Abstract
The COVID-19 pandemic poses a major burden on healthcare and economic systems across the globe. Even though a majority of the population develops only minor symptoms upon SARS-CoV-2 infection, a significant number are hospitalized at intensive care units (ICU) requiring critical care. While insights into the early stages of the disease are rapidly expanding, the dynamic immunological processes occurring in critically ill patients throughout their recovery at ICU are far less understood. Here, we have analysed whole blood samples serially collected from 40 surviving COVID-19 patients throughout their recovery in ICU using high-dimensional cytometry by time-of-flight (CyTOF) and cytokine multiplexing. Based on the neutrophil-to-lymphocyte ratio (NLR), we defined four sequential immunotypes during recovery that correlated to various clinical parameters, including the level of respiratory support at concomitant sampling times. We identified classical monocytes as the first immune cell type to recover by restoration of HLA-DR-positivity and the reduction of immunosuppressive CD163 + monocytes, followed by the recovery of CD8 + and CD4 + T cell and non-classical monocyte populations. The identified immunotypes also correlated to aberrant cytokine and acute-phase reactant levels. Finally, integrative analysis of cytokines and immune cell profiles showed a shift from an initially dysregulated immune response to a more coordinated immunogenic interplay, highlighting the importance of longitudinal sampling to understand the pathophysiology underlying recovery from severe COVID-19.
Similar content being viewed by others
Introduction
The SARS-CoV-2 coronavirus outbreak has infected more than 93 million people worldwide causing more than 2 million deaths and was officially declared a pandemic by the World Health Organization (WHO) [1]. Due to the lack of immunity to SARS-CoV-2, and while awaiting a comprehensive vaccination strategy, the unpredictable clinical outcome of COronaVirus-Induced Disease 2019 (COVID-19) has created significant strain on medical systems [2,3,4,5]. Over the past year, major research efforts have tried to unravel the pathobiology underlying the disease course [6, 7]. As such, it has been shown that COVID-19 leads to the combination of pathologically elevated levels of pro-inflammatory cytokines, coagulopathy and a dysregulated immune response [6, 8, 9]. Deep immune-profiling in severe COVID-19 patients revealed excessive amounts of dysfunctional neutrophils, decreased levels of lymphocytes and low levels of antigen-presenting receptors on monocytes and dendritic cells, hindering efficient adaptive immune responses [8, 10,11,12]. In contrast to asymptomatic or mild/moderate symptomatic COVID-19 patients, critically ill patients typically show a biphasic disease course with an early viral stage before ICU admission, followed by a hypoxic phase characterized by a cytokine storm and acute respiratory distress requiring (invasive) respiratory support, antivirals and immune modulation to prevent multiple organ failure and death.
Considering the potentially severe and unpredictable impact of a SARS-CoV-2 infection, research has also focused on predicting the disease course of COVID-19. Using a variety of multiplexing technologies, longitudinal blood sampling has revealed a dichotomous profile in cytokine expression levels that could differentiate patients prone to develop mild/moderate versus severe disease within the first-two weeks following SARS-CoV-2 infection [13,14,15]. In addition, severely diverging levels of immune cell populations, typically reported by aberrant neutrophil-over-lymphocyte ratios (NLR) and higher levels of immature neutrophil frequencies were also indicative of a more severe disease course [16,17,18,19,20,21,22]. Based on these insights, various interventional trials were initiated, from which the use of corticosteroids showed promising results [23] although the optimal timing of these interventions remains to be determined and side effects (i.e. secondary infections) may complicate their use. Therefore, until the population gets vaccinated, patients exhibiting severe pathology will continue to require extensive medical care and need to be admitted to ICU, with the associated burden on the healthcare system.
In contrast to the early stages of the disease, the biology from ICU admission to the recovery phase remains far less understood. A systems-biology study with a longitudinal follow-up of 10 ICU patients suggested involvement of eosinophils early in the disease [24]. Various immune cell populations were also found to be increased in relative proportions during recovery, while excessive cytokine levels returned to normal at the time of discharge from ICU [24, 25]. Restoration of T-cell functionality, including the induction of effector-memory cells, has also been described during the recovery phase of COVID-19 [26]. However, the exact sequence of changes occurring during the restitution of the peripheral immune system after severe infection remains elusive.
Within the transdisciplinary clinical COntAGIouS trial (COvid-19 Advanced Genetic and Immunologic Sampling, NCT04327570), we have analysed serially collected whole blood and serum samples of critically ill COVID-19 patients that survived an ICU stay at the University Hospitals Leuven, Belgium. By using high-dimensional analysis technologies, we identified four immunotypes (i.e. blood cell composition profiles) based on the NLR, defining a common biological trajectory during patient recovery which correlated well to various clinical parameters including the required level of respiratory support at concomitant sampling times. We identified classical monocytes as the first cell type to recover, followed by CD8 + and CD4 + T cell and non-classical monocyte populations, while the NLR-defined groups also correlated to cytokine and acute-phase reactant (APR) levels. Finally, an integrated analysis of cytokines and immune cell profiles allowed us to monitor recovery from this dysregulated immune response, highlighting the importance of sampling at multiple timepoints to better understand the pathophysiology underlying recovery from severe COVID-19.
Methods
Patient cohort and sampling
Forty COVID-19 patients that had been admitted to the ICU were enrolled in the ‘COntAGIouS’ trial (NCT04327570). Thirty-eight patients were confirmed positive for SARS-CoV-2 infection by nasopharyngeal swab and/or by bronchoalveolar lavage (BAL) sampling and subsequent PCR quantification. Two patients were confirmed positive for SARS-CoV-2 via CT scan. In line with previous reports, CT scans are used to classify “probable positive” cases in the absence of a positive RT-PCR test based on the “ground-glass” characteristics which are a known hallmark of early COVID-19 infection [27]. Routine clinical laboratory tests at the time of sampling were performed as outlined within the ‘COntAGIouS’ study design [12]. The patients enrolled in the study were sampled at several clinically relevant time points: upon admission to the ICU (0–2 days post admission), at 6 ± 2 days post admission, and eventually at ICU discharge. Of note, if the medical condition of the patient was not improving, a bronchoscopy procedure was performed during which whole blood samples were again collected for concomitant immune-profiling.
The healthy control group consisted of volunteers (recruited from UZ Hospital staff) with no prior diagnosis or recent symptoms compatible with COVID-19. This group was not tested by nasopharyngeal swab, but antibody assessment of plasma samples revealed an absence of COVID-19 IgG antibodies.
Sample processing and staining procedure for CyTOF
Whole blood (WB) samples were drawn into sterile anticoagulant lithium heparin blood collection tubes and processed for mass cytometry staining within 2–4 h of collection. WB was stained with the Maxpar® Direct™ Immune Profiling Assay (DIPA) kit from Fluidigm© according to manufacturer instructions by following the workflow outlined for whole blood staining. Additional antibodies not included in the Maxpar DIPA kit were titrated on WB samples after RBC lysis from healthy donors to determine the optimal staining index. The antibodies are outlined in Suppl. Table 1. The additional markers were added after the red blood cell-removal step. The last step of the protocol is 193-Ir intercalation, which was performed overnight at 4 °C. The samples that could not be acquired on the instrument the next day were immediately cryopreserved in Maxpar Fix/Perm buffer + 193-Ir at − 80 °C. This cryopreservation technique was validated in triplicate by dividing aliquots of donor samples stained on the same day and comparing cell viability and immune profiles between the fresh and cryopreserved samples. Batch effects were evaluated by running a reference sample derived from an aliquot of the same healthy donor’s blood over the period of the study.
Data acquisition
Cells stained for mass cytometry were acquired the day after the staining procedure or within a week of cryopreservation. For CyTOF acquisition, the samples were pelleted in Maxpar Cell Acquisition Buffer (CAS) on the day of acquisition and transferred to the KU Leuven Flow and Mass Cytometry Facility. Prior to acquisition, 1 million cells/ml were resuspended in CAS containing EQ beads (1:10) and filtered through 35 µm cell strainer cap tubes. The samples were acquired at a rate of 250–300 events per second on a Helios® Mass Cytometer (Fluidigm). CyTOF software version 6.7.1016 and the Maxpar Direct Immune Profiling Assay.tem template were used to acquire and normalize data from the stained samples.
