Abstract
Fibroblast growth factor 2 (FGF2) plays an important role in the development of osteoarthritis (OA) through the regulation of cartilage degradation. However, the molecular mechanism underlying FGF2-induced OA is poorly characterized. MicroRNAs (miRNAs) maintain cartilage homeostasis. To examine whether FGF2 regulates OA through the modulation of miRNA, we screened potential miRNA molecules that could be regulated through FGF2 using microarray analysis. The results showed that microRNA-105 (miR-105) was significantly downregulated in chondrocytes stimulated with FGF2. Runt-related transcription factor 2 (Runx2), a key transcription factor involved in OA, has been identified as a novel potential target of miR-105. FGF2 suppressed miR-105 expression through the recruitment of the subunit of the nuclear factor kappa B transcription complex p65 to the miR-105 promoter. The knockdown of Runx2 mimicked the effect of miR-105 and abolished the ability of miR-105 to regulate the expression of a disintegrin-like and metalloproteinase with thrombospondin 4 (ADAMTS4), ADAMTS5, ADAMTS7 and ADAMTS12, both of which are responsible for the degradation of collagen 2A1 (COL2A1) and aggrecan (ACAN). miR-105 is also required for FGF2/p65-induced Runx2 activation and ADAMTS expression. Moreover, miR-105 expression was downregulated in OA patients and inversely correlated with the expression of Runx2, ADAMTS7 and ADAMTS12, which were upregulated in OA patients. These data highlight that the FGF2/p65/miR-105/Runx2/ADAMTS axis might play an important role in OA pathogenesis and that miR-105 might be a potential diagnostic target and useful strategy for OA treatment.
Key message
-
Runx2 was identified as a novel direct target of miR-105.
-
FGF2 inhibits miR-105 transcription through recruitment of p65 to miR-105 promoter.
-
p65/miR-105 is essential for FGF2-mediated Runx2 and ADAMTS upregulation.
-
miR-105 is downregulated in OA and inversely correlated with Runx2 expression.






Similar content being viewed by others
References
Conaghan PG, Kloppenburg M, Schett G, Bijlsma JW, EULAR osteoarthritis ad hoc committee (2014) Osteoarthritis research priorities: a report from a EULAR ad hoc expert committee. Ann Rheum Dis 73:1442–1445
Juhl C, Christensen R, Roos EM, Zhang W, Lund H (2014) Impact of exercise type and dose on pain and disability in knee osteoarthritis: a systematic review and meta-regression analysis of randomized controlled trials. Arthritis Rheumatol 66:622–636
Nieminen HJ, Salmi A, Karppinen P, Haeggstrom E, Hacking SA (2014) The potential utility of high-intensity ultrasound to treat osteoarthritis. Osteoarthritis Cartilage 22:1784–1799
Ellman MB, Yan D, Ahmadinia K, Chen D, An HS, Im HJ (2013) Fibroblast growth factor control of cartilage homeostasis. J Cell Biochem 114:735–742
Yates LA, Norbury CJ, Gilbert RJ (2013) The long and short of microRNA. Cell 153:516–519
Le LT, Swingler TE, Clark IM (2013) Review: The role of microRNAs in osteoarthritis and chondrogenesis. Arthritis Rheum 65:1963–1974
Miyaki S, Asahara H (2012) Macro view of microRNA function in osteoarthritis. Nat Rev Rheumatol 8:543–552
Akhtar N, Makki MS, Haqqi TM (2015) MicroRNA-602 and microRNA-608 regulate sonic hedgehog expression via target sites in the coding region in human chondrocytes. Arthritis Rheumatol 67:423–434
Miyaki S, Sato T, Inoue A, Otsuki S, Ito Y, Yokoyama S, Kato Y, Takemoto F, Nakasa T, Yamashita S et al (2010) MicroRNA-140 plays dual roles in both cartilage development and homeostasis. Genes Dev 24:1173–1185
Akhtar N, Haqqi TM (2012) MicroRNA-199a* regulates the expression of cyclooxygenase-2 in human chondrocytes. Ann Rheum Dis 71:1073–1080
