet 131(7):1405–1422. https://doi.org/10.1007/s00122-018-3086-6
Kaler AS, Purcell LC (2019) Estimation of a significance threshold for genome-wide association studies. BMC Genomics 20:618. https://doi.org/10.1186/s12864-019-5992-7
Karlström A, Calle F, Salazar S, Morante N, Dufour D, Ceballos H (2016) Biological implications in cassava for the production of amylose-free starch: impact on root yield and related traits. Front Plant Sci 7:604. https://doi.org/10.3389/fpls.2016.00604
Kawano K, Fukuda WMG, Cenpukdee U (1987) Genetic and environmental effects on dry matter content of cassava root. Crop Sci 27(1):69–74
Kawano K (2003) Thirty years of cassava breeding for productivity – biological and social factors for success. Crop Sci 43:1325–1335. https://doi.org/10.2135/cropsci2003.1325
Kayondo SI, Pino Del Carpio D, Lozano R, Ozimati A, Wolfe M et al (2018) Genome-wide association mapping and genomic prediction for CBSD resistance in Manihot esculenta. Sci Rep 8:1549. https://doi.org/10.1038/s41598-018-19696-1
Kulembeka HP, Ferguson M, Herselman L, Kanju E, Mkamilo G et al (2012) Diallel analysis of field resistance to brown streak disease in cassava (Manihot esculenta Crantz) landraces from Tanzania. Euphytica 187:277–288
Kuon J-E, Qi W, Schläpfer P, Hirsch-Hoffmann M, Rogalla P, von Bieberstein A, Patrignani LP et al (2019) Haplotype-resolved genomes of geminivirus-resistant and geminivirus-susceptible african cassava cultivars. BMC Biol 17(1):75
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Liaw A. 2013 Breiman and Cutler’s random forests for classification and regression. Available 403 at: http://cran.r-project.org/web/packages/randomForest/index.html.
Lipka AE, Tian F, Wang Q, Peiffer J, Li M, Bradbury PJ et al (2012) GAPIT, genome association and prediction integrated tool. Bioinformatics 28:2397–2399. https://doi.org/10.1093/bioinformatics/bts444
Liu X, Huang M, Fan B, Buckler ES, Zhang Z (2016) Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS Genet 12(2):e1005767. https://doi.org/10.1371/journal.pgen.1005767
Lorenz AJ, Smith KP, Jannink J-L (2012) Potential and optimization of genomic selection for Fusarium head blight resistance in six-row barley. Crop Sci 52:1609–1621
Lorenz AJ (2013) Resource allocation for maximizing prediction accuracy and genetic gain of genomic selection in plant breeding: a simulation experiment. G3 Genes|Genomes|Genetics 3:481–91
Maenhout S, De Baets B, Haesaert G, Van Bockstaele E (2007) Support vector machine regression for the prediction of maize hybrid performance. Theor Appl Genet 115:1003–1013
McKey D, Elias M, Pujol B, Duputié A (2010) The evolutionary ecology of clonally propagated domesticated plants. New Phytol 186(2):318–332
Meuwissen THE, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829
Momen M, Mehrgardi AA, Sheikhi A, Kranis A, Tusell L et al (2018) Predictive ability of genome-assisted statistical models under various forms of gene action. Sci Rep 8:12309. https://doi.org/10.1038/s41598-018-30089-2
Morota G, Gianola D (2014) Kernel-based whole-genome prediction of complex traits: a review. Front Gene 5:363
Newport Scientific operation manual of the series 4 rapid visco analyzer. Australia. 1995. p 93
Nwokocha LM, Aviara NA, Senan C, Williams PA (2009) A comparative study of some properties of cassava and cocoyam starches. Carbohydr Polym 76:362–367. https://doi.org/10.1016/j.carbpol.2008.10.034
