Abstract.
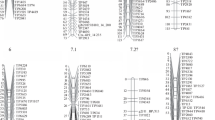
Fifty sequence-tagged microsatellite site (STMS) markers and a resistant gene-analog (RGA) locus were integrated into a chickpea (Cicer arietinum L., 2n = 2x = 16 chromosomes) genetic map that was previously constructed using 142 F6-derived recombinant inbred lines (RILs) from a cross of C. arietinum × Cicer reticulatum Lad. The map covers 1,174.5 cM with an average distance of 7.0 cM between markers in nine linkage groups (LGs). Nine markers including the RGA showed distorted segregation (P < 0.05). The majority of the newly integrated markers were mapped to marker-dense regions of the LGs. Six co-dominant STMS markers were integrated into two previously reported major quantitative trait loci (QTLs) conferring resistance to Ascochyta blight caused by Ascochyta rabiei (Pass.) Labr. Using common STMS markers as anchors, three maps developed from different mapping populations were joined, and genes for resistance to Ascochyta blight, Fusarium wilt (caused by Fusarium oxysporum Schlechtend.: Fr. f. sp. ciceris), and for agronomically important traits were located on the combined linkage map. The integration of co-dominant STMS markers improves the map of chickpea and makes it possible to consider additional fine mapping of the genome and also map-based cloning of important disease resistance genes.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Tekeoglu, .M., Rajesh, .P. & Muehlbauer, .F. Integration of sequence tagged microsatellite sites to the chickpea genetic map. Theor Appl Genet 105, 847–854 (2002). https://doi.org/10.1007/s00122-002-0993-2
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00122-002-0993-2