Luan Y, Kong L, Howell DR, Ilalov K, Fajardo M, Bai XH, Di Cesare PE, Goldring MB, Abramson SB, Liu CJ (2008) Inhibition of ADAMTS-7 and ADAMTS-12 degradation of cartilage oligomeric matrix protein by alpha-2-macroglobulin. Osteoarthritis Cartilage 16:1413–1420
Xu X, Fan Z, Kang L, Han J, Jiang C, Zheng X, Zhu Z, Jiao H, Lin J, Jiang K et al (2013) Hepatitis B virus X protein represses miRNA-148a to enhance tumorigenesis. The J Clin Invest 123:630–645
Pan X, Zhou T, Tai YH, Wang C, Zhao J, Cao Y, Chen Y, Zhang PJ, Yu M, Zhen C et al (2011) Elevated expression of CUEDC2 protein confers endocrine resistance in breast cancer. Nat Med 17:708–714
Bao JP, Chen WP, Feng J, Hu PF, Shi ZL, Wu LD (2010) Leptin plays a catabolic role on articular cartilage. Mol Biol Rep 37:3265–3272
Swingler TE, Wheeler G, Carmont V, Elliott HR, Barter MJ, Abu-Elmagd M, Donell ST, Boot-Handford RP, Hajihosseini MK, Munsterberg A et al (2012) The expression and function of microRNAs in chondrogenesis and osteoarthritis. Arthritis Rheum 64:1909–1919
Akhtar N, Rasheed Z, Ramamurthy S, Anbazhagan AN, Voss FR, Haqqi TM (2010) MicroRNA-27b regulates the expression of matrix metalloproteinase 13 in human osteoarthritis chondrocytes. Arthritis Rheum 62:1361–1371
Matsukawa T, Sakai T, Yonezawa T, Hiraiwa H, Hamada T, Nakashima M, Ono Y, Ishizuka S, Nakahara H, Lotz MK et al (2013) MicroRNA-125b regulates the expression of aggrecanase-1 (ADAMTS-4) in human osteoarthritic chondrocytes. Arthritis Res Ther 15:R28
Vonk LA, Kragten AH, Dhert WJ, Saris DB, Creemers LB (2014) Overexpression of hsa-miR-148a promotes cartilage production and inhibits cartilage degradation by osteoarthritic chondrocytes. Osteoarthritis Cartilage 22:145–153
Honeywell DR, Cabrita MA, Zhao H, Dimitroulakos J, Addison CL (2013) miR-105 inhibits prostate tumour growth by suppressing CDK6 levels. PLoS One 8:e70515
Zhou W, Fong MY, Min Y, Somlo G, Liu L, Palomares MR, Yu Y, Chow A, O’Connor ST, Chin AR et al (2014) Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell 25:501–515
Chhana A, Callon KE, Pool B, Naot D, Watson M, Gamble GD, McQueen FM, Cornish J, Dalbeth N (2011) Monosodium urate monohydrate crystals inhibit osteoblast viability and function: implications for development of bone erosion in gout. Ann Rheum Dis 70:1684–1691
Chen CG, Thuillier D, Chin EN, Alliston T (2012) Chondrocyte-intrinsic Smad3 represses Runx2-inducible matrix metalloproteinase 13 expression to maintain articular cartilage and prevent osteoarthritis. Arthritis Rheum 64:3278–3289
Shen J, Li J, Wang B, Jin H, Wang M, Zhang Y, Yang Y, Im HJ, O’Keefe R, Chen D (2013) Deletion of the transforming growth factor beta receptor type II gene in articular chondrocytes leads to a progressive osteoarthritis-like phenotype in mice. Arthritis Rheum 65:3107–3119
Huang RL, Yuan Y, Tu J, Zou GM, Li Q (2014) Opposing TNF-alpha/IL-1beta- and BMP-2-activated MAPK signaling pathways converge on Runx2 to regulate BMP-2-induced osteoblastic differentiation. Cell Death Dis 5:e1187
Sonomoto K, Yamaoka K, Oshita K, Fukuyo S, Zhang X, Nakano K, Okada Y, Tanaka Y (2012) Interleukin-1beta induces differentiation of human mesenchymal stem cells into osteoblasts via the Wnt-5a/receptor tyrosine kinase-like orphan receptor 2 pathway. Arthritis Rheum 64:3355–3363
Yoon WJ, Cho YD, Kim WJ, Bae HS, Islam R, Woo KM, Baek JH, Bae SC, Ryoo HM (2014) Prolyl isomerase Pin1-mediated conformational change and subnuclear focal accumulation of Runx2 are crucial for fibroblast growth factor 2 (FGF2)-induced osteoblast differentiation. J Biol Chem 289:8828–8838
Altorok N, Tsou PS, Coit P, Khanna D, Sawalha AH (2015) Genome-wide DNA methylation analysis in dermal fibroblasts from patients with diffuse and limited systemic sclerosis reveals common and subset-specific DNA methylation aberrancies. Ann Rheum Dis 74:1612–1620