Ogbonna AC, Braatz de Andrade LR, Mueller LA et al (2021) Comprehensive genotyping of a Brazilian cassava (Manihot esculenta Crantz) germplasm bank: insights into diversification and domestication. Theor Appl Genet. https://doi.org/10.1007/s00122-021-03775-5
Okeke UG, Akdemir D, Rabbi I et al (2017) Accuracies of univariate and multivariate genomic prediction models in African cassava. Genet Sel Evol 49:88. https://doi.org/10.1186/s12711-017-0361-y
Ozimati AR, Kawuki W, Esuma IS, Kayondo M, Wolfe et al (2018) Training population optimization for prediction of cassava brown streak disease resistance in west african clones. G3 Genes|Genomes|Genetics 8:3903–3913
Ozimati A, Kawuki R, Esuma W, Kayondo SI, Pariyo A, Wolfe M, Jannink JL (2019) Genetic variation and trait correlations in an East African cassava breeding population for genomic selection. Crop Sci 59(2):460–473
Park T, Casella G (2008) The Bayesian lasso. J Am Stat Assoc 103:681–686
Pérez JC, Ceballos H, Calle F, Morante N, Gaitán W et al (2005) Within-family genetic variation and epistasis in cassava (Manihot esculenta Crantz) adapted to the acid-soils environment. Euphytica 145:77–85
Pérez P, de Loscampos G (2014) Genome-wide regression and prediction with the bglr statistical package. Genetics 198:483–495. https://doi.org/10.1534/genetics.114.164442
Pérez-Rodríguez P, Gianola D, González-Camacho JM, Crossa J, Manès Y, Dreisigacker S (2012) Comparison between linear and non-parametric regression models for genome-enabled prediction in wheat. G3 Genes|Genomes|Genetics 2(12):1595–1605. https://doi.org/10.1534/g3.112.003665
Pérez L, Soto E, Farré G, Juanos J, Villorbina G, Bassie L, Medina V, Serrato AJ, Sahrawy M, Rojas JA, Romagosa I, Muñoz P, Zhu C, Christou P (2019) CRISPR/Cas9 mutations in the rice Waxy/GBSSI gene induce allele specific and zygosity dependent feedback effects on endosperm starch biosynthesis. Plant cell Reports 38:417–433. https://doi.org/10.1007/s00299-019-02388-z
R Core Team (2017). R: A language and environment for statistical computing. R foundation for statistical computing, Vienna, Austria. URL https://www.R-project.org/.
Rabbi IY, Udoh LI, Wolfe M, Parkes EY, Gedil MA, Dixon A, Ramu P, Jannink J-L, Kulakow P (2017) Genome wide association mapping of correlated traits in cassava: dry matter and total carotenoid content. Plant Genome 10(3):944. https://doi.org/10.3835/plantgenome2016.09.0094
Rabbi IY, Kayondo SI, Bauchet G et al (2020) Genome-wide association analysis reveals new insights into the genetic architecture of defensive, agro-morphological and quality-related traits in cassava. Plant Mol Biol. https://doi.org/10.1007/s11103-020-01038-3
Raemakers K, Schreuder M, Suurs L et al (2005) Improved cassava starch by antisense inhibition of granule-bound starch synthase I. Mol Breed 16:163–172. https://doi.org/10.1007/s11032-005-7874-8
Ramu P, Esuma W, Kawuki R, Rabbi IY, Egesi C, Bredeson JV, Bart RS, Verma J, Buckler ES, Fei Lu (2017) Cassava haplotype map highlights fixation of deleterious mutations during clonal propagation. Nat Genet. https://doi.org/10.1038/ng.3845
Riedelsheimer C et al (2012) Genomic and metabolic prediction of complex heterotic traits in hybrid maize. Nat Genet 44:217–220
Rojanaridpiched, C., V. Vichukit, S. Thongsri, O. Boonseng, A. Limsila and D. Suparhan (2010) Recent progress in cassava breeding and varietal adoption in Thailand. In: Howeler, R. (ed.) A new future for cassava in Asia: its use as food and fuel to benefit the poor. Proceedings of the 8th regional workshop, Vientiane, Lao PDR, pp. 202–210
