Abstract
Squamous cell carcinoma (SCC) is an aggressive malignancy that can originate from various organs. TP63 is a master regulator that plays an essential role in epidermal differentiation. It is also a lineage-dependent oncogene in SCC. ΔNp63α is the prominent isoform of TP63 expressed in epidermal cells and SCC, and overexpression promotes SCC development through a variety of mechanisms. Recently, ΔNp63α was highlighted to act as an epidermal-specific pioneer factor that binds closed chromatin and enhances chromatin accessibility at epidermal enhancers. ΔNp63α coordinates chromatin-remodeling enzymes to orchestrate the tissue-specific enhancer landscape and three-dimensional high-order architecture of chromatin. Moreover, ΔNp63α establishes squamous-like enhancer landscapes to drive oncogenic target expression during SCC development. Importantly, ΔNp63α acts as an upstream regulator of super enhancers to activate a number of oncogenic transcripts linked to poor prognosis in SCC. Mechanistically, ΔNp63α activates genes transcription through physically interacting with a number of epigenetic modulators to establish enhancers and enhance chromatin accessibility. In contrast, ΔNp63α also represses gene transcription via interacting with repressive epigenetic regulators. ΔNp63α expression is regulated at multiple levels, including transcriptional, post-transcriptional, and post-translational levels. In this review, we summarize recent advances of p63 in epigenomic and transcriptional control, as well as the mechanistic regulation of p63.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Squamous cell carcinomas (SCCs) are a series of aggressive malignancies from various tissue origins, including the skin, head and neck, lung, and esophagus [1, 2]. Squamous subtype cancers can also be identified in subsets of pancreatic [3,4,5], urothelial [6], and prostate cancer [7]. SCCs from various organs share some similar features, including their genomic landscapes, morphological characteristics, and molecular alterations [2, 8]. Human papillomavirus infection and consumption of alcohol and tobacco are defined etiology factors for SCCs [9]. To date, SCCs are highly aggressive and current therapeutics have limited effect, and do not provide a satisfactory clinical outcome [2].
Lineage dependency occurs where cancer cells depend on survival and self-renewal mechanisms co-opted from the original healthy tissues from where they arose [10]. Transcription factor (TF) TP63 is a p53 family member which plays a crucial role in epidermal development. Alternative usage of different promoters of TP63 gene results in two major isoforms, TAp63 and NH2-terminal truncated ΔNp63, which lacks the canonical transactivating domain (TA1) (Fig. 1a) [11]. However, it is proposed that ∆Np63 has an intrinsic transcription transactivation ability, which is conferred by a second transactivation domain (TA2) located between exons 11 and 12 [12]. Alternative splicing at the COOH terminal further generates variants of each isoform (α, β, γ) [11]. The δ variant is generated by the skipping of exons 12 and 13, whereas the ε variant arises from a premature transcriptional termination in intron 10 [13] (Fig. 1b). All p63 isoforms contain a central DNA-binding domain (DBD) and an oligomerization domain (OD). The α proteins are the longest isoforms, containing a COOH-terminal sterile alpha motif (SAM) domain, which mediates protein–protein interaction. SAM domain is followed by an inhibitory domain (ID), which auto-inhibits the transcriptional activity of the TA1 domain [14]. The β isoforms contain the TA2 domain, but lack both SAM and ID domains. The γ variants contain an OD domain, followed by a unique sequence derived from intron 10 (Fig. 1b) [13].
The p63 gene and protein structures. a Genomic structure of human TP63 gene. Alternative promoters (P1 and P2) are indicated. Alternative splicing events at the COOH terminus generate variants α, β, and γ. Exons skipping or premature transcription termination produces variants δ and ε, respectively. b Schematic diagrams of p63 protein isoforms structures. TA1 transactivating domain, TA* truncated transactivating domain of ΔN isoforms, DBD DNA-binding domain, OD oligomerization domain, TA2 secondary transactivating domain, SAM sterile alpha motif, ID inhibitory domain
The TAp63 and ∆Np63 isoforms regulate distinct target gene sets and often exert opposing regulatory functions [15, 16]. Heterozygosity of p63 prevents spontaneous and chemical-induced SCC formation [17, 18], indicating that TP63 acts as a lineage-survival oncogene in SCCs. ΔNp63α is the prominent isoform of the TP63 gene expressed in keratinocytes and basal cells of diverse epithelia [19,20,21] and various SCCs [2]. ΔNp63α promotes SCC development through regulating different target genes, including cell growth and proliferation [22, 23], extracellular matrix (ECM)–receptor interaction [24], cell adhesion [25,26,27], glucose metabolism [28, 29], anti-oxidant defense [30, 31], DNA damage repair [32, 33], terminal differentiation [34, 35], and inflammation and angiogenesis [36,37,38,39,40,41] (Fig. 2). Sustained expression of ∆Np63α tends to promote the well-differentiated SCC phenotype and restricts epithelial–mesenchymal transition induced by TGF-β [42]. However, both suppressive [43,44,45] and promotive roles [46] of ∆Np63α in cell invasion and metastasis have been demonstrated in various cell contexts, suggesting a cell context-specific role in cancer metastasis. Growing evidence demonstrates that ΔNp63 exerts both a suppressive and promotive role on target genes expression [47,48,49]. Recent studies established the master regulator role of p63 in controlling chromatin accessibility and enhancer reprogramming in keratinocytes [50, 51]. Various epigenetic modulators, including chromatin-remodeling complexes and epigenetic enzymes, are implicated in p63-mediated epigenomic reprogramming during keratinocyte differentiation and SCC development [52]. In this review, we focus on the role and mechanisms of p63 in chromatin remodeling and enhancer reprogramming during epidermal differentiation and SCC development.