Jeffries MA, Donica M, Baker LW, Stevenson ME, Annan AC, Humphrey MB, James JA, Sawalha AH (2014) Genome-wide DNA methylation study identifies significant epigenomic changes in osteoarthritic cartilage. Arthritis Rheumatol 66:2804–2815
Uzuki M, Sawai T, Ryan LM, Rosenthal AK, Masuda I (2014) Upregulation of ANK protein expression in joint tissue in calcium pyrophosphate dihydrate crystal deposition disease. J Rheumatol 41:65–74
Tetsunaga T, Nishida K, Furumatsu T, Naruse K, Hirohata S, Yoshida A, Saito T, Ozaki T (2011) Regulation of mechanical stress-induced MMP-13 and ADAMTS-5 expression by RUNX-2 transcriptional factor in SW1353 chondrocyte-like cells. Osteoarthritis Cartilage 19:222–232
Liu CJ (2009) The role of ADAMTS-7 and ADAMTS-12 in the pathogenesis of arthritis. Nat Clin Pract Rheumatol 5:38–45
Li W, Liu Z, Chen L, Zhou L, Yao Y (2014) MicroRNA-23b is an independent prognostic marker and suppresses ovarian cancer progression by targeting runt-related transcription factor-2. FEBS Lett 588:1608–1615
Huang Q, Jiang Z, Meng T, Yin H, Wang J, Wan W, Cheng M, Yan W, Liu T, Song D et al (2014) MiR-30a inhibits osteolysis by targeting RunX2 in giant cell tumor of bone. Biochem Biophys Res Commun 453:160–165
van der Deen M, Taipaleenmaki H, Zhang Y, Teplyuk NM, Gupta A, Cinghu S, Shogren K, Maran A, Yaszemski MJ, Ling L et al (2013) MicroRNA-34c inversely couples the biological functions of the runt-related transcription factor RUNX2 and the tumor suppressor p53 in osteosarcoma. The J Biol Chem 288:21307–21319
Viticchie G, Lena AM, Latina A, Formosa A, Gregersen LH, Lund AH, Bernardini S, Mauriello A, Miano R, Spagnoli LG et al (2011) MiR-203 controls proliferation, migration and invasive potential of prostate cancer cell lines. Cell Cycle 10:1121–1131
Kadri A, Funck-Brentano T, Lin H, Ea HK, Hannouche D, Marty C, Liote F, Geoffroy V, Cohen-Solal ME (2010) Inhibition of bone resorption blunts osteoarthritis in mice with high bone remodelling. Ann Rheum Dis 69:1533–1538
Fukuyo S, Yamaoka K, Sonomoto K, Oshita K, Okada Y, Saito K, Yoshida Y, Kanazawa T, Minami Y, Tanaka Y (2014) IL-6-accelerated calcification by induction of ROR2 in human adipose tissue-derived mesenchymal stem cells is STAT3 dependent. Rheumatology (Oxford) 53:1282–1290
Im HJ, Muddasani P, Natarajan V, Schmid TM, Block JA, Davis F, van Wijnen AJ, Loeser RF (2007) Basic fibroblast growth factor stimulates matrix metalloproteinase-13 via the molecular cross-talk between the mitogen-activated protein kinases and protein kinase Cdelta pathways in human adult articular chondrocytes. J Biol Chem 282:11110–11121
Wu B, Bi W (2015) Role of microRNA 503 in the suppression of osteosarcoma cell proliferation and migration via modulation of fibroblast growth factor 2. Mol Med Rep 12:7433–7438
Cheng Z, Ma R, Tan W, Zhang L (2014) MiR-152 suppresses the proliferation and invasion of NSCLC cells by inhibiting FGF2. Exp Mol Med 46:e112
Wang MM, Fang MX, Chen LG, Wang HQ, Liu HJ, Tang HL (2015) Differential expression of microRNA in endothelial cells incubated with serum of hypertension patients with blood-stasis syndrome. Chin J Integr Med 21:817–822
Ebrahimi-Barough S, Kouchesfehani HM, Ai J, Mahmoodinia M, Tavakol S, Massumi M (2013) An endothelial apelin-FGF link mediated by miR-424 and miR-503 is disrupted in pulmonary arterial hypertension. Nat Med 19:74–82
Rigoglou S, Papavassiliou AG (2013) The NF-kappaB signalling pathway in osteoarthritis. Int J Biochem Cell Biol 45:2580–2584
Yik JH, Hu Z, Kumari R, Christiansen BA, Haudenschild DR (2014) Cyclin-dependent kinase 9 inhibition protects cartilage from the catabolic effects of proinflammatory cytokines. Arthritis Rheumatol 66:1537–1546