Rutkoski J, Benson J, Jia Y, Brown-Guedira G, Jannink JL, Sorrells M (2012) Evaluation of genomic prediction methods for Fusarium head blight resistance in wheat. Plant Genome 5:51–61. https://doi.org/10.3835/plantgenome2012.02.0001
Salas-Tovar JA, Flores-Gallegos AC, Contreras-Esquivel JC et al (2017) Analytical methods for pectin methylesterase activity determination: a review. Food Anal Methods 10:3634–3646. https://doi.org/10.1007/s12161-017-0934-y
Sánchez T, Dufour D, Moreno IX, Ceballos H (2010) Comparison of pasting and gel stabilities of waxy and normal starches from potato, maize, and rice with those of a novel waxy cassava starch under thermal, chemical, and mechanical stress. J Agric Food Chem 58:5093–5099
Sauvage C, Segura V, Bauchet G, Stevens R, Do PT, Nikoloski Z, Fernie AR, Causse M (2014) Genome-wide association in tomato reveals 44 candidate loci for fruit metabolic traits. Plant Physiol 165(3):1120–1132. https://doi.org/10.1104/pp.114.241521
Schirmer M, Höchstötter A, Jekle M, Arendt E, Becker T (2013) Physicochemical and morphological characterization of different starches with variable amylose/amylopectin ratio. Food Hydrocoll 32:52–63
Segura V, Vilhjálmsson BJ, Platt A, Korte A, Seren Ü, Long Q, Nordborg M (2012) An efficient multi-locus mixed-model approach for genome-wide association studies in structured populations. Nat Genet 44(7):825–30
Somo M, Kulembeka H, Mtunda K et al (2020) Genomic prediction and quantitative trait locus discovery in a cassava training population constructed from multiple breeding stages. Crop Sci 60:896–913. https://doi.org/10.1002/csc2.20003
Spindel J, Begum H, Akdemir D, Virk P, Collard B, Redoña E et al (2015) Genomic selection and association mapping in rice (Oryza sativa): effect of trait genetic architecture, training population composition, marker number and statistical model on accuracy of rice genomic selection in elite. Tropical Rice Breed Lines Plos Genet 11(2):e1004982. https://doi.org/10.1371/journal.pgen.1004982
Stapleton G. (2012). Global starch market outlook and competing starch raw materials for by product segment and region. Pricing outlook and cassava growth potential. Cassava Starch World 2010. Centre for Management Technology (CMT), Phnom Penh
Su G, Christensen OF, Janss L, Lund MS (2014) Comparison of genomic predictions using genomic relationship matrices built with different weighting factors to account for locus-specific variances. J Dairy Sci 97(10):6547–6559. https://doi.org/10.3168/jds.2014-8210
Svetnik V, Liaw A, Tong C, Culberson JC, Sheridan RP, Feuston BP (2003) Random forest: a classification and regression tool for compound classification and QSAR modeling. J Chem Inf Comput Sci 43:1947–1958
Swarts K, Li H, Romero Navarro JA, An D, Romay MC, Hearne S, Acharya C, Glaubitz JC, Mitchell S, Elshire RJ, Buckler ES, Bradbury PJ (2014) Novel methods to optimize genotypic imputation for low-coverage, next-generation sequence data in crop plants. Plant Genome. https://doi.org/10.3835/plantgenome2014.05.0023
Thanyasiriwat T, Sraphet S, Whankaew S, Boonseng O, Bao J, Lightfoot DA, Tangphatsornruang S, Triwitayakorn K (2014) Quantitative trait loci and candidate genes associated with starch pasting viscosity characteristics in cassava (Manihot esculenta Crantz). Plant Biol (stuttg) 16(1):197–207. https://doi.org/10.1111/plb.12022 (Epub 2013 Apr 24 PMID: 23614826)
