Abstract
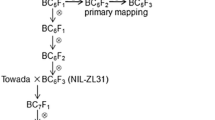
Low temperature at the booting stage is a serious abiotic stress in rice, and cold tolerance is a complex trait controlled by many quantitative trait loci (QTL). A QTL for cold tolerance at the booting stage in cold-tolerant near-isogenic rice line ZL1929-4 was analyzed. A total of 647 simple sequence repeat (SSR) markers distributed across 12 chromosomes were used to survey for polymorphisms between ZL1929-4 and the cold-sensitive japonica cultivar Towada, and nine were polymorphic. Single marker analysis revealed that markers on chromosome 7 were associated with cold tolerance. By interval mapping using an F2 population from ZL1929-4 × Towada, a QTL for cold tolerance was detected on the long arm of chromosome 7. The QTL explained 9 and 21% of the phenotypic variances in the F2 and F3 generations, respectively. Recombinant plants were screened for two flanking markers, RM182 and RM1132, in an F2 population with 2,810 plants. Two-step substitution mapping suggested that the QTL was located in a 92-kb interval between markers RI02905 and RM21862. This interval was present in BAC clone AP003804. We designated the QTL as qCTB7 (quantitative trait locus for cold tolerance at the booting stage on chromosome 7), and identified 12 putative candidate genes.


Similar content being viewed by others
References
Andaya VC, Mackill DJ (2003) QTLs conferring cold tolerance at the booting stage of rice using recombinant inbred lines from a japonica × indica cross. Theor Appl Genet 106:1084–1090
Ashikari M, Sakakibara H, Lin S, Yamamoto T, Takashi T, Nishimura A, Angeles ER, Qian Q, Kitano H, Matsuoka M (2005) Cytokinin oxidase regulates rice grain production. Science 309:741–745
Brzobohaty B, Moore I, Kristoffersen P, Bako L, Campos N, Schell J, Palme K (1993) Release of active cytokinin by a β-glucosidase localized to the maize root meristem. Science 262:1051–1054
Chen W, Provart NJ, Glazebrook J, Katagiri F, Chang HS, Eulgem T, Mauch F, Sh Luan, Zou G, Whitham SA, Budworth PR, Tao Y, Xie Z, Chen X, Lam S, Kreps JA, Harper JF, Si-Ammour A, Mauch-Mani B, Heinlein M, Kobayashi K, Hohn T, Dangl JL, Wang X, Zhu T (2002) Expression profile matrix of arabidopsis transcription factor genes suggests their putative functions in response to environmental stresses. Plant Cell 14:559–574
Cheng KS (1993) Rice genetic resources in Yunnan. In: Wu Zhengyi symposium on biodiversity in Yunnan. Yunnan Province Science and Technology Press, Kunming, China, pp 90–94
Chong J, Baltz R, Schmitt C, Beffa R, Fritig B, Saindrenan P (2002) Downregulation of a pathogen-responsive tobacco UDP-Glc: phenylpropanoid glucosyltransferase reduces scopoletin glucoside accumulation, enhances oxidative stress, and weakens virus resistance. Plant Cell 14:1093–1107
Cicek M, Esen A (1998) Structure and expression of a dhurrinase (b-glucosidase) from sorghum. Plant Physiol 116:1469–1478
Dai LY, Kariya K, Ye CR, Ise K, Tanno HI, Yu TQ, Xu FR (2002) Studies on cold tolerance of rice, Oryza sativa L. II. Evaluation on cold tolerance of Yunnan rice genetic resources. Southwest China J Agri Sci 15:47–52
Dai LY, Lin XH, Ye CR, Ise K, Saito K, Kato A, Xu FR, Yu TQ, Zhang DP (2004) Identification of quantitative trait loci controlling cold tolerance at the reproductive stage in Yunnan landrace of Rice, Kunmingxiaobaigu. Breeding Sci 54:253–258