Chemokine and cytokine assays
Chemokine and cytokine levels in plasma were assessed according to manufacturer instructions (Meso Scale Discovery) using the V-plex human cytokine 30-plex kit, complemented with Human IL-1RA (V-plex) and human IL-18 (U-plex) kits.
Data analysis
To analyse the CyTOF data, we used the FlowSOM models built for [12]. A general overview of our pipeline is shown in Supp. Figure 1, Part A. To generate these models, normalized.fcs files were first transferred to the Maxpar Pathsetter™ software (version 2.0.45), which performs a standardized, automatic and unsupervised quality check (bead removal and high-quality singlet selection) and gating of live single cells. Samples with fewer than 50,000 cells were discarded, as were four samples which showed-up as outliers in a PCA analysis of the 25, 50 and 75% quantiles of the marker values. The FlowSOM analysis was subsequently performed in R, using 123 cleaned.fcs files, including eight healthy controls and longitudinal samples of 40 COVID-19 patients, throughout various stages of disease. Samples were first pre-processed: margin events were filtered out, data was transformed with an arcsinh transformation with cofactor 5 and the PeacoQC algorithm (manuscript in preparation, tool available at https://github.com/saeyslab/peacoQC) was applied to remove any unstable signal regions during the measurement. On this cleaned data a first FlowSOM model [28] was trained, using a random selection of cells for all samples, resulting in 3,000,093 cells to train on. The clustering made use of 11 markers (CD3, CD4, CD8a, CD11c, CD14, CD19, CD20, CD45, CD66b, TCRƴδ and NCAM), mapped the data onto a 10 by 10 SOM grid and resulted in 30 meta-clusters. 22 meta-clusters were selected as having CD66b values lower than 2 or CD45 values higher than 4, corresponding to non-granulocytes, while eight meta-clusters were labelled as granulocytes. The full files were mapped onto this model, and for each of them new.fcs files were generated corresponding to the two subsets of cells. These were then used to build two separate FlowSOM models, including either the non-granulocyte or the granulocyte cells, again using only a subset of 2,949,946 (granulocyte: 3,000,093) cells for training mapped onto a 10-by-10 SOM grid, this time using 33 markers (CD3, CD4, CD8a, CD11c, CD14, CD16, CD19, CD20, CD27, CD28, CD38, CD45, CD45RA, CD45RO, CD57, CD66b, CD69, CD294, CD161, CD163, CCR4, CCR6, CCR7, CXCR3, CXCR5, HLA-DR, IgD, IL-2Rα, IL-3R, IL-7Rα, NCAM, NKG2A and TCRƴδ). To be able to identify small populations, no meta-clustering was applied on these second models, and the 100 clusters were manually annotated by three independent experts according to their mean metal intensity (MMI) values. In the non-granulocyte model, six clusters were manually identified as still being mixtures of different cell types, and split into two or three clusters, resulting in 108 non-granulocyte clusters and 100 granulocyte clusters.
For this study, all samples were again fully mapped onto these models to identify their immune profiles. During this process, we noticed some files had a sub-optimal granulocyte/non-granulocyte split. Therefore, we redefined this split using the file-specific MMI values of the metaclusters, selecting all metaclusters with CD66b values lower than 3 and CD45 values higher than 3 as non-granulocytes. For one file, three metaclusters were still incorrectly assigned when inspecting the density distribution of the cells, so for this file the CD45 boundary was adapted to 4. Once all cells were assigned to the 208 clusters from the existing models, we used a manual annotation of these clusters to aggregate them in 24 cell populations, describing the immune profile of each sample by the percentage of live cells each of these populations represented. In addition, the total lymphocyte percentage (i.e. the sum of all populations which were annotated as lymphocyte subsets) was added, bringing the total to 25 parameters per sample. Importantly, the values determined by CyTOF corresponded well to the general blood counts routinely assessed by the haematology analysers available at the clinic, as can be seen by the representation of the data as % of cells vs number of cells per volume of blood (Supp. Figure 2). An overview of the mapping of these 24 cell populations on a tSNE representation of the data is shown in Supplemental Figure 3-Part A. The samples were then grouped in four equal-sized groups depending on their NLR: 31 samples with a ratio higher than 7.1 (R4), 30 samples with a ratio between 7.1 and 4.8 (R3), 31 samples with a ratio between 4.8 and 2.7 (R2) and 31 samples with a ratio lower than 2.7 (R1). The four profiles were ordered by decreasing neutrophil/lymphocyte ratio, and are throughout the figures in this paper coloured as black, green, blue and cyan, respectively. We chose to split the data in four equal-sized groups, as clustering approaches are known to perform suboptimally on data sets in which gradual/continuous changes are expected. As described throughout the manuscript dividing the samples in the identified groups resulted in the discovery of relevant patterns.
The differences among these four groups, which included both patient and control samples, were then further characterized. To compute statistics, we made use of linear mixed models, and for the cytokines and APR values we used mixed models with random effect for the patient, to correct for the fact that patients can have multiple longitudinal samples, spread over one or more groups. Correlations were calculated using the Spearman Rank test. All p values were taken together to be corrected for multiple testing with the Benjamini–Hochberg (FDR) method. Curve-fitting analysis was done using the drc package in R from which the inflection points and the corresponding p values were collected. In addition, we also used the built-in immuno profiling tools of the Maxpar Pathsetter™ software to analyse the overall immune cell profile in a standardized and automated way. The Pathsetter-based blood profiles were grouped into 12 clusters using k-means clustering, and manually further grouped into four final profiles. These profiles strongly corresponded with the NLR results and led to similar biological conclusions, thus giving additional validation in our grouping strategy.
The code used to generate the results is available at https://github.com/saeyslab/CYTOF_covid19_study. The raw.fcs files have been uploaded and annotated via Flow Repository and the data is available at Flow Repository ID: FR-FCM-Z34U.
To create the correlation matrices, we applied the Morpheus software (https://software.broadinstitute.org/morpheus) and used Pearson correlation metric for clustering. For clustering, only those values for COVID-19 patients that were matched across both serum cytokine/chemokine screening as well as mass cytometry were considered. Unmatched patients were excluded.
Results
Mapping the disease course of COVID-19 survivors at ICU
To unravel the sequential recovery of the immune system throughout the course of a severe COVID-19 infection, we have analysed a cohort of 40 patients that survived a stay at the intensive care unit (ICU) at the University Hospitals Leuven, Belgium. These patients were enrolled in the COntAGIouS trial (NCT04327570) through which whole blood samples were serially collected during their ICU stay (see methods and Table 1 for patient demographics and characteristics; a set of healthy controls was included for reference).
For each patient, a detailed timeline of their ICU stay was generated to display the most clinically relevant events, including onset of symptoms, hospital admission, ICU admission, start/stop of therapeutic interventions and finally discharge from ICU and hospital. If applicable, discharge from a specialized revalidation centre where the patient continued his/her recovery after hospital discharge is also included. Occasionally, blood was drawn at additional time points (such as during bronchoscopy) and is also indicated on the timeline. From this graphical representation of individual time points (Fig. 1a), a large range in the duration of ICU stay (from 2 to 72 days) was evident, indicating a highly variable recovery rate across patients. Of the 40 enrolled patients, only nine were women, which corresponds to previous observations of males experiencing increased severity and hospitalization rates once infected with SARS-CoV-2 [29,30,31] (Fig. 1b). The age range and overall time spent at ICU was, however, comparable between male and female patients (Fig. 1a, b and Table 1). Within this critically ill patient cohort, we did not find a correlation between BMI and the overall duration of ICU stay (r = − 0.15, p = 0.35) despite > 40% of the patients having a BMI above 30 (Table 1, Fig. 1b) [32].