Diverse functions of ΔNp63α in SCCs. ΔNp63α promotes the development of malignant features of SCC through multiple mechanisms, including stimulation of cell growth and survival, inhibition of terminal differentiation, reprogramming of glucose metabolism and maintaining anti-oxidative homeostasis, promotion of DNA damage repair and triggering of inflammation and angiogenesis. ΔNp63α is also understood to regulate cell adhesion and remodeling of the ECM in tumor microenvironments. Representative target genes of ΔNp63α in SCC are shown. Red or green represents positively or negatively regulated target gene
TP63 in epidermal commitment and differentiation
TP63 plays essential roles in lineage commitment during epidermal development through a variety of epigenetic mechanisms (Table 1). Mutations of TP63 are associated with various developmental disorders, including ectrodactyly–ectodermal dysplasia–cleftlip/palate (EEC) syndrome [53], ankyloblepharon–ectodermal defects–cleft lip/palate (AEC) syndrome and split-hand/foot malformation-IV syndrome. Ablation of all p63 isoforms in mice results in the absence of stratified epithelia and their derivatives [54,55,56], and specific ablation of ΔNp63 isoform leads to severe developmental anomalies similar to that existing in p63-null mice. These abnormalities include truncated forelimbs and the absence of hind limbs [57], indicating that the TP63 gene is required for skin and epithelial development. However, both p63- and ΔNp63-deficient epidermis expresses the terminal differentiation markers loricrin, filaggrin, and involucrin [54,55,56], although there is a premature expression pattern of terminal differentiation markers at E15.5 in ΔNp63 isoform-specifically ablated mice [57]. This suggests that TP63 or ΔNp63 is dispensable for epidermal lineage commitment. Intriguingly, introducing p63 mutants from EEC patients into induced pluripotent stem cells (iPSCs) dramatically impairs the induction of epidermal marker K14 in an epidermal commitment model [58] induced by retinoic acid (RA) and bone morphogenetic protein 4 (BMP4) [59], arguing that TP63 is essential for epidermal commitment. These opposite interpretations may indicate that other lineage-determining factors could compensate for p63 function in a p63-null context, whereas mutant p63 in iPSCs abrogates the function of the normal p63. Although p63 is dispensable to drive human embryonic stem cells (hESCs) to differentiate to surface ectoderm progenitor cells, it is required for further differentiation toward functional keratinocytes upon RA/BMP4 treatment [60]. It has been shown that genes expressed at an early differentiation stage are not under the control of p63. The p63 protein predominantly regulates genes during the specification switch from the multipotent state to the epidermal fate [59]. Thus, these studies suggest that p63 plays a prominent role in maturation, rather than the initiation stage of skin differentiation triggered by inductive morphogens (Table 1).
TP63 acts as an epidermal pioneer factor to open chromatin regions
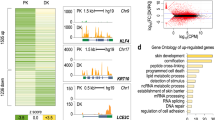
It has been speculated that TP53 family members may exert intrinsic pioneer factor activity to specify epithelial lineage-specific chromatin landscape [61], and this may be the case with p63. The p63 protein preferentially binds nucleosome-enriched regions marked with active histone modifications in epidermal keratinocytes, which are inaccessible in other lineages without p63 expression [51, 62, 63], implying that p63 is required for establishing an epidermal-specific chromatin architecture at its binding regions. The primary keratinocytes from EEC syndrome patients harboring heterozygous mutations in the p63 DNA-binding domain display a dramatic reduction of chromatin accessibility at p63-binding sites compared to normal cells [64]. By using single-cell transcriptomic and epigenomic profiling techniques, Fan et al. demonstrated that ΔNp63α binding generates open chromatin regions near or within the genes involved in epidermal fate specification of skin [65]. Furthermore, Lin-Shiao et al. used p63 mutants, lacking DNA-binding activity, to demonstrate that ΔNp63α DNA binding is required to establish keratinocyte-specific enhancers [66]. During zebrafish embryo development, p63 binding occurs prior to chromatin opening at p63-binding sites (Table 1). Remarkably, the pioneered p63-binding sites are preferentially associated with epidermal-expressing genes, as is the case of lama5. The chromatin accessibility of the pioneered p63-binding sites is significantly reduced in tp63−/− mutants. Consequently, the levels of expression of these epidermal genes are significantly decreased in the tp63−/− mutant, such as col18a1a [67].
Loss of p63 reduces the chromatin accessibility of keratinocyte-specific genes, such as KRT5. Furthermore, depletion of p63 reduces the chromatin accessibility and occupancy probability of other maturation-associated TFs, including p53, RFX, and Kruppel-like factor 4 (KLF4) [68], indicating that p63 exerts intrinsic pioneer factor activity to allow other lineage-specific TFs to bind (Fig. 3). Interestingly, ΔNp63 (−/−) epidermal cells express a number of transcripts associated with embryonic stem cells (ESCs) and induced pluripotent stem cells (iPSCs) and display compromised epithelial identity [69], suggesting that ΔNp63 exerts repressive role on genes expressed at the early differentiation stage once surface ectoderm progenitor cells enter maturation stage. Mechanistically, p63 reduces chromatin accessibility at morphogen-dependent accessible sites through promoting deposition of H3K27me3 at these sites during the maturation stage [60]. These studies consistently highlight that p63 exerts intrinsic pioneer factor activity, to open inaccessible chromatin, and it cooperates with specific TFs during keratinocyte maturation.