Ding X, Cao Y, Huang L, Zhao J, Xu C, Li X, Wang S (2008) Activation of the indole-3-acetic acid–amido synthetase GH3-8 suppresses expansin expression and promotes salicylate- and jasmonate-independent basal immunity in rice. Plant Cell 20:228–240
Dong CH, Agarwal M, Zhang Y, Xie Q, Zhu JK (2006) The negative regulator of plant cold responses, HOS1, is a RING E3 ligase that mediates the ubiquitination and degradation of ICE1. Proc Natl Acad Sci USA 103:8281–8286
Esen A (1993) β-glucosidases: biochemistry and molecular biology. Oxford University Press, Oxford
Futsuhara Y, Toriyama K (1966) Genetic studies on cool tolerance in rice. III. Linkage relations between genes controlling cool tolerance and marker genes of Nagao and Takahashi. Jpn J Breed 16:19–30
Gerardi C, Blando F, Santino A, Zacheo G (2001) Purification and characterisation of a beta-glucosidase abundantly expressed in ripe sweet cherry (Prunus avium L.) fruit. Plant Sci 160:795–805
Glaszmann JC, Kaw RN, Khush GS (1990) Genetic divergence among cold tolerant rices (Oryza sativa L.). Euphytica 45:95–104
Guilfoyle TJ (1999) Auxin-regulated genes and promoters. In: Hooykaas PJJ, Hall MA, Libbenga KR (eds) Biochemistry and molecular biology of plant hormones. Elsevier, Amsterdam, pp 423–459
Hannah MA, Heyer AG, Hincha DK (2005) A global survey of gene regulation during cold acclimation in Arabidopsis thaliana. PLoS Genet 1:179–196
Hayase H, Satake T, Nishiyama I, Ito N (1969) Male sterility caused by cooling treatment at the young microspore stage in rice plants. II. The most sensitive stage to cooling and the fertilizing ability of pistils. Proc Crop Sci Soc Jpn 38:706–711
He GM, Luo XJ, Tian F, Li KG, Zhu ZF, Su W, Qian XY, Fu YC, Wang XK, Sun CQ, Yang JS (2006) Haplotype variation in structure and expression of a gene cluster associated with a quantitative trait locus for improved yield in rice. Genome Res 16:618–626
Horisue N, Kunihiro Y, Higashi T, Oyamada Z, Wang H, Xiong J, Zhang S, Li Z, Wang Y (1988) Screening for cold tolerance of Chinese and Japanese rice varieties and selection of standard varieties. Trop Agric Res Ser 21:76–87
Inc SPSS (2002) SPSS 11.0 for Macintosh. SPSS Inc, Chicago
Institute SAS (2000) SAS user’s guide version 8.0. SAS Institute, Cary
International Rice Genome Sequencing Project (2005) The map-based sequence of the rice genome. Nature 436:793–799
Ishitani M, Xiong L, Lee H, Stevenson B, Zhu JK (1998) HOS1, a genetic locus involved in cold-responsive gene expression in Arabidopsis. Plant Cell 10:1151–1162
Jain M, Khurana JP (2009) Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J 276:3148–3162
Jain M, Kaur N, Tyagi AK, Khurana JP (2006) The auxin-responsive GH3 gene family in rice (Oryza sativa). Funct Integr Genomics 6:36–46
Konishi S, Izawa T, Lin SY, Ebana K, Fukuta Y, Sasaki T, Yano M (2006) An SNP caused loss of seed shattering during rice domestication. Science 312:1306–1392
Kuroki M, Saito K, Matsuba S, Yokogami N, Shimizu H, Ando I, Sato Y (2007) A quantitative trait locus for cold tolerance at the booting stage on rice chromosome 8. Theor Appl Genet 115:593–600
Li TG, Guo WM (1993) Identification and study on tolerance in main stresses of China cultivated rice germplasm resource. In: Ying CS (ed) Rice germplasm resources in China. China Agricultural Science and Technology Press, Beijing, pp 71–75
Li HB, Wang J, Liu AM, Liu KD, Zhang Q, Zou JS (1997) Genetic basis of low- temperature-sensitive sterility in indica-japonica hybrids of rice as determined by RFLP analysis. Theor Appl Genet 95:1092–1097