Toae R, Sriroth K, Rojanaridpiched C, Vichukit V, Chotineeranat S, Wansuksri R, Chatakanonda P, Piyachomkwan K (2019) Outstanding characteristics of Thai non-gm bred waxy cassava starches compared with normal cassava starch, waxy cereal starches and stabilized cassava starches. Plants 8(11):447. https://doi.org/10.3390/plants8110447
Tibshirani R (1996) Regression shrinkage and selection via the Lasso. J R Stat Soc Ser B Methodol 58:267–288
Tumuhimbise R, Melis R, Shanahan P (2014) Diallel analysis of early storage root yield and disease resistance traits in cassava (Manihot esculenta Crantz). F Crop Res 167:86–93
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423
Wang X, Yang Z, Xu C (2015) A comparison of genomic selection methods for breeding value prediction. Sci Bull 60:925–935
Wang X, Li L, Yang Z, Zheng X, Yu S, Xu C, Hu Z (2017) Predicting rice hybrid performance using univariate and multivariate GBLUP models based on North Carolina mating design II. Heredity 118:302–310
Wickham H (2016). ggplot2: Elegant graphics for data analysis. Springer-Verlag New York. ISBN 978–3–319–24277–4, https://ggplot2.tidyverse.org.
Wolfe MD, Rabbi IY, Egesi C, Hamblin M, Kawuki R, Kulakow P et al (2016) Genome-wide association and prediction reveals the genetic architecture of cassava mosaic disease resistance and prospects for rapid genetic improvement. Plant Genome 9(2):1–13. https://doi.org/10.3835/plantgenome2015.11.0118
Wolfe MD, Kulakow P, Rabbi IY, Jannink J-L (2016b) Marker-based estimates reveal significant nonadditive effects in clonally propagated cassava (Manihot esculenta): implications for the prediction of total genetic value and the selection of varieties. G3 Genes|Genomes|Genetics 6:3497. https://doi.org/10.1534/g3.116.033332
Wolfe MD, Del Carpio DP, Alabi O, Ezenwaka LC, Ikeogu UM, Kayondo IS et al (2017) Prospects for genomic selection in cassava breeding. Plant Genome 10(3):1–19
Wolfe MD, Bauchet GJ, Chan AW, Lozano R, Ramu P et al (2019) Historical introgressions from a wild relative of modern cassava improved important traits and may be under balancing selection. Genetics 213:1237–1253
Xu S (2017) Predicted residual error sum of squares of mixed models: an application for genomic prediction. G3 Genes|Genomes|Genetics 7:895–909
Xu S, Zhu D, Zhang Q (2014) Predicting hybrid performance in rice using genomic best linear unbiased prediction. Proc Natl Acad Sci USA 111:12456–12461
Xu Y, Wang X, Ding X, Zheng X, Yang Z, Xu C, Hu Z (2018) Genomic selection of agronomic traits in hybrid rice using an NCII population. Rice 10, 11(1):32. https://doi.org/10.1186/s12284-018-0223-4. PMID: 29748895; PMCID:PMC5945574
Yabe S, Hara T, Ueno M, Enoki H, Kimura T, Nishimura S, Yasui Y, Ohsawa R, Iwata H (2018) Potential of genomic selection in mass selection breeding of an allogamous crop: an empirical study to increase yield of common buckwheat. Front Plant Sci. https://doi.org/10.3389/fpls.2018.00276
Yasui T, Ashida K (2011) Waxy endosperm accompanies increased fat and saccharide contents in bread wheat (Triticum aestivum L.) grain. Cereal Sci 53:104–111. https://doi.org/10.1016/j.jcs.2010.10.004
Yang N, Lu Y, Yang X, Huang J, Zhou Y, Ali F et al (2014) Genome wide association studies using a new nonparametric model reveal the genetic architecture of 17 agronomic traits in an enlarged maize association panel. PLoS Genet 10(9):e1004573. https://doi.org/10.1371/journal.pgen.1004573