p63 acts as an epidermal pioneer factor. P63 binds to compacted chromatin regions and recruits multiple epigenetic regulators, including COMPASS complex, histone acetyltransferases p300/CBP, SNF/SWF chromatin remodeling complex, DNMT3 and TETs family members, leading to an increase in chromatin accessibility and formation of active enhancers at epidermal genes
TP63 establishes epidermal enhancer landscape
Enhancers are cis-regulatory elements defined by H3K4me1 modification. Enhancers can be categorized as poised or active depending on the co-occupancy of H3K27me3 or H3K27ac, respectively [70, 71]. The specific spatiotemporal regulation of developmental gene transcription is mainly determined by dynamic alterations in enhancer activity [70, 72, 73]. Active enhancers produce bidirectional enhancer RNAs (eRNAs), which can interact with various TFs and stabilize enhancer–promoter loops [74]. Remarkably, the chromatin state of enhancer regions changes more prominently than the promoter regions, as iPSCs gradually differentiate to keratinocytes [68]. Enhancers associated with genes that determine the ectoderm lineage specification, including ITGA6, KRT5, and KRT14, are dramatically decommissioned as epidermal stem cells (EpSCs) differentiate to keratinocytes, whereas enhancers driving the expression of epidermal differentiation genes such as IVL and SPRR1A are activated de novo, which is accompanied by a shift of Dnmt3a genomic binding from the former enhancers cluster to the latter [75].
Chromatin immunoprecipitation-sequencing (ChIP-seq) has revealed that, while the majority of p63-bound regulatory elements and p63-bound open chromatin regions in epidermal keratinocytes are active enhancers, and only a small number of p63-bound regions belong to the promoter [50, 51, 63, 64, 76], suggesting a predominant p63 role in enhancer-mediated transactivation. The p63-bound enhancers are dynamically changed during epidermal commitment. It has been demonstrated that p63 and KLF4 gain accessibility to their target enhancers as surface ectoderm progenitor differentiate to mature keratinocytes [68]. Furthermore, by using an inducible transdifferentiation model, Lin-Shiao et al. demonstrated that inducible expression of ΔNp63α in fibroblasts leads to de novo H3K27ac and an increase in chromatin accessibility at p63-bound sites, as well as expression of genes important for epithelial lineage specification [66]. Co-expression of ΔNp63α and KLF4 in fibroblasts is sufficient to activate transcription of keratin 14 and convert fibroblasts into keratinocyte-like cells. Interestingly, either ΔNp63α or KLF4 alone could not induce expression of keratin 14 [66], suggesting the interaction between ΔNp63α and KLF4 is required for full epidermal commitment. EEC syndrome patients display keratinocytes harboring p63 mutations, which lack DNA-binding ability, and display a loss of enhancers at epidermal-expressing genes, indicating that ΔNp63α binding of DNA is necessary to establish the epidermal enhancer landscape and epidermal commitment [64, 77]. Furthermore, the majority of p63-bound epidermal enhancers lose their accessibility in p63 knockout ESC during in vitro differentiation [59]. Taken together, these studies indicate that the formation and maintenance of epidermal enhancers at p63-binding sites are heavily dependent on p63 expression and its binding to DNA (Table 1).
TP63 represses nonepithelial genes
It has been demonstrated that embryonic epithelia from p63-null mice display compromised epithelial fate, concomitant with the activation of mesodermal-specific genes [78], suggesting that p63 represses the nonepithelial transcriptional program. Introducing p63 mutants from EEC patients into iPSCs impairs the acquisition of epithelial identity, but also activates mesodermal genes expression. In contrast, inhibition of mesodermal induction by suramin enhances epidermal differentiation of iPSCs carrying the p63 mutation [59]. Cbx4, a component of polycomb repressive complex 1 (PRC1), is a target of p63 and contributes to repression of nonepithelial genes during epidermal differentiation [79]. In the early stages of embryonic development, p63 prevents Sox3 binding to pre-established neural genes enhancers and represses neural gene expression, demonstrating that p63 restricts neural lineage determination during the early stages of embryonic development [67] (Table 1). Thus, p63 safeguards epithelial lineage commitment, not only directly activating epidermal genes, but also indirectly repressing nonepithelial genes.
TP63 establishes a squamous subtype-specific enhancer landscape in SCCs
Chromatin states are closely associated with tumorigenesis and tumor phenotypes in SCCs [42]. There are dramatic epigenetic changes between SCC and its healthy counterpart, with gain or loss of H3K27ac occupancy at various chromatin regions [80]. As a lineage-survival oncogene in SCCs, ΔNp63α is exploited by tumor cells to establish squamous-specific chromatin architecture, which favors the development of SCC [81, 82]. ΔNp63α is highly expressed and predicts unfavorable overall prognosis in squamous subtype of pancreatic cancer (PC) or pancreatic ductal adenocarcinoma (PDA) [81, 83]. ΔNp63α selectively exerts an oncogenic role in squamous-like PC [81, 82]. H3K27ac profiling demonstrates that ΔNp63α-expressing PDAs display a distinct enhancer landscape resembling the squamous cell lineage, whereas ΔNp63α occupancy in squamous-like PC cells mainly distributes on active enhancers marked with H3K27ac and open chromatin regions, rather than transcriptional start sites. Loss of ΔNp63α in squamous-like BxPC3 PDA cells selectively reduces H3K27ac at p63-bound squamous elements, whereas it exerts no effect on H3K27ac at the control regions [81]. Remarkably, a large set of ΔNp63α-occupied regions are characterized as super enhancers (SE) [82], which are believed to drive oncogene expression in multiple cancers [84, 85]. H3K27ac occupancy at SEs associated with FAT2 and NECTIN1 is significantly reduced upon depletion of ΔNp63α in squamous-like PC cells, as well as downregulation of FAT2 and NECTIN1 [82]. This indicates that SE regions are associated with the squamous subtype and display a high dependence on ΔNp63α. Nasopharyngeal carcinoma (NPC) is a unique subtype of head and neck cancer [86, 87]. ΔNp63α is the primary isoform of TP63 expressed in NPC cells [21, 88, 89]. Our works recognized that ΔNp63α is enriched in SEs associated with oncogenes in NPC, including EGFR, CD44, etc.[90, 91]. Overexpression of ΔNp63α led to preferential expression of basal cell-specific proteins, including basal-type keratins, in NPC, through establishing basal-specific SEs [90]. In addition to protein-coding genes, ΔNp63α also activates transcription of a number of long non-coding RNAs, including CCAT1 [92] and LINC01503 [93], through establishment of squamous-specific SEs in SCCs. Consequently, overexpression of CCAT1 or LINC01503 promotes progression of SCCs [92,93,94]. New findings suggest that phase-separated condensates of TFs and coactivators play a role in the formation of SE [95, 96]. It has been demonstrated that activation domains of diverse TFs facilitate formation of phase-separated condensates with coactivators to drive gene transcription [97]. SEs are sensitive to chemical disruption of phase separation by 1,6-hexanediol (1,6-HD) [96, 98]. It is proposed that phase separation could be a target of next-generation drugs for chromatin biology [99]. Thus, enrichment of ΔNp63α in SEs associated with oncogenes provides an opportunity to target SCC cells by disrupting phase separation.
