Abstract
Epistasis is defined as interactions between alleles of two or more genetic loci. Detection of epistatic interactions is the key to understand the genetic architecture and gene networks underlying complex traits. Here, we examined the extent of epistasis for seven quantitative traits with an association mapping approach in a large population of elite sugar beet lines. We found that correction for population stratification is required and that in terms of reducing the false-positive rate the mixed model approach including the kinship matrix performed best. In genome-wide scans, we detected both main effects and epistatic QTL. For physiological traits, the detected digenic and higher-order epistasis explained a considerable proportion of the genotypic variance. We illustrate that the identified epistatic interactions define comprehensive genetic networks, which may serve as starting points towards a systems-oriented approach to understand the regulation of complex traits.






Similar content being viewed by others
References
Bernardo R (1993) Estimation of coefficient of coancestry using molecular markers in maize. Theor Appl Genet 85:1055–1062
Brem RB, Kruglyak L (2005) The landscape of genetic complexity across 5.700 gene expression traits in yeast. Proc Natl Acad Sci USA 102:1572–1577
Carlborg Ö, Haley CS (2004) Epistasis: too often neglected in complex trait studies? Nat Rev Genet 5:618–625
Carlborg Ö, Jacobsson L, Ahgren P, Siegel P, Andersson L (2006) Epistasis and the release of genetic variation during long-term selection. Nat Genet 38:418–420
Chao S, Zhang W, Dubcovsky J, Sorrells ME (2007) Evaluation of genetic diversity and genome-wide linkage disequilibrium among U.S. wheat (Triticum aestivum L.) germplasm representing different market classes. Crop Sci 47:1018–1030
Costanzo M, Baryshnikova A, Bellay J, Kim Y, Spear ED et al (2010) The genetic landscape of a cell. Science 327:425–431
Draycott AP (2006) Sugar beet, 1st edn. Blackwell, Oxford
Eshed Y, Zamir D (1996) Less-than-additive epistatic interactions of quantitative trait loci in tomato. Genetics 143:1807–1817
Gilmour AR, Gogel BJ, Cullis BR, Thompson R (2006) ASReml User Guide Release 2.0 VSN International Ltd, Hermel Hempstead, UK
He X, Qian W, Wang Z, Li Y, Zhang J (2010) Prevalent positive epistasis in Escherichia coli and Saccharomyces cerevisiae metabolic networks. Nat Genet 42:272–276
Hill WG, Robertson A (1968) Linkage disequilibrium in finite populations. Theor Appl Genet 38:226–231
Holm S (1979) A simple sequentially rejective multiple test procedure. Scand J Stat 6:65–70
Kraakman ATW, Niks RE, Van den Berg PMMM, Stam P, Van Eeuwijk FA (2004) Linkage disequilibrium mapping of yield and yield stability in modern spring barley cultivars. Genetics 168:435–446
Li Z, Pinson SR, Park WD, Paterson AH, Stansel JW (1997) Epistasis for three grain yield components in rice (Oryza sativa L.). Genetics 145:453–465
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits. Sinauer Assoc., Sunderland
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Maurer HP, Melchinger AE, Frisch M (2008) Population genetic simulation and data analysis with Plabsoft. Euphytica 161:133–139
Melchinger AE, Utz HF, Schön CC (1998) Quantitative trait locus (QTL) mapping using different testers and independent population samples in maize reveals low power of QTL detection and larger bias in estimates of QTL effects. Genetics 149:383–403
Montooth KL, Marden JH, Clark AG (2003) Mapping determinants of variation in energy metabolism, respiration and flight in Drosophila. Genetics 165:623–635
Phillips PC (2008) Epistasis—the essential role of gene interactions in the structure and evolution of genetic systems. Nat Rev Genet 9:855–867
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA et al (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Schadt EE, Monks SA, Drake TA, Lusis AJ, Che N et al (2003) Genetics of gene expression surveyed in maize, mouse and man. Nature 422:297–302
