Abstract
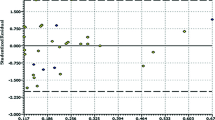
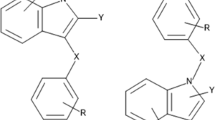
Indolylarylsulfones (IASs) have received considerable interest during the last decades due to high potency against HIV-1 as non-nucleoside reverse transcriptase inhibitors. In present work, quantitative structure–activity relationship (QSAR) and molecular docking analyses were performed to model the anti-HIV-1 activity of 36 IASs. 2D and 3D-descriptors, genetic algorithm, internal and external validations were used to develop statistically robust four-parametric QSAR models. The best QSAR model is with R 2tr = 0.8608. The QSAR analysis reveals that the activity of IASs depends on the presence of electronegative and heavy atoms at the internal atmosphere of the compounds. The docking analysis reveals that lipophilic and H-bonding interactions are the prominent interactions among IASs and the receptor. The QSAR analysis proved to be a useful tool in the prediction of anti-HIV-1 activity of congeneric compounds and some important insights were also found that will be useful to guide for the synthesis of new IASs with improved activity.






Similar content being viewed by others
Abbreviations
- QSAR:
-
Quantitative structure–activity relationship
- NNRTIs:
-
Non-nucleoside reverse transcriptase inhibitors
- IASs:
-
Indolylarylsulfones
- WHO:
-
World Health Organization
- AIDS:
-
Acquired Immunodeficiency Syndrome
- HAART:
-
Highly active antiretroviral therapy
- RT:
-
Reverse transcriptase
- GA:
-
Genetic algorithm
References
Chirico N, Gramatica P (2011) Real external predictivity of QSAR models: how to evaluate it? comparison of different validation criteria and proposal of using the concordance correlation coefficient. J Chem Inf Model 51(9):2320–2335
Chirico N, Gramatica P (2012) Real external predictivity of QSAR models. Part 2. New intercomparable thresholds for different validation criteria and the need for scatter plot inspection. J Chem Inf Model 52(8):2044–2058
Doweyko AM (2008) QSAR: dead or alive? J Comput Aided Mol Des 22(2):81–89
Golbraikh A, Tropsha A (2002) Beware of q2! J Mol Graph Model 20(4):269–276
Gramatica P (2013) On the development and validation of QSAR models. Methods Mol Biol 930:499–526
Gramatica P, Papa E (2007) Screening and ranking of POPs for global half-life: QSAR approaches for prioritization based on molecular structure. Environ Sci Technol 41(8):2833–2839
Gramatica P, Pilutti P, Papa E (2007) Approaches for externally validated QSAR modelling of nitrated polycyclic aromatic hydrocarbon mutagenicity. SAR QSAR Environ Res 18(1–2):169–178
Kim JH, Gramatica P, Kim MG, Kim D, Tratnyek PG (2007) QSAR modelling of water quality indices of alkylphenol pollutants. SAR QSAR Environ Res 18(7–8):729–743
Kovarich S, Papa E, Li J, Gramatica P (2012) QSAR classification models for the screening of the endocrine-disrupting activity of perfluorinated compounds. SAR QSAR Environ Res 23(3–4):207–220
La Regina G, Coluccia A, Brancale A, Piscitelli F, Gatti V, Maga G, Samuele A, Pannecouque C, Schols D, Balzarini J, Novellino E, Silvestri R (2011) Indolylarylsulfones as HIV-1 non-nucleoside reverse transcriptase inhibitors: new cyclic substituents at indole-2-carboxamide. J Med Chem 54(6):1587–1598
Liu H, Gramatica P (2007) QSAR study of selective ligands for the thyroid hormone receptor beta. Bioorg Med Chem 15(15):5251–5261
Liu H, Papa E, Gramatica P (2008) Evaluation and QSAR modeling on multiple endpoints of estrogen activity based on different bioassays. Chemosphere 70(10):1889–1897
Mahajan DT, Masand VH, Patil KN, Ben Hadda T, Jawarkar RD, Thakur SD, Rastija V (2012) CoMSIA and POM analyses of anti-malarial activity of synthetic prodiginines. Bioorg Med Chem Lett 22(14):4827–4835
Mahajan DT, Masand VH, Patil KN, Hadda TB, Rastija V (2013) Integrating GUSAR and QSAR analyses for antimalarial activity of synthetic prodiginines against multi drug resistant strain. Med Chem Res 22:2284–2292
Martin TM, Harten P, Young DM, Muratov EN, Golbraikh A, Zhu H, Tropsha A (2012) Does rational selection of training and test sets improve the outcome of QSAR modeling? J Chem Inf Model 52(10):2570–2578
Masand VH, Jawarkar RD, Patil KN, Nazerruddin GM, Bajaj SO (2010) Correlation potential of Wiener index and molecular refractivity vis-a`-vis antimalarial activity of xanthone derivatives. Org Chem 6(1):30–38
Masand VH, Jawarkar RD, Mahajan DT, Hadda TB, Sheikh J, Patil KN (2012) QSAR and CoMFA studies of biphenyl analogs of the anti-tuberculosis drug (6S)-2-nitro-6-{[4-(trifluoromethoxy) benzyl]oxy}-6,7-dihydro-5H-imidazo[2,1-b][1,3]oxazine(PA-824). Med Chem Res 21:2624–2629
Masand VH, Mahajan DT, Patil KN, Hadda TB, Youssoufi MH, Jawarkar RD, Shibi IG (2013) Optimization of antimalarial activity of synthetic prodiginines: QSAR, GUSAR, and CoMFA analyses. Chem Biol Drug Des 81(4):527–536
Sushko I, Novotarskyi S, Korner R, Pandey AK, Cherkasov A, Li J, Gramatica P, Hansen K, Schroeter T, Muller KR, Xi L, Liu H, Yao X, Oberg T, Hormozdiari F, Dao P, Sahinalp C, Todeschini R, Polishchuk P, Artemenko A, Kuz’min V, Martin TM, Young DM, Fourches D, Muratov E, Tropsha A, Baskin I, Horvath D, Marcou G, Muller C, Varnek A, Prokopenko VV, Tetko IV (2010) Applicability domains for classification problems: benchmarking of distance to models for Ames mutagenicity set. J Chem Inf Model 50(12):2094–2111
Tropsha A (2010) Best practices for QSAR model development, validation, and exploitation. Mol Inf 29:476–488
WHO Report (2011) http://www.who.int/. Accessed 23 Nov 2012
Acknowledgments
We are thankful to Dr. K. N. Patil, Amravati and Dr. Ashraf Ali, Singapore for useful discussions. Sincere thanks to Dr. F. C. Raghuwanshi for encouragement and facilities.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Masand, V.H., Mahajan, D.T., Hadda, T.B. et al. Molecular docking and quantitative structure–activity relationship (QSAR) analyses of indolylarylsulfones as HIV-1 non-nucleoside reverse transcriptase inhibitors. Med Chem Res 23, 417–425 (2014). https://doi.org/10.1007/s00044-013-0647-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-013-0647-8