It has been demonstrated that ΔNp63α physically interacts with SOX2 in SCC cells [100], which is a lineage-survival oncogene frequently amplified in SCC from various organ sites [101,102,103,104]. Notably, ChIP-seq analysis revealed that SOX2 and p63 co-ordinate to occupy a large set of distal enhancers in SCC cells, but co-occupancy of SOX2 and p63 is rare in embryonic stem (ES) cells, which is in line with the absence of p63 in ES cells. Depletion p63 attenuates SOX2 enrichment at the enhancer of target genes, such as ETV4, which is an oncogene co-regulated by these two proteins. However, loss of p63 exerts little effect on other SOX2-occupied regions without p63 binding [100]. Thus, this study indicates that p63 might act as a pioneer factor to establish open chromatin architecture to facilitate access by other lineage-survival factors, such as SOX2, to bind and activate target gene transcription during SCC development. Recently, it has been demonstrated that p63 cooperates with SOX2 to activate intronic enhancer cluster of GLUT1 (SLC2A1), which in turn facilitates GLUT1-mediated glucose influx and generation of NADPH and GSH in SCCs [28]. The p63 protein also binds to the enhancer region of ZNF185 and promotes its expression in keratinocyte during epithelial differentiation. However, ZNF185 is downregulated in HNSCC, esophageal and cervical SCC even though p63 is commonly amplified in these cancers [27]. It is not clear why ZNF185 is decreased in SCC, and may be due to loss of function mutations on coactivators bound to p63, which results in a decrease of enhancer activity of ZNF185 in SCC cells. In contrast, the upstream enhancer of FANCD2 is inactive in primary keratinocytes, but aberrantly activated by ΔNp63α in SCC [32].
Mechanisms of ΔNp63α mediated chromatin remodeling and enhancer formation
There are multiple epigenetic modulators implicated in regulating enhancer and spatiotemporal transcription of lineage-specific genes, including enzymatic regulators histone methylases, histone acetyltransferases, histone deacetylases, polycomb repressive complex (PRC), DNA methylase and a number of chromatin remodelers. The p63 activates enhancers and increases chromatin accessibility through interaction with these various epigenetic modulators, which will be discussed in the section below.
Complex of proteins associated with set1-like complex (COMPASS)
COMPASS is a multiple-protein complex, which exerts histone H3K4 methylase activity. In mammalian cells, incorporation of H3K4me1 at enhancers is mainly accomplished by histone lysine methyltransferase 2C (KMT2C/MLL3) and 2D (KMT2D/MLL4) COMPASS-related complexes [105]. UTX is a putative demethylase within the MLL3/4 COMPASS-like complex, which erases H3K27me2/3 and facilitates CBP/p300-mediated H3K27ac deposition at poised enhancers [106]. It has been demonstrated that KMT2D interacts with p63 on chromatin and occupies p63-bound enhancers (Fig. 3). Specifically, KMT2D physically interacts with the α, β, and γ isoforms of ΔNp63, indicating that the C-terminal SAM or the TA domain is not necessary for this interaction. Depletion of KMT2D in keratinocytes reduces H3K4me1 and H3K27ac at p63-binding enhancers, as well as repressing p63 target genes. Bioinformatic analysis demonstrates that KMT2D-dependent p63 target genes that lose enhancer histone modifications are enriched for development and differentiation [107]. Functionally, keratinocytes with a KMT2D deficiency display premature and highly disorganized morphologies [107]. Although a high level of protein expression was achieved, overexpression of p63 in KMT2D-depleted keratinocytes was unable to recapitulate differentiation gene expression patterns [107]. This study indicates that recruitment of KMT2D by p63 is a prerequisite to establish the active enhancer of its target genes. Given that KMT2D is broadly implicated in enhancer formation and transcription regulation, it is proposed that KMT2D might affect epidermal differentiation through multiple TFs, rather than solely through p63. KMT2D is frequently mutated in human cancers, including SCC from various organ origins [108,109,110], but it is not clear whether p63-KMT2D interaction occurs in cancer cells, or whether this might be the case for other histone lysine methyltransferase members.