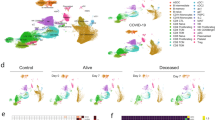
Detailed timeline of the COVID-19 patient cohort highlighting clinical milestones. a Line plot on which clinical milestones are highlighted according to their relative time (days) from each patient’s admission to ICU. Patients are ordered (top to bottom) according to their overall length of stay in ICU. b Dot plot indicating the gender and BMI of each included patient. c, d Line plot indicating the level of required respiratory support and corresponding WHO scores at concomitant sampling times (d) as indicated in panel a. Panel c highlights the maximal level of respiratory support each patient required at concomitant sampling times (c). e Mapping of the defined NLR groups (1–4) on top of the indicated timeline as defined in panel a. f Indication of the treatments received by each of the patients during their stay at ICU. The colour and symbol code are indicated in each separate panel. CS corticosteroid, a-IL1 anti-IL-1, a-IL6 anti-IL6, HFNC high flow nasal cannula, NIV non-invasive ventilation, IMV invasive mechanical ventilation, ECMO extracorporeal membrane oxygenation, iNO inhaled nitric oxide, BMI body mass index
This timeline was further expanded to include the level of respiratory support that was required at concomitant sampling times (Fig. 1c, d). Here, respiratory support levels were classified from 0 to 5 where level 0 indicates no support, while level 4 and 5 indicates the need for mechanical ventilation [with level 5 indicating patients that were ventilated in prone position and/or requiring inhalation NO therapy or extracorporeal membrane oxygenation (ECMO)] (Table 1; the corresponding WHO scores are indicated in Fig. 1d). Subsequent correlation analysis showed that the maximal level of respiratory support correlated well to the overall duration of ICU stay (r = 0.67, p = 7e–17, Fig. 1c, d, supp. Figure 4).
Identifying peripheral blood profiles using repeated sampling
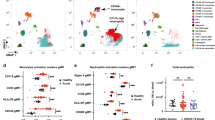
Next, we performed deep immune-profiling on collected blood samples using high dimensional cytometry by time-of-flight (CyTOF) analysis. From this data, 33 surface markers were used to map changes in composition and phenotype of white blood cells during ICU stay. All cells were clustered using FlowSOM, an unsupervised gating method, and these clusters were manually annotated to identify 25 immune cell subtypes to reconstruct potential dynamic changes. However, given the variation in duration of hospitalization at the ICU in this patient cohort, mapping immune profiles on an absolute time scale did not produce coherent results. Alternatively, we ranked all samples, both from COVID-19 patients and healthy controls, based on the neutrophil-to-lymphocyte ratio (NLR). To allow comparison between and identify evolutions across different NLR levels, we defined four equal-sized groups (Fig. 2a, b) across all collected samples, regardless of when a sample was taken. As such, we defined four “NLR” stages (R1–4) ranging from severe lymphopenia and neutrophilia (profile R4, which showed the highest NLR values) to immune profiles resembling those of healthy controls, defined by the lowest NLR values (R1) (Fig. 2a, b and Suppl. Figure 5). In addition, the highest NLR group (R4) had increased levels of the acute phase reactants C-reactive protein (CRP) and ferritin (Fig. 2c, d), biomarkers of inflammation that are commonly monitored in ICU patients and define a critical state.
Longitudinal immunoprofiling of peripheral blood profiles and integration with clinical features. a Overview of immune cell population percentages, grouped by NLR, where R4 indicates the most severe neutrophilia/lymphopenia and R1 shows similar values to healthy controls (HC). (n: R4 = 31, R3 = 30, R2 = 31, R1 = 31) b Neutrophil and lymphocyte percentages, ordered by decreasing NLR. (n = 123) c, d Levels of acute phase reactants, C-reactive protein and ferritin, across all ratio groups. (n: R4 = 30, R3 = 29, R2 = 31, R1 = 22) e Distribution of respiratory support levels (n: R4 = 30, R3 = 29, R2 = 31, R1 = 22) and f SOFA score of the first 24 h upon admission to ICU over the NLR groups. (n: R4 = 12, R3 = 4, R2 = 4, R1 = 4) g Longitudinal sampling time points of patients from admission into Ward and ICU, mid-stay and discharge as grouped by NLR. (n: R4 = 31, R3 = 30, R2 = 31, R1 = 31) h Days of revalidation upon discharge from ICU as grouped by NLR. (n: R4 = 3, R3 = 11, R2 = 11, R1 = 14) a–h Colour indicates NLR group: black: R4, green: R3, blue: R2, cyan R1. HC are indicated as black open circles. ***p ≤ 0.001, **p ≤ 0.01, *p ≤ 0.05
Integrating longitudinal immune-profiling with clinical features
Next, we assessed how the identified NLR groups corresponded to a variety of clinical parameters. As indicated above, the level of respiratory support was measured at each sampling time point and correlation analysis of this feature with the four NLR profiles revealed that patients exhibiting R4 required increased respiratory support at that moment compared to lower level NLRs (Fig. 2e). Further correlation analysis also showed that the overall clinical status of each patient upon admission to ICU, as determined by calculating the Sequential Organ Failure Assessment Score (SOFA) [33, 34], was significantly correlated to the NLR groups (Fig. 2f; r = − 0.23, p = 0.03). We did not find any correlations of age or BMI with the NLR groups (Suppl. Figure 6A-D).
Using these groups, we reconstructed a detailed sequence of events for each individual patient and investigated how these evolved along their ICU stay (Figs. 1e, 2g, Suppl. Figure 7). For 35 out of 40 patients, we observed a recovery (overall downwards shift of the NLR score) or stabilisation of their immune profile, along their stay at ICU (Suppl. Figure 7), with an enrichment of R1 towards ICU discharge (Fig. 2g). Strikingly, the NLR values at ICU discharge ranged between 1 and 4, suggesting that the physical condition of these patients remained heterogeneous at that moment. In line with this observation, we found that patients exhibiting a higher NLR value upon discharge from ICU required significantly longer revalidation (either at the regular hospital ward and/or a specialized revalidation center) following their stay at ICU (Fig. 2h, r = − 0.4, p = 0.02). In addition, Charlson comorbidity indices [35] were low for all patients (Table 1), indicating that the majority of patients admitted to ICU were in good general health before their infection with SARS-CoV-2. We also did not observe correlations of the Charlson comorbidity indices to either the NLRs or the required level of respiratory support (not shown).
Finally, we tracked alterations in patient immune profiles who were treated with diverse immunomodulatory regimens. Commonly, the glucocorticoid methylprednisolone (MP) was given as a monotherapy (n = 19). In several cases, MP was prescribed in combination with either anti-IL-1 (anakinra; n = 3) or anti-IL-6 (tocilizumab; n = 5). One patient received anti-IL-6 exclusively while the remaining patients in our cohort (n = 12) relied on supportive care alone (Table 1; Fig. 1f). Unsurprisingly, our data analysis confirmed that patients who underwent steroid treatment had a greater tendency towards a longer ICU stay. Correlation analysis of the NLR profiles with MP treatment revealed that patients exhibited a higher NLR upon ICU discharge compared to patients that did not receive MP (Suppl. Figure 6E). Finally, while the cohort of patients that received anti-IL-1 and anti-IL-6 was small, the reconstitutional trajectory of their immune systems followed a similar pattern to those patients receiving standard-of-care treatment without immunomodulatory drugs and/or MP.
Reconstructing the cellular recovery of critically ill COVID-19 patients
Considering that the NLR levels correlated well with the clinical condition of the patients (see above), we next investigated the recovery of more specific immune cell populations to eventually define a sequence of events. To do so, we used a dual approach. First, we performed curve-fitting along the trajectory of the four NLR profiles, in which we determined the inflection points of each identified immune cell population to estimate the moment recovery would begin (Fig. 3a, b, Suppl. Table 2). The data from this approach were then, in a second phase, combined with the statistical comparison of the four NLR groups (Fig. 2a, suppl. Table 3; see methods for statistical procedure) to define the eventual order by which specific cell populations were recovering. We found that, on average across this cohort of patients, classical monocytes were the first immune cell population to recover followed by naive CD8 + T cells, naive and Th2-polarized CD4 + T cells. As indicated by our analysis, effector and memory T cell populations were only restored at later stages, including the anti-viral Th1-polarized CD4 + T cells [36]. Following this first phase, the non-classical monocytes began recovering. From the other professional antigen-presenting cells, plasmacytoid dendritic cells (pDC) began recovering immediately after the early T cell response while myeloid DCs (mDC) only recovered towards R2/R1, similar to overall B and NK cell populations. This suggests that monocytes, as implied by other studies, could become important targets in both understanding COVID-19 disease progression as well as improving recovery at the early phase of ICU admission [37].