Li SC, Han JW, Chen KC, Chen CS (2001) Purification and characterization of isoforms of b-galactosidases in mung bean seedlings. Phytochemistry 57:349–359
Lincoln S, Daly M, Lander E (1992) Whitehead Institute Technical Report, Whitehead Institute, Cambridge, MA, USA
Liu FX, Sun CQ, Tan LB, Fu YC, Li DJ, Wang XK (2003) Identification and mapping of quantitative trait loci controlling cold-tolerance of Chinese common wild rice (O. rufipogon Griff.) at booting to flowering stages. Chin Sci Bull 48:2068–2071
Manly KF, Cudmore RH, Meer JM (2001) Map Manager QTX: cross-platform software for genetic mapping. Mamm Genome 12:930–932
McCouch SR, Kochert G, Yu ZH, Wang ZY, Khush GS, Coffman WR, Tanksley SD (1988) Molecular mapping of rice chromosomes. Theor Appl Genet 76:815–829
McCouch SR, Teytelman L, Xu Y, Lobos KB, Clare K, Walton M, Fu B, Maghirang R, Li Z, Xing Y, Zhang Q, Kono I, Yano M, Fjellstrom R, DeClerck G, Schneider D, Cartinhour S, Ware D, Stein L (2002) Development of 2,240 new SSR markers for rice (Oryza sativa L.). DNA Res 9:199–207
Nishiyama I (1976) Male sterility caused by cooling treatment at the young microspore stage in rice plants. XIII. Ultrastructure of tapetal hypertrophy without primary wall. Proc Crop Sci Soc Jpn 45:270–278
Nishiyama I (1982) Male sterility caused by cooling treatment at the young microspore stage in rice plants. XXIII. Anther length, pollen number and the difference in susceptibility to coolness among spikelets on the panicle. Jpn J Crop Sci 51:462–469
Nishiyama I (1995) Damage due to extreme temperatures. In: Matsuo T, Kumazawa K, Ishii R, Ishihara H, Hirata H (eds) Science of the rice plant. Food and Agriculture Policy Research Centre, Tokyo, Japan, pp 769–812
Paterson AH, De-Verna JW, Lanini B, Tanksley SD (1990) Fine mapping of quantitative trait loci using selected overlapping recombinant chromosomes, in an interspecific cross of tomato. Genetics 124:735–742
Rogers OS, Bendich AJ (1988) Extraction of DNA from plant tissues. Plant Mol Biol Manual A6:1–10
Saito K, Miura K, Nagano K, Hayano-saito Y, Araki H, Kato A (2001) Identification of two closely linked quantitative trait loci for cold tolerance on chromosome 4 of rice and their association with anther length. Theor Appl Genet 103:862–868
Saito K, Miura K, Hayanosaito Y, Maruyama-Funatsuki W, Sato Y, Kato A (2004) Physical mapping and putative candidate gene identification of a quantitative trait locus Ctb1 for cold tolerance at the booting stage of rice. Theor Appl Genet 109:515–522
Satake T (1989) Male sterility caused by cooling treatment at the young microspore stage in rice plants, XXIX. The mechanism of enhancement in cool tolerance by raising water temperature before the critical stage. Jpn J Crop Sci 8:240–245
Satake T, Hayase H (1970) Male sterility caused by cooling treatment at the young microspore stage in rice plants. V. Estimation of pollen developmental stage and the most sensitive stage to coolness. Proc Crop Sci Soc Jpn 39:468–473
Song Y, Wang L, Xiong L (2009) Comprehensive expression profiling analysis of OsIAA gene family in developmental processes and in response to phytohormone and stress treatments. Planta 229:577–591
Spano G, Rinaldi A, Ugliano M, Moio L, Beneduce L, Massa S (2005) A β-glucosidase gene isolated from wine Lactobacillus plantarum is regulated by abiotic stresses. J Appl Microbiol 98:855–861