Histone acetyltransferases p300
Deposition of H3K27ac at the H3K4me1-defined enhancer is proposed to be catalyzed by the cAMP response element-binding protein (CBP) or p300 histone acetyltransferases [111]. Transcriptional co-activator p300 interacts with the N-terminal domain of p63γ and stimulates p63γ transcriptional activity in an acetylase-dependent manner [112]. The p300 also physically interacts with ΔNp63α and catalyzes its acetylation on lysine 193 (K193), leading to enhanced ΔNp63α protein stability [113]. A new finding demonstrated that the C-terminal domain of p63α interacts with p300 to activate β-catenin [114]. Inhibition of p300 activity suppresses HNSCC tumor growth [115, 116]. Thus, we propose that ΔNp63α might recruit p300/CBP to its bound regions to deposit H3K27ac at H3K4me1-defined enhancer (Fig. 3). Paradoxically, loss of function mutations of CBP/p300 are frequent in cutaneous SCC, suggesting a tumor-suppressor role. Loss of CBP/p300 in mouse keratinocytes exacerbates HrasS35-induced skin tumorigenesis [117]. Thus, it cannot be ruled out that other histone acetyltransferase members mediate ΔNp63α-dependent enhancer formation in SCC; this may be the case with general control nonrepressed protein 5 (GCN5), which is a histone acetyltransferase upregulated in ESCC [118]. It has been demonstrated that GCN5 cooperates with chromatin remodeler Switch/sucrose nonfermentable (SWI/SNF) to activate gene transcription [119, 120]. Furthermore, recruitment of SWI/SNF by GCN5 plays an essential role in the DNA damage response to double-strand breaks [121]. This suggests that the interaction between histone acetyltransferase members and the SWF/SNF complex may contribute to epigenomic reprogramming in SCC development.
SWI/SNF/Baf complex
The mammalian SWI/SNF complex is an ATP-dependent chromatin remodeling and histone acetylation complex highly expressed in pluripotent stem cells [122]. The mSWI/SNF or BAF complex contains one catalytic subunit (Brg1 or Brm) and 14 regulatory subunits [123, 124]. It has been shown that SWI/SNF complexes interact with p300 to modulate H3K27ac and are essential for the maintenance of lineage-specific enhancers [125]. The BAF complex is required to maintain epidermal cell type-specific open chromatin sites. BAF complex and p63 protein in keratinocyte are physically close to each other (Fig. 3). A large number of open chromatin regions in epidermal cells are co-occupied by BAF and p63. BAF depletion selectively reduces the chromatin accessibility at p63 motif regions, but not at motif regions of other factors, such as CTCF and KLF4, suggesting that BAF is required for maintaining the open status of p63-bound chromatin regions. Loss of p63 selectively attenuates BAF binding to p63 regulatory elements, without affecting BAF binding to the motif of CTCF and KLF4 [51]. These data strongly suggest that ΔNp63α recruits the BAF complex to p63-bound chromatin regions, which in turn establishes an open chromatin landscape in epidermal cells. Brg1 is required for nuclear internalization of the epidermal differentiation complex (EDC) locus, which is a ~ 3.1 Mb gene-rich region of mouse chromosome 3 encoding multiple genes essential for epidermal stratification and barrier formation [126]. During development of the epidermis, EDC is relocated from the nuclear membrane toward the nuclear interior, accompanied by a developmentally linked increase in the transcriptional activity of genes within the EDC locus. It has been shown that p63 directly activates Brg1 transcription. Nuclear internalization of the EDC locus is markedly impaired in the skin epithelium of p63−/− mice and is accompanied by insufficient expression of EDC genes in epidermal progenitor cells [126]. The specialized adenine and thymine-rich binding protein 1 (Satb1) is another direct target of p63 implicated in remodeling chromatin architecture of the EDC locus. Satb1 ablation, strikingly, expands the length of the whole EDC locus and its central domain, accompanied by insufficient expression of epidermis-specific genes and epidermal morphology alterations similar to p63 deficiency in skin [127]. Thus, these studies uncover an essential role of p63 in nuclear positioning of EDC locus and regulating higher-order chromatin structure of EDC through its direct target Brg1 [126] and Satb1 [127], respectively.
DNA methyltransferase 3A (Dnmt3a) and methylcytosine dioxygenase TET2
In addition to histone modifications, the status of DNA methylation also influences the activity of enhancers. Early studies suggest that patterns of 5-methylcytosine can mark enhancers. It is widely appreciated that lineage-specific enhancer regions are usually hypomethylated [128,129,130,131]. It has been demonstrated that genomic 5-hydroxymethylcytosine (5-hmC) signals are distinctly enriched at enhancers marked with H3K4me1 in human and mouse embryonic stem cells [132, 133], suggesting a potential role in the regulation of enhancer formation. 5-hmC is enriched at the poised enhancers in mouse embryonic stem cells, rather than active enhancers with H3K27ac marks [134]. However, another study indicates that eRNA-producing enhancers in mouse embryonic stem cells are usually marked with H3K27ac, decreased DNA methylation, and are enriched in the DNA hydroxylase Tet1 [135]. In pancreatic cancer cells, 5-hmC-enriched loci specifically overlap with H3K4me1 marked enhancers and open regions of chromatin. Gain of 5-hmC is correlated with upregulation of the cognate transcripts known to be important to cancer development [136]. DNA methylation in the human genome is catalyzed by DNA methyltransferase (DNMT), including Dnmt1, Dnmt3a, and Dnmt3b. Dnmt3a and Dnmt3b catalyze de novo methylation, whereas Dnmt1 is responsible for the maintenance of genome methylation patterns [137]. Three Tet proteins catalyze DNA hydroxymethylation [138,139,140] and are required for eRNA production [135]. It has been demonstrated that Dnmt3a and Dnmt3b bind to active enhancers in a histone H3K36me3-dependent manner in human EpSCs. Strikingly, Dnmt3a predominantly locates at the center of its target enhancers, whereas Dnmt3b peaks broadly distribute at the enhancer center and body. Both Dnmt3a and Dnmt3b promote enhancer activity and favor binding to SEs rather than typical enhancers. The center of enhancers bound by Dnmt3a exhibit high levels of 5-hmC. Silencing Dnmt3a reduces the levels of 5-hmC at its target enhancers, whereas silencing Tet2 specifically restores the level of DNA methylation at Dnmt3a-bound enhancers, indicating that Dnmt3a-bound enhancers are sequentially methylated by Dnmt3a and then are hydroxymethylated by Tet2 to license their active state. Remarkably, Dnmt3a, not Dnmt3b, physically interacts with p63 in EpSCs [75]. Co-occupancy by p63 was seen in approximately 50% of the Dnmt3a-bound enhancers (Fig. 3), which associate with the expression of genes implicated in keratinocyte proliferation and cellular identity specification. Notably, depletion of p63 reduces Dnmt3a localization to its target enhancers [75]. During epidermal differentiation, the genomic localization of Dnmt3a shifts from the enhancers closest to genes involved in stem cell proliferation to those that regulate genes involved in differentiation [75]. We speculate that p63 sequentially recruits Dnmt3a and Tet2 localization to the center of its target enhancers, establishing high levels of 5-hmC at those sites, which facilitates subsequent deposition of H3K4me1 and H3K27ac modifications and eRNA transcription (Fig. 3). It is worth noting that ΔNp63α binds to Dnmt3a, HDAC9, and KDM4C in cisplatin-resistant SCC-11 cells, suggesting that ΔNp63α may recruit these enzymes to shape the epigenome of SCC cells during acquisition of chemoresistance [141].