Overview of cell population abundances in peripheral blood profiles. a Abundance of selected populations shown, fitted with a dose response curve using a three-parameter logistic model (red line). Individual samples coloured by NLR, black open circles for healthy controls (HC). (n: R4 = 31, R3 = 30, R2 = 31, R1 = 31) b Median abundance per NLR group. Values are scaled per row, dark green indicating in which group the cell population is the most abundant. CD4_Tregs had a median of 0 in all profiles, indicated in grey. c Overview of all FlowSOM clusters labelled as monocytes. On the left, the relative abundance per NLR group (similar to b); on the right, the median marker expression for those clusters
Functional recovery of monocytes
Antigen-presentation by monocytes, pDCs and/or mDCs plays a vital role in the initiation of an efficient adaptive immune response to viral infections [38,39,40,41,42]. As previously shown, declined levels of HLA-DR expression and hence reduced antigen presentation seems to be an early hallmark of a trajectory towards severe COVID-19 compared to mild/moderate disease [10, 12]. Considering that monocytes are among the first immune cell types to recover, we further investigated the functionality of these cells across the four NLR profiles during recovery. As such, we observed a regain in both the numbers and the antigen-presenting capabilities of the monocytes towards discharge (Fig. 3c). The first subset to be re-established were the classical HLA-DR + monocytes (Cluster (Cl) 3, Cl33 and Cl12; Fig. 3c), while it was only in later stages of disease progression that the HLA-DR + mDCs were restored (see Cl11, Fig. 3c; see also statistics in Fig. 2a).
The early phase of severe COVID19 is characterized by a functional shift towards a more immunosuppressive spectrum of monocytes, as seen by a downregulation of HLA-DR and an enrichment of CD163 + monocytes in R4 (Fig. 3) [10, 12]. However, longitudinal follow-up shows that this shift is reversible, as seen by the recovery of HLA-DR expression (suppl. Figure 8) and a relative reduction in the amount of immunosuppressive CD163 + monocytes (see Cl53 and Cl32; Fig. 3c), leading to a restoration of the antigen-presenting phenotype of these cells.
Longitudinal cytokines profiling
In addition to immunophenotyping, we also performed multiplexed analysis of soluble serum proteins, which included 32 pro-inflammatory cytokines and chemokines in serum samples prepared at concomitant sampling times. The levels of these analytes were subsequently compared to the NLR profiles. In line with previous observations [14, 15, 43, 44], the initial R4 stage of patients arriving at ICU was characterized by an increase in IFN-ƴ, TNF-α, IL-2, IL-6, IL-7, IL-10, IL-15, IP-10, MCP-1, MIP-1α, MIP-1β levels and a reduction in TARC, MDC and IL1-α, suggestive of a pro-inflammatory cytokine signature, often referred to as a “cytokine storm” (Fig. 4a). This signature was steadily reversed as patients attained a normal R1 stage. Similarly, CRP and ferritin were also significantly higher in R4 and gradually decreased as patients reached R1, suggestive of a return to baseline following an acute phase induction of the immune system (Fig. 2c, d). Clustering analysis to uncover patterns in cytokine profiles across the various NLR groups further confirmed that pro-inflammatory cytokine levels collectively reduced along the duration of an ICU stay (Fig. 4b). On the other hand, MDC and TARC, two constitutive chemokines designated as CCL17 and CCL22 that are regulated at a post-translational level, increased back to normal levels [45].
Cytokine and chemokine profiling according to the peripheral blood profiles. Serum levels of 32 pro-inflammatory soluble proteins were determined by multiplexed ELISA concurrent with the whole blood sampling times. a Box plots illustrate the correlation of the level of the significantly different cytokine/chemokine with the outlined R1–4 (log10 scale, filled circle presents a COVID-19 patient, black empty circle illustrates a healthy control patient). (n: R4 = 29, R3 = 25, R2 = 28, R1 = 25) b Heatmap generated by clustering analysis of the cytokine expression signatures across R1–4. ***p ≤ 0.001, **p ≤ 0.01, *p ≤ 0.05
Even though the levels of cytokines within the R1 profile remained comparable between COVID-19 patients and healthy individuals, we observed altered expression levels of TNF-α, IL-18, IL1-α, MCP-1 and MIP-1α, between the two study groups (Fig. 4a; healthy controls are indicated as hollow black circles in the figure). It remains to be investigated how long it would take for these levels to normalise; however, despite their aberrant profiles, patients were still able to leave the ICU for their recovery.
Integrative mapping of the immune response during recovery
Considering the important interplay between immune cells, cytokines and chemokines during severe COVID-19, we also performed an integrative, similarity matrix-based correlative statistical modelling analyses (Fig. 5) to uncover associative patterns along the duration of a patient’s recovery at ICU. As recently determined, critically ill COVID-19 patients are characterized by a strong dysregulation of the immune reaction against the SARS-CoV-2 virus, whereby normally highly concerted interplays of cyto/chemokines and specific immune cell populations become disentangled [8]. In line with above observations, our similarity matrix analyses also revealed an intense correlation between various immune cells and cyto/chemokines (a putative indication of ongoing immunological interactions) in the R4 critically-ill COVID-19 patients (Fig. 5a), such that these correlations progressively “normalized” or decreased in terms of number of intense clusters when traversing from R4 to R1 (Fig. 5a–d), thereby indicating that apart from above quantitative shifts, there was also a qualitative shift in possible immune-interactions when going from critically-ill to recovering COVID-19 patients.
Integration analysis of the blood profiles and cytokine levels. Pearson correlation-driven similarity/correlation matrix analysis of cytokines/chemokines and mass cytometry data across the different COVID-19 patient’s NLR subgroups i.e., R4 (a), R3 (b), R2 (c) and R1 (d). This matrix is a statistical modelling representation wherein statistically stable correlations between variables across different patients facilitates their co-clustering as indicated by the dendrograms. The diagonal correlation value is 1, which denotes the highest possible statistically-significant correlation value that is “centering” our correlation matrices
Furthermore, the immunological characteristics of these clusters within the four blood profiles were of particular interest considering the importance of both neutrophils and lymphocytes in marking the health status of COVID-19 patients. Interestingly, neutrophils exhibited proficient correlation with mainly adaptive immunity-relevant cytokines (e.g., IFN-ƴ, IP-10/CXCL10, IL-2, IL-13, IL-15), pro-neutrophilic IL-8 and CD163 + classical-monocytes in R4 (Fig. 5a). However, in R3, apart from the above-mentioned cytokines, neutrophils gained correlation with TNF-α, IL-10, MCP-1/CCL2, ƴδT cells, IL-17 and effector/naive CD4 T cells, a sign of slightly better (albeit pro-inflammatory) cross-talk with adaptive immune cells and some degree of immunoregulation (IL-10/IL-17) (Fig. 5b). However, the neutrophil-centred clusters considerably “contracted” in R2/R1 (Fig. 5c, d); ultimately resulting in a ‘homeostatic’ cluster in the R1 subgroup, wherein neutrophils co-clustered with naive CD4 + /CD8 + T cells, a sign of immune response resolution and pro-homeostatic orientation [46].
On the level of lymphocytes, the most striking phenotypes were observed in the R4 profile (Fig. 5a); whereas, patterns in R3/R2 were largely transitional and ultimately culminated into a “contracted” lymphocytic correlative-compartment in R1 (Fig. 5b–d). More specifically, in R4 patients (Fig. 5a), the typically infection-resolving lymphocytic compartment (e.g. Th1/Th2 cells, effector/memory CD4/CD8 T cells) had relatively fewer correlations with various effector-function cytokines, including IFN-ƴ, TNF-α, IL-2, IL-15, thereby indicating a certain degree of immunological dysregulation. Most of these lymphocytes also had considerable negative correlation with neutrophils. Interestingly TNF-α, Th17 and Tregs formed a cluster together which might be a sign of pro-inflammatory signalling since Tregs and Th17 have been shown to reciprocally stimulate each other via TNF-signalling pathway [47]. We believe that in the current context, this crosstalk might play a disease-potentiating role in COVID-19 severity. These discords were largely ameliorated from R3 to R1 (e.g. Th17 gaining correlation with Th1/Th2/Tfh and activated-CD8 T cells in R1) thereby indicating that better lymphocytic activity/regulation is beneficial for the recovery of COVID-19 patients.
In conclusion, the above qualitative analyses revealed that neutrophil/lymphocyte-associated inflammation undergoes considerable changes in terms of co-associative immune-components between R4 and R1, such that a relatively contracted neutrophil cluster with a pro-homeostatic orientation and a contracted lymphocyte cluster defines favourable recovery for COVID-19 patients.