Su SC, Tanimoto K, Tanne Y, Kunimatsu R, Hirose N, Mitsuyoshi T, Okamoto Y, Tanne K (2014) Celecoxib exerts protective effects on extracellular matrix metabolism of mandibular condylar chondrocytes under excessive mechanical stress. Osteoarthritis Cartilage 22:845–851
Wang JH, Shih KS, Wu YW, Wang AW, Yang CR (2013) Histone deacetylase inhibitors increase microRNA-146a expression and enhance negative regulation of interleukin-1beta signaling in osteoarthritis fibroblast-like synoviocytes. Osteoarthritis Cartilage 21:1987–1996
Rathore MG, Saumet A, Rossi JF, de Bettignies C, Tempe D, Lecellier CH, Villalba M (2012) The NF-kappaB member p65 controls glutamine metabolism through miR-23a. Int J Biochem Cell Biol 44:1448–1456
Liang M, Yao G, Yin M, Lu M, Tian H, Liu L, Lian J, Huang X, Sun F (2013) Transcriptional cooperation between p53 and NF-kappaB p65 regulates microRNA-224 transcription in mouse ovarian granulosa cells. Mol Cell Endocrinol 370:119–129
Acknowledgments
The authors would like to thank Dr. Jiying Chen and Dr. Zhigang Wang for collecting the data and Dr. Min Wei for valuable comments and sample provision. The work was financially supported through grants from the National Natural Science Foundation (81330053, 81472589, 81101387, 81371976, 31100604 and 81372161), Beijing Natural Science Foundation (7152135) and Beijing Nova Program (Z141102001814055). The General Hospital of Chinese People’s Liberation Army and Beijing Institute of Biotechnology contributed equally to this work.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Quanbo Ji, Xiaojie Xu, Yameng Xu and Zhongyi Fan contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Potential target genes of miR-105 screened. a Candidate target genes of miR-105 were found using publicly available databases (TargetScan and miRanda). b Immunoblot analysis showing the expression of the candidate target genes in chondrocytes infected with miR-105. (TIF 369 kb)
Fig. S2
FGF2/p65 promotes Runx2 and ADAMTS expression through miR-105 in human chondrocytes. Human cultured chondrocytes (passage 1) were infected with miR-105 mimics or miR-105 inhibitor and the non-target control for 48 h as described in the Methods section. Then, the infected cells were stimulated with or without FGF2. Chondrocytes were subjected to quantitative RT-PCR to determine the expression of miR-105 and the mRNA levels of Runx2, ADAMTS4. ADAMTS5, ADAMTS7 and ADAMTS12. Values are the mean of at least three independent experiments performed in triplicate ± standard deviation. * P < 0.05; ** P < 0.01. (TIF 307 kb)
Fig. S3
Knockdown of Runx2 in chondrocytes results in decreased pro-catabolic responses. a Human cultured chondrocytes (passage 1) were infected with Runx2, Runx2 shRNA or a non-target control for 48 h as described in the Methods section. ADAMTS7, ADAMTS12, ADAMTS4 or ADAMTS5 expression was examined by western blot. b Quantitative RT-PCR was performed to evaluate the mRNA levels of ADAMTS4, ADAMTS5, ADAMTS7 and ADAMTS12. Values are the mean of at least three independent experiments performed in triplicate ± standard deviation. * P < 0.05; ** P < 0.01. (TIF 397 kb)
Fig. S4
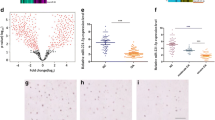
Expression of miR-105, Runx2, ADAMTS7 and ADAMTS12 in patients with CA. Runx2, ADAMTS7 and ADAMTS12 scores and miR-105 expression in CA and normal tissue were plotted and compared. (TIF 290 kb)
Rights and permissions
About this article
Cite this article
Ji, Q., Xu, X., Xu, Y. et al. miR-105/Runx2 axis mediates FGF2-induced ADAMTS expression in osteoarthritis cartilage. J Mol Med 94, 681–694 (2016). https://doi.org/10.1007/s00109-016-1380-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-016-1380-9