Mechanisms underlying ΔNp63α mediated transcription repression
Not only does ΔNp63α act as a dominant negative competitor to repress p53-mediated transcription [11], but it also exerts a repressive effect on transcription through interaction with diverse epigenetic regulators, which have been discussed in detail below.
HDAC1 and HDAC2
By using a tandem affinity purification strategy, Ramsey et al. observed that ΔNp63α interacts with histone deacetylase (HDAC) 1 and HDAC2 via its transactivation inhibitory domain in SCC cells. ΔNp63α recruits HDAC1 binding to promoter region of PUMA and represses its transcription. Treatment with the HDAC inhibitor, trichostatin A (TSA), alleviates repression of PUMA transcription by ΔNp63α and induces substantial apoptotic cell death in SCC cells. TSA treatment increases H4 acetylation levels at the p63-binding site within the PUMA promoter, whereas silencing ΔNp63α promotes deposition of H4 acetylation level at PUMA promoter and induces PUMA expression, indicating that ΔNp63α represses gene transcription through HDAC1-mediated histone deacetylation (Fig. 4a) [142]. An earlier study showed that deletion of ectodermal Hdac1 and Hdac2 leads to severe defects in hair follicle specification and epidermal proliferation and stratification, which is similar to phenotypes of p63-deficient mice [143]. Ectodermal Hdac1/2 deficiency activates the expression of targets of p63-mediated repression, rather than affecting expression of p63 positively regulated basal cell targets [143]. HDAC complexes can bind both promoter and enhancer regions of target genes [144]. While there are no studies demonstrating co-occupancy of p63 and HDAC1/2 complex on enhancers, we speculate that removal of H3K27ac by HDAC1/2 might be implicated in ΔNp63α-mediated repression of specific enhancers (Fig. 4a), since loss of ΔNp63α also results in unexpected gain of enhancer activity at RUNX1-enriched regions [77]. It has been demonstrated that the balance of EP300 and HDAC1 activities controls nucleosome eviction by the BRG1-containing SWI/SNF complex, which in turn affects transcription of DNA repair enzymes in macrophages [145]. Furthermore, recruitment of BRG1 by HDAC2 contributes to the transcriptional repression function of the SWF/SNF complex [146, 147]. Thus, it is proposed that the interaction between HDACs and the chromatin-remodeling SWF/SNF complex may be involved in ΔNp63α-mediated transcription repression.
Epigenetic mechanisms underlying p63 mediated transcription repression. a ΔNp63α physically interacts with HDAC1/2 to remove H3K27ac at enhancer and promoter of target genes, resulting in chromatin compaction and transcriptional repression. b ΔNp63α normally cooperates with the SNF/SWF complex to activate gene transcription, whereas the interaction between ΔNp63α and ACTL6A prevents the SNF/SWF complex from binding to target genes, leading to repression of growth inhibitory gene WWC1. c ΔNp63α physically interacts with Cbx4, a component of the PRC1 complex, to repress the transcription of nonepidermal lineage genes. d ΔNp63α promotes incorporation of histone variant HA2.Z through recruiting SRCAP to p63-binding sites at tumor-suppressor genes. e ΔNp63α activates Lsh expression and cooperates with Ras to bypass OIS through repressing p21WAF1/Cip1
ACTL6A/Baf53a
ACTL6A/BAF53a is a regulatory subunit of the SWI/SNF complex implicated in embryonic stem cells pluripotency and epidermal progenitor cells state [148,149,150]. An early study shows Baf53 forms a distinct histone acetyltransferase complex and acts as a cofactor for c-myc in oncogenic transformation [151]. High expression of BAF53a blocks myogenic differentiation and promotes cell proliferation in rhabdomyosarcoma [152], suggesting that ACTL6A/BAF53a is implicated in cancer development. The p63 and ACTL6A proteins are rarely co-expressed in normal epidermis, but are co-amplified in a subset of HNSCC (~ 19%) [153]. ACTL6A and p63 physically interact, cooperatively controlling a transcriptional program that promotes proliferation and suppresses differentiation, in part through activation of the Hippo–YAP pathway via regulators including WWC1 (Fig. 4b). Thus, ACTL6A and p63 collaborate as oncogenic drivers in HNSCC [153]. ACTL6a suppresses epidermal differentiation of adult stem cells by preventing the SWI/SNF complex from binding to genes controlling differentiation [148]. Given that more than 20% of human malignancies carry mutations in the mSWI/SNF complex [154, 155], overexpression of ACTL6A in SCCs may severely disrupt the epidermal differentiation program controlled by the p63-mSWI/SNF complex.