Staswick PE, Tiryaki I, Rowe ML (2002) Jasmonate response locus JAR1 and several related Arabidopsis genes encode enzymes of the firefly luciferase superfamily that show activity on jasmonic, salicylic, and indole-3-acetic acids in an assay for adenylation. Plant Cell 14:1405–1415
Staswick PE, Serban B, Rowe M, Tiryaki I, Maldonado MT, Maldonado MC, Suza W (2005) Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell 17:616–627
Sue M, Ishihara A, Iwamura H (2000) Purification and characterization of a hydroxamic acid glucoside b-glucosidase from wheat (Triticum aestivum L.) seedlings. Planta 210:432–438
Suh JP, Jeung JU, Lee JI, Choi YH, Yea JD, Virk PS, Mackill DJ, Jena KK (2009) Identification and analysis of QTLs controlling cold tolerance at the reproductive stage and validation of effective QTLs in cold-tolerant genotypes of rice (Oryza sativa L.). Theor Appl Genet. doi:10.1007/s00122-009-1226-8
Takeuchi Y, Hayasaka H, Chiba B, Tanaka I, Shimano T, Yamagishi M, Nagano K, Sasaki T, Yano M (2001) Mapping quantitative trait loci controlling cool-temperature tolerance at booting stage in temperate japonica rice. Breed Sci 51:191–197
Temnykh S, Park WD, Ayres N, Ayres N, Cartinhour S, Hauck N, Lipovich L, Cho YG, Ishii T, McCouch SR (2000) Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theor Appl Genet 100:697–712
Thorlby G, Fourrier N, Warren G (2004) The SENSITIVE TO FREEZING2 gene, required for freezing tolerance in Arabidopsis thaliana, encodes a beta-glucosidase. Plant Cell 16:2192–2203
Ueda T, Sato T, Hidema J, Hirouchi T, Yamamoto K, Kumagai T, Yano M (2005) qUVR-10, a major quantitative trait locus for ultraviolet-B resistance in rice, encodes cyclobutane pyrimidine dimer photolyase. Genetics 171:1941–1950
Wang XS, Zhao XQ, Zhu J, Wu WR (2005) Genomewide investigation of intron length polymorphisms and their potential as molecular markers in rice (Oryza sativa L.). DNA Res 12:417–427
Xu LM, Zhou L, Zeng YW, Wang FM, Zhang HL, Shen SQ, Li ZC (2008) Identification and mapping of quantitative trait loci for cold tolerance at the booting stage in a japonica rice near-isogenic line. Plant Sci 174:340–347
Zeng YW, Li SC, Pu XY, Du J, Yang SM, Liu K, Gui M, Zhang H (2006) Ecological difference and correlation among cold tolerance traits at the booting stage for core collection of rice landrace in Yunnan, China. Chin J Rice Sci 20:265–271
Zhou GA, Chang RZ, Qiu LJ (2010) Overexpression of soybean ubiquitin-conjugating enzyme gene GmUBC2 confers enhanced drought and salt tolerance through modulating abiotic stress-responsive gene expression in Arabidopsis. Plant Mol Biol 72:357–367
Acknowledgments
We thank Professor Robert A McIntosh, University of Sydney, for critically reading and revising the manuscript. This work was funded by the “973” Project of the Ministry of Sciences and Technology of China (2010CB125900-G), the 863 project (2006AA10Z158, 2006AA100101), the National Natural Science Foundation of China (30571140), the Key Project of Transgenic Crop Improvement in China (2009ZX08009-073B) and China National Key Technologies R&D Program (2009BADA2B01, 2006BAD13B01-6).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. Close.
L. Zhou and Y. Zeng contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhou, L., Zeng, Y., Zheng, W. et al. Fine mapping a QTL qCTB7 for cold tolerance at the booting stage on rice chromosome 7 using a near-isogenic line. Theor Appl Genet 121, 895–905 (2010). https://doi.org/10.1007/s00122-010-1358-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-010-1358-x