Discussion
There has been great interest in defining an early laboratory parameter that can be predictive of disease severity in COVID-19 patients. The muLBSTA score was proposed as an approachable predictor of viral pneumonia severity prior to the COVID-19 pandemic [48], and has also been successfully applied to COVID-19 [49, 50]. However, this score does not take into account a key aspect of COVID-19 pathology, namely the contribution of neutrophils to severe/critical disease. The first clinical reports of COVID-19 patients in Wuhan described elevated neutrophil counts and reduced lymphocyte counts, particularly in patients with severe disease [49, 51]. This results in a higher neutrophil-to-lymphocyte ratio, which was also identified as an early prognostic marker for severe disease [16, 52]. Meta-analyses have confirmed the utility of this ratio not only as a measure of disease severity [53, 54], but also as a predictor of mortality [55]. Importantly, the NLR had a significantly higher adjusted odds ratio than cell counts alone, CRP, lymphocyte-to-monocyte ratio, or platelet-to-lymphocyte ratio [18], thus highlighting the benefit of this ratio over other comparative leukocyte ratios or routinely measured laboratory parameters.
Here, we have performed a longitudinal study in which consecutive peripheral blood samples of 40 surviving COVID-19 patients were analysed via high-dimensional mass cytometry analysis, thus allowing us to uncover detailed insights in the amount and functionality of the white blood cell (WBC) compartment. Importantly, using fresh, non-cryopreserved whole blood samples, our study allowed the analysis of both granulocytes, from which neutrophils take the mainstay and play an important role in COVID-19 [56] (see also below), in addition to the most important populations of monocytes and lymphocytes. Using a whole blood phenotyping approach, we could calculate neutrophil-to-lymphocyte ratios within the same blood samples used for our full analysis.
Using those insights, we grouped all measured samples based on the NLR, as such identifying four blood profiles that defined a biological trajectory through which patients evolved at a highly individualized pace during recovery. Importantly, despite random grouping, these NLR profiles correlated well to various clinical variables, including the level of respiratory support needed at concomitant sampling times. Another striking observation was the large range of NLR levels upon discharge from ICU. This indicates that patients with an increased NLR continue to exhibit a broad range of physical fitness as supported by the longer revalidation periods required following discharge from ICU.
Beyond general clinical features, this cohort also included patients that received various therapeutic interventions, from which the use of corticosteroids (CS) was the most prominent. Our analysis showed that the likelihood of patients-receiving CS therapy correlated well with the duration of ICU stay, indicative of a good clinical selection of patients for this intervention. In addition, correlation analysis of the NLR profiles between patient groups that did and did not receive CS therapy suggests that while both groups exhibited similar trajectories from high to lower ratio levels, the CS group showed a slower evolution towards a normal ratio. While it is known that CS therapy dampens the overall immune response [57], the excessive levels of cytokines and concurrent defective immune response as observed in COVID-19 warrants treatment with CS, in line with other studies [23]. Due to the small number of patients-receiving anti-IL1 or anti-IL6, we cannot make conclusions towards changes in the recovery patterns of their peripheral immune system, even though we did not observe obvious differences with those patients that did not receive these treatments. Overall, the timing of these therapeutic interventions remains critical and still requires further optimization [58, 59]. Indeed, one of the major problems seems to be the delay of the innate immune system in efficiently responding to SARS-CoV-2 infection, which further delays the subsequent adaptive immune response. While this cohort is small, it would be useful to track the phenotypes of severe “long-stay” vs “short-stay” patients as well as non-hospitalized COVID “long haulers” to identify early biomarkers to better predict immune responses to the virus [60].
The defined biological trajectory also allowed us to further elucidate the sequence of events and study the order in which particular immune cell populations reappear in the peripheral blood including the analysis of certain functionalities. Using this approach, we identified that classical monocytes recover first, followed by CD8 + and CD4 + T cells. While these observations are in line with other smaller studies [25, 37], they will need further corroboration in larger cohorts. Overall, the underlying biology suggests that protecting and/or reactivating the innate immune system (from which the monocytes seem first to recover), could be a potential therapeutic strategy. Indeed, based on these fundamental insights, therapies aimed at improving early monocyte functionality could be an interesting avenue to improve the disease course, as suggested by the role for MCP1/CCR2 interference therapy in SARS-CoV1 [12, 61] but also in COVID-19 as previously suggested [37]. We should, however, also look beyond monocytes; a reactivation of the other members of the innate immune system, including DCs and NK cells, could have the potential to ignite a stronger adaptive immune response and promote faster recovery. It also remains to be seen how specific the observed patterns are for COVID-19 and how they diverge from other severe respiratory diseases to tailor future interventions. Moreover, the addition of a comparator cohort on non-survivors could make these observations stronger.
In addition to the immune cell profiles, we also assessed concurrent levels of various cytokines and chemokines and acute phase reactants (i.e. CRP and ferritin). Clustering analysis revealed the gradual release of the pathologically elevated cytokine and chemokine levels, often referred to as a “cytokine storm”, even though the levels in COVID-19 patients are still lower compared to some auto-inflammatory disorders e.g. macrophage activation syndrome. When patients evolved toward lower NLR levels, the coordination of the immune system recovered with a better alignment of cytokines and inflammatory cell types. Indeed, qualitative, correlative matrix analyses further revealed that hyper-inflammatory neutrophil clusters and dysregulated lymphocyte clusters distinguished critically-ill COVID-19 patients. Accordingly, amelioration of these features (i.e., contraction of neutrophil-clusters down to neutrophil-specific immune-signalling and consolidation of lymphocyte-clusters to better crosstalk between different T cell subsets) marked the recovery phase of COVID-19.
Finally, neutrophils are a relatively short-lived and readily activated cell type, complicating their inclusion in detailed immunophenotyping analyses as samples need to be processed immediately after blood collection. Our approach in classifying patients by NLR, integrates deep immunophenotyping with concomitant neutrophil levels in the same blood samples, thus incorporating a key factor in severe COVID-19: the aggravated neutrophil response. Neutrophils provide a major source of reactive oxygen species in an inflammatory response, contributing to tissue damage via oxidative stress [62]. Neutrophils also provide a deadly link between the innate immune response and pathological blood clotting via the formation of neutrophil extracellular traps (NETs) [63]. NETs are comprised of an extracellular meshwork released from activated neutrophils that is both a toxic barrier to pathogens and a scaffold for thrombus formation. NETs likely contribute to the severe lung phenotype seen in COVID-19, with NETs found not only within vascular occlusions in the lung, but also elevated systemically in the blood circulation, especially in critically ill patients [64, 65]. Pro-inflammatory cytokines, activated platelets, and even SARS-CoV-2 itself can lead to activation of neutrophils toward NET release [65, 66]. Our results show significant elevation of several key cytokines contributing to NET formation, including IL-8, TNF-a, and IL-6. NETs also amplify cytokine production by macrophages [67], therefore, allowing potential further exacerbation of the ongoing inflammatory response in COVID-19.
With our results that included analysis of serial sampling from patients over their hospital stay, we could identify an immunocompromised state along increasing NLR subgroups (R2–R4), and also follow recovery of key immune populations together with a return to normal NLR (R1). Future studies integrating our approach with an in-depth analysis of NETosis in neutrophils could provide additional insight into key drivers of severe COVID-19 pathology. This approach using NLR for classification may be applicable not only for COVID-19, but also in other respiratory viral infections, including influenza. Community-acquired influenza A infections still account for significant morbidity and mortality worldwide, with the next pandemic influenza strain potentially arising in coming years. Outbreaks of H7N9 and H5N1 influenza strains have shown similar or more severe pneumonia as COVID-19, with lymphopenia [68, 69]. NLR was retrospectively seen to be a specific and selective marker for mortality in this context as well [70]. There has been tremendous progress in our understanding of COVID-19 pathology in recent months thanks to the global effort undertaken in record time, and this new knowledge can be applied to future viral infections with unfortunately inevitable pandemic potential.
Availability of data and materials
The datasets generated during the current study are available in the Flow repository, https://flowrepository.org/id/FR-FCM-Z34U.
Code availability
All code is available via Github through https://github.com/saeyslab/CYTOF_covid19_study.