Polycomb repressive complex 1 component Cbx4
Cbx4 belongs to PRC1, which is one of the two complexes formed by polycomb chromatin-remodeling proteins that exert a repressive role on gene transcription via compacting chromatin [156]. It has been shown that Cbx4 physically interacts with p63 in thymic epithelial cells and plays an essential role in in thymic organogenesis [157]. Cbx4 is a direct target of p63 and is upregulated in keratinocytes during epithelial stratification. Ablation impairs epidermal progenitor cell proliferation and terminal differentiation [79]. Mechanistically, Cbx4 represses nonepidermal lineage gene expression via interacting with H3K27me3 that is deposited by EZH2 from the PRC1 complex and promoting ubiquitination of H2AK119 at these sites [79]. As a consequence, ubiquitination of H2AK119 is required for efficient repression of target genes by the PRC1 complex [158]. Restoration of Cbx4 partially rescues the epidermal phenotype in embryonic skin explants upon p63 ablation [79]. Thus, p63 represses the nonepidermal lineage transcription program during keratinocyte development via PRC1-mediated transcriptional repression (Fig. 4c). Cbx4 is frequently amplified and plays an oncogenic role in esophageal SCC [159], suggesting that an interaction between p63 and Cbx4 might also contribute to SCC development. A new study highlights the cooperation between mSWI/SNF and PRC1 that ensures meiotic progression in spermatocytes [160]. Furthermore, the SWI/SNF complex has been demonstrated to mediate eviction of PRC1 and PRC2 from the INK4b-ARF-INK4a locus [161]. Thus, the close interconnectivity of diverse chromatin modulators may be involved in ΔNp63α-regulated dynamics of chromatin in cancer cells.
SRCAP chromatin regulatory complex
H2A.Z is a histone variant usually incorporated into chromatin associated with gene promoters and enhancers. SRCAP (SWR1) is the major chromatin-remodeling complex responsible for catalyzing the incorporation of H2AZ-H2B dimers into nucleosomes [162, 163]. Study revealed that ΔNp63α physically interacts with and recruits SRCAP chromatin-remodeling complex subunits, including DMAP1, RUVBL1, and RUVBL2 [164]. Recruitment of the SRCAP complex by ΔNp63α facilitates H2A.Z deposition at ΔNp63α target loci and represses its transcription (Fig. 4d). Silencing SRCAP subunits or H2A.Z specifically induces expression of ΔNp63α-repressed genes. Among those repressive target genes, SAMD9L was identified as a crucial proliferation-suppressive gene for SCC development, whose depletion alleviating proliferation arrest was caused by loss of ΔNp63α [164].
Lymphoid-specific helicase (LSH)
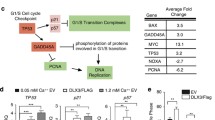
Lsh is an ATPase that belongs to the SNF2/helicases family involved in chromatin remodeling [165]. Bypassing oncogene-induced senescence (OIS) is a prerequisite step for malignant transformation [166]. A pilot study found that inducible p63 deficiency in the adult epidermis leads to cellular senescence and accelerated aging [167]. It has been shown that ΔNp63α expression in primary keratinocytes is downregulated in OIS induced by oncogenic H-Ras-V12. Forced expression of wild-type ΔNp63α, rather than TAp63a or ΔNp63αR279H mutant causing EEC syndrome, is sufficient to bypass OIS in Ras-expressing primary keratinocytes, indicating downregulation of ΔNp63α is required for OIS. Mechanistically, ΔNp63α cooperates with Ras to promote proliferation and survival of keratin 15-expressing stem cells. Lsh was identified as a direct target of ΔNp63α, which initiates senescence bypass, as silencing of the gene leads to a stronger senescent-like morphology [168]. Both p63 and Lsh are robustly detected in proliferating cells in DMBA-induced premalignant papillomas and HNSCC samples [168]. In addition to previously known functions regulating DNA methylation level at repeat elements and transcriptional silencing [169, 170], Lsh was recently recognized to regulate accessibility and H3 occupancy at a subset of putative enhancers associated with cellular identity, independent of DNA methylation [171, 172]. Lsh is highly expressed in various human cancers and drives cancer development [173,174,175]. It is proposed that Lsh may physically interact with ΔNp63α to regulate nucleosome positioning and H3 occupancy at cancer-associated genes (Fig. 4e).