Schneider K, Schäfer-Pregl R, Borchardt C, Salamini F (2002) Mapping QTLs for sucrose content, yield and quality in a sugar beet population fingerprinted by EST-related markers. Theor Appl Genet 104:1107–1113
St Onge RP, Mani R, Oh J, Proctor M, Fung E et al (2007) Systematic pathway analysis using high-resolution fitness profiling of combinatorial gene deletions. Nat Genet 39:199–206
Stich B, Melchinger AE, Frisch M, Maurer HP, Heckenberger M et al (2005) Linkage disequilibrium in European elite maize germplasm investigated with SSRs. Theor Appl Genet 111:723–730
Stich B, Melchinger AE, Heckenberger M, Möhring J, Schechert A et al (2008a) Association mapping in multiple segregating populations of sugar beet (Beta vulgaris L.). Theor Appl Genet 117:1167–1179
Stich B, Möhring J, Piepho HP, Heckenberger M, Buckler ES et al (2008b) Comparison of mixed-model approaches for association mapping. Genetics 178:1745–1754
Stich B, Yu J, Melchinger AE, Piepho HP, Utz HF et al (2008c) Power to detect higher-order epistatic interactions in a metabolic pathway using a new mapping strategy. Genetics 176:563–570
Stram DO, Lee JW (1994) Variance components testing in longitudinal mixed effects model. Biometrics 50:1171–1177
Tong AH, Lesage G, Bader GD, Ding H, Xu H et al (2004) Global mapping of the yeast genetic interaction network. Science 303:808–813
Utz HF, Melchinger AE, Schön CC (2000) Bias and sampling error of the estimated proportion of genotypic variance explained by quantitative trait loci determined from experimental data in maize using cross validation and validation with independent samples. Genetics 154:1839–1849
Weber WE, Borchardt DC, Koch G (2000) Marker analysis for quantitative traits in sugar beet. Plant Breed 119:97–106
Weir BS (1996) Genetic data analysis II, 2nd edn. Sinauer Associates, Sunderland
Wright S (1978) Evolution and genetics of populations, variability within and among natural populations, vol 4. The University of Chicago Press, Chicago, p 91
Xu S, Jia Z (2007) Genomewide analysis of epistatic effects for quantitative traits in barley. Genetics 175:1955–1963
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M et al (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zhao K, Aranzana MJ, Kim S, Lister C, Shindo C et al (2007) An Arabidopsis example of association mapping in structured samples. PLoS Genet 3:e4
Acknowledgments
This research was conducted within the Biometric and Bioinformatic Tools for Genomics based Plant Breeding project supported by the German Federal Ministry of Education and Research (BMBF) within the framework of GABI-FUTURE initiative.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by F. van Eeuwijk.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2011_1570_MOESM1_ESM.eps
Supplementary Figure S1 The 22 parents (1-22) with their male and female grandparents (23-40) are shown and the number of genotypes in the data set to which these parents contributed either as male or as female parent (progeny) (EPS 674 kb)
122_2011_1570_MOESM2_ESM.eps
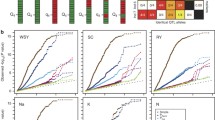
Supplementary Figure S2 Plot of observed vs. expected P values for the epistasis scan with the model including the kinship matrix (K). (WSY, white sugar yield, SY, sugar yield, SC, sugar content, BY, beet yield, K, potassium, Na, sodium, N, α-amino nitrogen) (EPS 3480 kb)
122_2011_1570_MOESM3_ESM.eps
Supplementary Figure S3 (A) 2-way epistatic QTL and (B) 3-way epistatic QTL. The chromosomes are depicted as black lines. Interacting loci are connected by lines where the line strength indicates the size of the QTL effects for white sugar yield (WSY), sugar yield (SY), sugar content (SC), beet yield (BY), potassium (K), sodium (Na), and α-amino nitrogen (N). (EPS 1257 kb)
Rights and permissions
About this article
Cite this article
Würschum, T., Maurer, H.P., Schulz, B. et al. Genome-wide association mapping reveals epistasis and genetic interaction networks in sugar beet. Theor Appl Genet 123, 109–118 (2011). https://doi.org/10.1007/s00122-011-1570-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-011-1570-3