References
World Health Organization (2020) COVID-19 weekly epidemiological update reports. Available online at: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports. Accessed Jan 2020
Ma X, Vervoort D (2020) Critical care capacity during the COVID-19 pandemic: global availability of intensive care beds. J Crit Care 58:96–97. https://doi.org/10.1016/j.jcrc.2020.04.012
Ma X, Vervoort D, Reddy CL et al (2020) Emergency and essential surgical healthcare services during COVID-19 in low- and middle-income countries: a perspective. Int J Surg 79:43–46. https://doi.org/10.1016/j.ijsu.2020.05.037
Bong C-L, Brasher C, Chikumba E et al (2020) The COVID-19 pandemic: effects on low- and middle-income countries. Anesth Analg 131:86–92. https://doi.org/10.1213/ANE.0000000000004846
Bielicki JA, Duval X, Gobat N et al (2020) Monitoring approaches for health-care workers during the COVID-19 pandemic. Lancet Infect Dis. https://doi.org/10.1016/S1473-3099(20)30458-8
Blanco-Melo D, Nilsson-Payant BE, Liu W-C et al (2020) Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181:1036-1045.e9. https://doi.org/10.1016/j.cell.2020.04.026
Mann ER, Menon M, Knight SB et al (2020) Longitudinal immune profiling reveals key myeloid signatures associated with COVID-19. Sci Immunol. https://doi.org/10.1126/sciimmunol.abd6197
Wauters E, Thevissen K, Wouters C et al (2020) Establishing a Unified COVID-19 “Immunome”: Integrating Coronavirus Pathogenesis and Host Immunopathology. Front Immunol 11:1642. https://doi.org/10.3389/fimmu.2020.01642
Pedersen SF, Ho Y-C (2020) SARS-CoV-2: a storm is raging. J Clin Invest 130:2202–2205. https://doi.org/10.1172/JCI137647
Schulte-Schrepping J, Reusch N, Paclik D et al (2020) Severe COVID-19 is marked by a dysregulated myeloid cell compartment. Cell. https://doi.org/10.1016/j.cell.2020.08.001
Wilk AJ, Rustagi A, Zhao NQ et al (2020) A single-cell atlas of the peripheral immune response in patients with severe COVID-19. Nat Med 26:1070–1076. https://doi.org/10.1038/s41591-020-0944-y
Vanderbeke L, Van Mol P, Van Herck Y et al (2020) Monocyte-driven atypical cytokine storm and aberrant neutrophil activation as key mediators of COVID19 disease severity. SSRN. https://doi.org/10.2139/ssrn.3646561
Lucas C, Wong P, Klein J et al (2020) Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584:463–469. https://doi.org/10.1038/s41586-020-2588-y
Zhao Y, Qin L, Zhang P et al (2020) Longitudinal COVID-19 profiling associates IL-1RA and IL-10 with disease severity and RANTES with mild disease. JCI Insight. https://doi.org/10.1172/jci.insight.139834
Del Valle DM, Kim-Schulze S, Huang H-H et al (2020) An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat Med 26:1636–1643. https://doi.org/10.1038/s41591-020-1051-9
Liu J, Liu Y, Xiang P et al (2020) Neutrophil-to-lymphocyte ratio predicts critical illness patients with 2019 coronavirus disease in the early stage. J Transl Med 18:206. https://doi.org/10.1186/s12967-020-02374-0
Fu J, Kong J, Wang W et al (2020) The clinical implication of dynamic neutrophil to lymphocyte ratio and D-dimer in COVID-19: A retrospective study in Suzhou China. Thromb Res 192:3–8. https://doi.org/10.1016/j.thromres.2020.05.006
Yang A-P, Liu J-P, Tao W-Q, Li H-M (2020) The diagnostic and predictive role of NLR, d-NLR and PLR in COVID-19 patients. Int Immunopharmacol 84:106504. https://doi.org/10.1016/j.intimp.2020.106504
Ma A, Cheng J, Yang J et al (2020) Neutrophil-to-lymphocyte ratio as a predictive biomarker for moderate-severe ARDS in severe COVID-19 patients. Crit Care 24:288. https://doi.org/10.1186/s13054-020-03007-0
Liu J, Li S, Liu J et al (2020) Longitudinal characteristics of lymphocyte responses and cytokine profiles in the peripheral blood of SARS-CoV-2 infected patients. EBioMedicine 55:102763. https://doi.org/10.1016/j.ebiom.2020.102763
Carissimo G, Xu W, Kwok I et al (2020) Whole blood immunophenotyping uncovers immature neutrophil-to-VD2 T-cell ratio as an early marker for severe COVID-19. Nat Commun 11:5243. https://doi.org/10.1038/s41467-020-19080-6
Silvin A, Chapuis N, Dunsmore G et al (2020) Elevated calprotectin and abnormal myeloid cell subsets discriminate severe from mild COVID-19. Cell 182:1401-1418.e18. https://doi.org/10.1016/j.cell.2020.08.002
RECOVERY Collaborative Group, Horby P, Lim WS et al (2020) Dexamethasone in hospitalized patients with Covid-19—preliminary report. N Engl J Med. https://doi.org/10.1056/NEJMoa2021436
Rodriguez L, Pekkarinen PT, Lakshmikanth T et al (2020) Systems-level immunomonitoring from acute to recovery phase of severe COVID-19. Cell reports Med 1:100078. https://doi.org/10.1016/j.xcrm.2020.100078
Wen W, Su W, Tang H et al (2020) Immune cell profiling of COVID-19 patients in the recovery stage by single-cell sequencing. Cell Discov 6:31. https://doi.org/10.1038/s41421-020-0168-9
Odak I, Barros-Martins J, Bošnjak B et al (2020) Reappearance of effector T cells is associated with recovery from COVID-19. EBioMedicine 57:102885. https://doi.org/10.1016/j.ebiom.2020.102885
Gietema HA, Zelis N, Nobel JM et al (2020) CT in relation to RT-PCR in diagnosing COVID-19 in The Netherlands: a prospective study. PLoS ONE 15:e0235844. https://doi.org/10.1371/journal.pone.0235844
Van Gassen S, Callebaut B, Van Helden MJ et al (2015) FlowSOM: using self-organizing maps for visualization and interpretation of cytometry data. Cytometry A 87:636–645. https://doi.org/10.1002/cyto.a.22625
Jin J-M, Bai P, He W et al (2020) Gender differences in patients with COVID-19: focus on severity and mortality. Front Public Heal 8:152. https://doi.org/10.3389/fpubh.2020.00152
Chen T, Wu D, Chen H et al (2020) Clinical characteristics of 113 deceased patients with coronavirus disease 2019: retrospective study. BMJ 368:m1091. https://doi.org/10.1136/bmj.m1091
Gemmati D, Bramanti B, Serino ML et al (2020) COVID-19 and individual genetic susceptibility/receptivity: role of ACE1/ACE2 genes, immunity, inflammation and coagulation. Might the double X-chromosome in females be protective against SARS-CoV-2 compared to the single X-chromosome in males? Int J Mol Sci. https://doi.org/10.3390/ijms21103474
Rod JE, Oviedo-Trespalacios O, Cortes-Ramirez J (2020) A brief-review of the risk factors for covid-19 severity. Rev Saude Publica 54:60. https://doi.org/10.11606/s1518-8787.2020054002481
Vincent JL, de Mendonça A, Cantraine F et al (1998) Use of the SOFA score to assess the incidence of organ dysfunction/failure in intensive care units: results of a multicenter, prospective study. Working group on “sepsis-related problems” of the European Society of Intensive Care Medicine. Crit Care Med 26:1793–1800. https://doi.org/10.1097/00003246-199811000-00016
Singer M, Deutschman CS, Seymour CW et al (2016) The third international consensus definitions for sepsis and septic shock (Sepsis-3). JAMA 315:801–810. https://doi.org/10.1001/jama.2016.0287
Tuty Kuswardhani RA, Henrina J, Pranata R et al (2020) Charlson comorbidity index and a composite of poor outcomes in COVID-19 patients: a systematic review and meta-analysis. Diabetes Metab Syndr 14:2103–2109. https://doi.org/10.1016/j.dsx.2020.10.022
Maloy KJ, Burkhart C, Junt TM et al (2000) CD4(+) T cell subsets during virus infection. Protective capacity depends on effector cytokine secretion and on migratory capability. J Exp Med 191:2159–2170. https://doi.org/10.1084/jem.191.12.2159