Yin L, Zhang H, Tang Z, Xu J, Yin D, Zhang Z, Yuan X, Zhu M, Zhao S, Li X (2021) rMVP: a memory-efficient, visualization-enhanced, and parallel-accelerated tool for genome-wide association study, genomics. Proteom Bioinform. https://doi.org/10.1016/j.gpb.2020.10.007
Yonis BO, Pino del Carpio D, Wolfe M et al (2020) Improving root characterisation for genomic prediction in cassava. Sci Rep 10:8003. https://doi.org/10.1038/s41598-020-64963-9
Yun MS, Kawagoe Y (2009) Amyloplast division progresses simultaneously at multiple sites in the endosperm of rice. Plant Cell Physiol 50(9):1617–1626. https://doi.org/10.1093/pcp/pcp104
Zacarias AM, Labuschagne MT (2010) Diallel analysis of cassava brown streak disease, yield and yield related characteristics in Mozambique. Euphytica 176:309–320
Zhang Z, Ersoz E, Lai CQ, Todhunter RJ, Tiwari HK, Gore MA, Bradbury PJ, Yu J, Arnett DK, Ordovas JM (2010) Buckler ES. Nat Genet 42(4):355–360
Zhang N et al (2015) Genome-wide association of carbon and nitrogen metabolism in the maize nested association mapping population. Plant Physiol 168:575–583
Zhang S, Chen X, Lu C, Ye J, Zou M, Lu K, Feng S, Pei J, Liu C, Zhou X, Ma P, Li Z, Liu C, Liao Q, Xia Z, Wang W (2018) Genome-wide association studies of 11 agronomic traits in cassava (Manihot esculenta Crantz). Front Plant Sci 9:503. https://doi.org/10.3389/fpls.2018.00503
Acknowledgements
We would like to thank James Tanaka for his assistance in the laboratory of Dr. Mark Sorrells.
Funding
This work was supported by Kasetsart University Research and Development Institute (KURDI), Grant Number รหัส ศ-ข(กษ)2.57, and was partially supported by the Center of Excellence on Agricultural Biotechnology, Office of the Permanent Secretary, Ministry of Higher Education, Science, Research and Innovation (AG-BIO/MHESI) Grant Number (60-005-001)J. The authors would also like to thank TTDI for providing supporting plant materials.
Author information
Authors and Affiliations
Contributions
CP and JC designed the experiment. PA, PN, SH, PJK, PK, WW, and CP conducted the field experiments. CR and VV provided germplasm. CK, SC, and KP conducted the starch pasting properties experiments. CP, PF, and PT performed data analysis. CP, SC, and PJK wrote the first draft of the manuscript. CP, MS, JJ, and MW revised the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Damaris Odeny.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
122_2021_3956_MOESM2_ESM.pdf
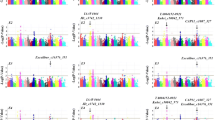
Manhattan and quantile–quantile plots (QQ) plots comparing different yield-related traits data using compressed mixed linear (CMLM) model (PDF 194 KB)
122_2021_3956_MOESM3_ESM.pdf
Manhattan and quantile–quantile plots (QQ) plots comparing different yield-related traits data using multi‐locus mixed model (MLMM) (PDF 85 KB)
122_2021_3956_MOESM4_ESM.pdf
Manhattan and quantile–quantile plots (QQ) plots comparing different waxy and wild-type starch cassava using compressed mixed linear (CMLM) model, multi‐locus mixed model (MLMM) model, and fixed and random model circulating probability unification (FarmCPU) model (PDF 61 KB)
Rights and permissions
About this article
Cite this article
Phumichai, C., Aiemnaka, P., Nathaisong, P. et al. Genome-wide association mapping and genomic prediction of yield-related traits and starch pasting properties in cassava. Theor Appl Genet 135, 145–171 (2022). https://doi.org/10.1007/s00122-021-03956-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-021-03956-2