Chua RL, Lukassen S, Trump S et al (2020) COVID-19 severity correlates with airway epithelium-immune cell interactions identified by single-cell analysis. Nat Biotechnol 38:970–979. https://doi.org/10.1038/s41587-020-0602-4
Cervantes-Barragan L, Züst R, Weber F et al (2007) Control of coronavirus infection through plasmacytoid dendritic-cell-derived type I interferon. Blood 109:1131–1137. https://doi.org/10.1182/blood-2006-05-023770
Scheuplein VA, Seifried J, Malczyk AH et al (2015) High secretion of interferons by human plasmacytoid dendritic cells upon recognition of Middle East respiratory syndrome coronavirus. J Virol 89:3859–3869. https://doi.org/10.1128/JVI.03607-14
Boyette LB, Macedo C, Hadi K et al (2017) Phenotype, function, and differentiation potential of human monocyte subsets. PLoS ONE 12:e0176460. https://doi.org/10.1371/journal.pone.0176460
Hou W, Gibbs JS, Lu X et al (2012) Viral infection triggers rapid differentiation of human blood monocytes into dendritic cells. Blood 119:3128–3131. https://doi.org/10.1182/blood-2011-09-379479
Lambotin M, Raghuraman S, Stoll-Keller F et al (2010) A look behind closed doors: interaction of persistent viruses with dendritic cells. Nat Rev Microbiol 8:350–360. https://doi.org/10.1038/nrmicro2332
Abers MS, Delmonte OM, Ricotta EE et al (2021) An immune-based biomarker signature is associated with mortality in COVID-19 patients. JCI Insight. https://doi.org/10.1172/jci.insight.144455
Costela-Ruiz VJ, Illescas-Montes R, Puerta-Puerta JM et al (2020) SARS-CoV-2 infection: the role of cytokines in COVID-19 disease. Cytokine Growth Factor Rev 54:62–75. https://doi.org/10.1016/j.cytogfr.2020.06.001
Griffith JW, Sokol CL, Luster AD (2014) Chemokines and chemokine receptors: positioning cells for host defense and immunity. Annu Rev Immunol 32:659–702. https://doi.org/10.1146/annurev-immunol-032713-120145
Garg AD, Vandenberk L, Fang S et al (2017) Pathogen response-like recruitment and activation of neutrophils by sterile immunogenic dying cells drives neutrophil-mediated residual cell killing. Cell Death Differ 24:832–843. https://doi.org/10.1038/cdd.2017.15
Chen X, Oppenheim JJ (2014) Th17 cells and Tregs: unlikely allies. J Leukoc Biol 95:723–731. https://doi.org/10.1189/jlb.1213633
Guo L, Wei D, Zhang X et al (2019) Clinical features predicting mortality risk in patients with viral pneumonia: the MuLBSTA score. Front Microbiol 10:2752. https://doi.org/10.3389/fmicb.2019.02752
Chen N, Zhou M, Dong X et al (2020) Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet (London, England) 395:507–513. https://doi.org/10.1016/S0140-6736(20)30211-7
Tusha J, Khanam V, Tegeltija V, Kumar S (2020) The mulbsta score: predicting risk of mortality and disease severity in patients with COVID-19 pneumonia. Chest 158:A300. https://doi.org/10.1016/j.chest.2020.08.299
Huang C, Wang Y, Li X et al (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet (London, England) 395:497–506. https://doi.org/10.1016/S0140-6736(20)30183-5
Kong M, Zhang H, Cao X et al (2020) Higher level of neutrophil-to-lymphocyte is associated with severe COVID-19. Epidemiol Infect 148:e139. https://doi.org/10.1017/S0950268820001557
Lagunas-Rangel FA (2020) Neutrophil-to-lymphocyte ratio and lymphocyte-to-C-reactive protein ratio in patients with severe coronavirus disease 2019 (COVID-19): a meta-analysis. J Med Virol 92:1733–1734. https://doi.org/10.1002/jmv.25819
Feng X, Li S, Sun Q et al (2020) Immune-inflammatory parameters in COVID-19 cases: a systematic review and meta-analysis. Front Med 7:301. https://doi.org/10.3389/fmed.2020.00301
Li X, Liu C, Mao Z et al (2020) Predictive values of neutrophil-to-lymphocyte ratio on disease severity and mortality in COVID-19 patients: a systematic review and meta-analysis. Crit Care 24:647. https://doi.org/10.1186/s13054-020-03374-8
Wang J, Li Q, Yin Y et al (2020) Excessive neutrophils and neutrophil extracellular traps in COVID-19. Front Immunol 11:2063. https://doi.org/10.3389/fimmu.2020.02063
Cain DW, Cidlowski JA (2017) Immune regulation by glucocorticoids. Nat Rev Immunol 17:233–247. https://doi.org/10.1038/nri.2017.1
Galluccio F, Ergonenc T, Garcia Martos A et al (2020) Treatment algorithm for COVID-19: a multidisciplinary point of view. Clin Rheumatol 39:2077–2084. https://doi.org/10.1007/s10067-020-05179-0
Menzella F, Biava M, Barbieri C et al (2020) Pharmacological treatment of COVID-19: lights and shadows. Drugs Context. https://doi.org/10.7573/dic.2020-4-6
Park A, Iwasaki A (2020) Type I and type III interferons—induction, signaling, evasion, and application to combat COVID-19. Cell Host Microbe 27:870–878. https://doi.org/10.1016/j.chom.2020.05.008
Channappanavar R, Fehr AR, Vijay R et al (2016) Dysregulated type I interferon and inflammatory monocyte-macrophage responses cause lethal pneumonia in SARS-CoV-infected mice. Cell Host Microbe 19:181–193. https://doi.org/10.1016/j.chom.2016.01.007
Laforge M, Elbim C, Frère C et al (2020) Tissue damage from neutrophil-induced oxidative stress in COVID-19. Nat Rev Immunol 20:515–516. https://doi.org/10.1038/s41577-020-0407-1
Sorvillo N, Cherpokova D, Martinod K, Wagner DD (2019) Extracellular DNA NET-works with dire consequences for health. Circ Res 125:470–488. https://doi.org/10.1161/CIRCRESAHA.119.314581
Middleton EA, He X-Y, Denorme F et al (2020) Neutrophil extracellular traps contribute to immunothrombosis in COVID-19 acute respiratory distress syndrome. Blood 136:1169–1179. https://doi.org/10.1182/blood.2020007008
Nicolai L, Leunig A, Brambs S et al (2020) Immunothrombotic dysregulation in COVID-19 pneumonia is associated with respiratory failure and coagulopathy. Circulation 142:1176–1189. https://doi.org/10.1161/CIRCULATIONAHA.120.048488
Veras FP, Pontelli MC, Silva CM et al (2020) SARS-CoV-2-triggered neutrophil extracellular traps mediate COVID-19 pathology. J Exp Med. https://doi.org/10.1084/jem.20201129
Warnatsch A, Ioannou M, Wang Q, Papayannopoulos V (2015) Inflammation. Neutrophil extracellular traps license macrophages for cytokine production in atherosclerosis. Science 349:316–320. https://doi.org/10.1126/science.aaa8064
Gao H-N, Lu H-Z, Cao B et al (2013) Clinical findings in 111 cases of influenza A (H7N9) virus infection. N Engl J Med 368:2277–2285. https://doi.org/10.1056/NEJMoa1305584
Yuen KY, Chan PKS, Peiris M et al (1998) Clinical features and rapid viral diagnosis of human disease associated with avian influenza A H5N1 virus. Lancet 351:467–471. https://doi.org/10.1016/S0140-6736(98)01182-9
Zhang Y, Zou P, Gao H et al (2019) Neutrophil-lymphocyte ratio as an early new marker in AIV-H7N9-infected patients: a retrospective study. Ther Clin Risk Manag 15:911–919. https://doi.org/10.2147/TCRM.S206930
Acknowledgements
We would like to thank Tatjana Verbeke, Marleen Derweduwe, Annelies Claeys, Reena Chinnaraj and Jenny Sprooten for technical support for the CyTOF experiments. This work was supported by FWO grant I007418N, KULeuven grant C14/17/084, KOOR funding and ‘het Leuven Kankerinstituut’. S.V.G. is an ISAC Marylou Ingram Scholar and supported by an FWO postdoctoral research grant (Research Foundation—Flanders). L. V. is supported by an FWO PhD fellowship (Research Foundation—Flanders, E11E9819N). We particularly thank Dr. Toine Mercier for his help in the design of the patient and sample collection database.
Additional Contagious consortium members
Michael Casaer, Dieter Dauwe, Jan Gunst, Greet Hermans, Stephanie Humblet-Baron, Diether Lambrechts, Adrian Liston, Natalie Lorent, Philippe Meersseman, Johan Neyts, Paul Proost, Jeroen Raes, Stephen Rex, Sabine Tejpar, Karin Thevissen, Thomas Tousseyn, Birgit Weynand, Alexander Wilmer, Carine Wouters.
Funding
This work was supported by FWO grant I007418N, KULeuven grant C14/17/084, KOOR funding and ‘het Leuven Kankerinstituut’. S.V.G. is an ISAC Marylou Ingram Scholar and supported by an FWO postdoctoral research grant (Research Foundation—Flanders). L. V. is supported by an FWO PhD fellowship (Research Foundation—Flanders, E11E9819N).
Author information
Authors and Affiliations
Consortia
Contributions
Conceptualization, FDS; methodology, PAP, SVG, DP; software/data Analysis, SVG, KQ, AE, AA, YS, FMB, FDS; investigation, PAP, FDS; clinical trial design, data acquisition and interpretation, LV, JW, YVH, EW, PVM; cyto/chemokine data acquisition, JF, BMD, KA; integrative mapping and interpretation of cyto/chemokine data, ADG, DMB; writing–original draft, PAP, SVG, DP, ADG, FDS; writing–review and editing, PAP, SVG, DP, LV, YVH, PVM, EW, PM, KM, ADG, JW, and FDS. This work was fully supported by the Contagious Consortium.
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts of interest to declare.
Ethics approval and consent to participate
All enrolled patients signed an informed consent to participate. The trial (S63881) was approved by the Ethical committee Research of UZLeuven/KULeuven on March 26 2020 and is registered on clinicaltrials.gov (NCT04327570)—https://clinicaltrials.gov/ct2/show/NCT04327570. All patients that entered the trial gave explicit prior consent. All patients were enrolled at the ward of the hospital prior to them entering ICU.
Consent for publication
Not applicable. Overall, identifying details (names, dates of birth, identity numbers, biometrical characteristics (such as facial features, fingerprint, writing style, voice pattern, DNA or other distinguishing characteristic) and other information) of the participants that were studied are not published.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Members of The CONTAGIOUS consortium are listed in the Acknowledgement section.
Supplementary Information
Below is the link to the electronic supplementary material.
18_2021_3808_MOESM1_ESM.tiff
Supplementary file1 Supplementary Figure 1 Parts A and B – Gating and clustering analysis for CyTOF data. (A) 123 .fcs files containing data collected from patients (n=40) and healthy controls (n=8) were pre-processed by Maxpar PathsetterTM software. Quality Control analysis was performed using the PeacoCQ algorithm to remove aberrant cells. Using 11 markers, FlowSOM model 1 was generated and data mapping was done by splitting neutrophils/eosinophils (black dots) from the rest of the white blood cells (turquoise). Two additional FlowSOM models were built using 33 markers (non-granulocytes and granulocytes) which generated 208 clusters represented in two heatmaps, manually annotated as 24 cell populations. Neutrophil and lymphocyte percentages were extracted and used to calculate the NLR. Final statistics were obtained using a linear mixed model to account for random patient effects. (B) Two heatmaps generated from 2 FlowSOM models using 33 markers and manually annotated: 100 granulocyte clusters, left panel and 108 non-granulocyte clusters, right panel. (TIFF 55364 KB)
18_2021_3808_MOESM2_ESM.tiff
Supplementary file2 Supplementary Figure 2 - Overview of cell counts, as measured by CyTOF (left column) or using a standard clinical cytometer (middle column). These two independent measurements show a strong correlation (as shown in the last column). Overall, this figure illustrates that the decrease in lymphocytes and increase in neutrophils is not only happening from the relative point of view, but also in absolute cell counts. (TIFF 168762 KB)
18_2021_3808_MOESM3_ESM.tiff
Supplementary file3 Supplementary Figure 3 Parts A and B - tSNE visualisation of the CyTOF data. 25,000 cells were selected from both the granulocyte and the non-granulocyte FlowSOM model, for a resulting total of 50,000 cells. The measurements for the 30 DIP channels were used to compute a tSNE dimensionality reduction, using default parameters (perplexity = 30). (A) Coloured by the labels assigned to the FlowSOM clusters. (B) Coloured by arcsinh transformed marker expression for the 30 DIP markers. (TIFF 39378 KB)
18_2021_3808_MOESM4_ESM.tiff
Supplementary file4 Supplementary Figure 4 - Scatter plot of maximum respiratory support vs. length of stay. Scatter plot showing the maximum level of respiratory support patients received from 1-5 where 1=oxygen, 2=high flow nasal cannula, 3=non-invasive ventilation, 4=invasive mechanical ventilation (IMV) and 5=IMV+Prone/extracorporeal membrane oxygenation/inhaled nitric oxide versus total length of ICU stay in days (n=40). (TIFF 52739 KB)
18_2021_3808_MOESM5_ESM.tiff
Supplementary file5 Supplementary Figure 5 - Heatmap overview of CyTOF measurements. For all 123 samples used in this study, the abundance of 25 cell populations was determined based on FlowSOM clustering. The percentage of each of these populations is visualized in the heatmap. We labeled the samples according to their neutrophil over lymphocyte ratio, ordered from R4 (strong increase in neutrophils) to R1 (approx. normal neutrophil/lymphocyte ratio). These groups correlate with additional clinical properties of the samples, as outlined further in the manuscript. (TIFF 42191 KB)
18_2021_3808_MOESM6_ESM.tiff
Supplementary file6 Supplementary Figure 6 - Correlation analysis of various clinical features to the blood profiles. (A) Boxplot representation of the Body Mass Index (BMI) according to the highest NLR measured in each patient. (B) Boxplot representation of each patient’s age according to the highest NLR measured in each patient. (C-D) Continuous NLR value compared to BMI and age. Pearson correlations of 0.196 (BMI) and 0.005 (age) (E) Boxplot representation comparing patients that did not (left) or did (right) receive treatment with corticosteroids during their stay at ICU. Open circles indicate which patients received additional anti-IL-1 or anti-IL-6 treatment. (n: R4 = 3, R3 = 11, R2 = 11, R1 = 14) (A-D) n: R4 = 18, R3 = 12, R2 = 6, R1 = 4 (A-E) Colour codes used in this graph correlate to the measured NLR groups as indicated in this manuscript, black: R4, green: R3, blue: R2, cyan R1. (TIFF 16740 KB)
18_2021_3808_MOESM7_ESM.tiff
Supplementary file7 Supplementary Figure 7 - Detailed clinical trajectory of each individual patient. Line plot representation indicating the clinical evolution of each individual patient during their stay at ICU according to a timeline starting at their admission to ICU. The Y-axis of each graph indicates the NLR group (R1-4) as measured in each individual sample. The colour code of each dot indicates the time point of sampling: grey: admission to ward, red: admission ICU, orange: intermediate ICU, green: discharge ICU. (TIFF 56255 KB)
18_2021_3808_MOESM8_ESM.tiff
Supplementary file8 Supplementary Figure 8 - HLA-DR levels in the monocyte populations across R immune types. Boxplots showing the mean metal intensity (MMI) of HLA-DR expression in the various monocyte populations (classical monocytes, intermediate monocytes, myeloid dendritic cells (mDC), non-classical monocytes and plasmacytoid dendritic cells (pDC)) across the R1-4 immune types. Colour codes are as follows: forest green: classical monocytes, yellow: intermediate monocytes, lime-green: mDC, turquoise-green: non-classical monocytes and orange: pDC. (TIFF 84381 KB)
Rights and permissions
About this article
Cite this article
Penttilä, P.A., Van Gassen, S., Panovska, D. et al. High dimensional profiling identifies specific immune types along the recovery trajectories of critically ill COVID19 patients. Cell. Mol. Life Sci. 78, 3987–4002 (2021). https://doi.org/10.1007/s00018-021-03808-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-021-03808-8