Regulation of p63 in SCC
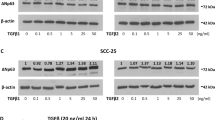
The N6-methyladenosine (m6A) modification, which is added by methyltransferase-like 3 (METTL3), promotes the increased expression of ΔNp63α mRNA and accelerates proliferation of cutaneous SCC [176]. There are data to support the hypothesis that it does so via stabilization of the ΔNp63α mRNA transcript [177]. Along with transcriptional and post-transcriptional control, TP63 is post-translationally regulated (Table 2). Nucleoporin 62 (NUP62), a component of nuclear pore complex, interacts with ΔNp63α and promotes ΔNp63α nuclear import and ΔNp63α-dependent target gene expression. In contrast, activated ROCK1 abolishes NUP62–ΔNp63α interaction and ΔNp63α nuclear accumulates in SCC cells through phosphorylating the FG domain of NUP62 [178]. Ubiquitin–proteasome system regulated p63 protein stability is intensively studied. To date, at least three E3 ligases, including anaphase-promoting complex/cyclosome (APC/C) complex, Itch (HECT, homologous E6-AP carboxyl terminus), and WWP1, have been recognized as mediators of p63 degradation in different contexts [179]. The ΔNp63α protein is preferentially degraded during early mitosis by the APC/C complex. ΔNp63α physically associates with the 29 amino acids at the C-terminal of Cdc20, which is an important subunit for APC/C activation. Ablation of Cdc20 substantially increases the half-life of the ΔNp63α protein in proliferating keratinocytes. However, in differentiating keratinocytes, Cdh1-APC/C is required for degradation of ΔNp63α protein and maintaining terminal differentiation capacity [180]. Syntaxin binding protein 4 (Stxbp4) is a PDZ domain containing adapter protein that associates with ΔNp63α [181, 182]. It has been demonstrated that Stxbp4 prevents degradation of ΔNp63α by the Cdc20-APC/C complex and suppresses epidermis differentiation [180]. High expression of Stxbp4 protein correlates with ΔNp63α protein accumulation in both skin SCC [180] and lung SCC [182], and predicts unfavorable overall survival (OS) and progression-free survival (PFS). Itch is a Nedd4-like ubiquitin protein E3 ligase belonging to the C2-WW-HECT type E3 family, which includes the members NEDD4, WWP1, SMURF1, and SMURF2 [183, 184]. Itch associates with both TAp63α and ΔNp63α via its WW2 domain, but preferentially to the latter [185]. Co-expression of Itch together with either TAp63α or ΔNp63α leads to increase of p63 ubiquitination, whereas a catalytically inactive Itch mutant (C830A mutant) still binds to TAp63α or ΔNp63α but fails to induce its ubiquitination. The ΔNp63α protein level increases in Itch knockout (KO) primary keratinocytes, whereas reintroduction of wild-type Itch in Itch KO cells leads to reduction of endogenous ΔNp63α protein level. The conserved threonine (T538) phosphorylation of a (T/S)P motif near the PY motif of p63 stabilizes the binding of PPPPY motif with the WW2 domain of Itch, facilitating subsequent ubiquitination and degradation [185]. However, a post-phosphorylation conformation switch from trans to cis catalyzed by Pin1, a peptidyl-prolyl isomerase, prevents the binding of p63 with Itch, thus stabilizing p63 protein [186]. These studies suggested the binding of p63 with WW domain containing E3 ligases is regulated by a Pin1-mediated cis/trans conformation switch, since E3 ligases favor the trans conformation [187]. Stxbp4 also inhibits Itch-mediated ΔNp63α degradation. However, silencing endogenous Itch fails to stabilize ΔNp63α protein upon loss of Stxbp4 in keratinocytes, whereas silencing of the receptor for protein kinase C1 (RACK1) partially rescues ΔNp63α degradation triggered by Stxbp4 deficiency [181]. This suggests that different E3 ligases control p63 protein turnover in a context-dependent manner.
The WW domain containing E3 ubiquitin protein ligase 1 (WWP1) is a member of HECT type E3 family. WWP1 physically associates with both TAp63α and ΔNp63α via PY/WW motif interaction, leading to ubiquitination and degradation of TAp63α and ΔNp63α proteins [188]. Phosphorylation of the conserved threonine (T538) close to the PxY motif of p63α is critical for Itch-p63α binding [186], but is not required for WWP1 mediated p63α degradation, as T538A substitution fails to affect WWP1-mediated proteasomal degradation [189]. Pin1 directly binds to p63α, but not the γ isotypes, and prevents p63α degradation by E3 ligase WWP1 [189]. A recent study revealed that WWP1 contributes to metformin-induced ΔNp63α degradation in HNSCC FaDu cells [190].
RACK1 is a WD-40 repeat-containing scaffold protein binding to p63α [181, 191]. Upon DNA damage, ΔNp63α protein is exported from the nucleus and into the cytoplasm, where it is recognized by RACK1. RACK1 binds the C-terminal SAM of p63 via its two WD40 repeats in C-terminus and promotes p63 proteasomal degradation [181, 191]. RACK1-dependent degradation of the ΔNp63α protein can be inhibited by Stxbp4 [181]. Upon DNA damage, stratifin physically interacts with ΔNp63α at its SAM motif and facilitates its nuclear export into the cytoplasm, and then facilitates degradation of ΔNp63α, mediated by RACK1 [191].
Conclusions and perspectives
Squamous cell carcinoma remains a great challenge to human health due to its highly aggressive and limited therapeutic options. SCC from various organs is highly dependent on ΔNp63α, which is recently highlighted for its pioneer activity and ability to establishing active enhancers through interaction with epigenetic regulators. The existence of p63 in chromatin-remodeling complexes provides an opportunity to target p63-positive SCCs via epigenetic therapy. However, many of the interaction partners of p63 in keratinocytes, including KMT2D and SWI/SNF components, are frequently mutated in SCC, implying that the p63-mediated differentiation transcriptional program is disturbed during SCC. It remains unclear how ΔNp63α cooperates with epigenetic regulators to activate enhancer activity of oncogenes in SCC. Additional efforts are necessary to fully understand the complexity of the interactions between ΔNp63α and epigenetic regulators in the control of SCC development. Characterization of ΔNp63α-binding partners in SCCs may enable the development novel strategies to indirectly interfere with ΔNp63α activity. Disruption of the interaction between ΔNp63α and diverse coactivators in SCC cells will broadly affect ΔNp63α-dependent transcription and may provide a novel therapeutic approach for SCC. ΔNp63α expression is sufficient to establish a squamous subtype-specific SE to drive oncogene expression in SCC. Thus, another area worthy of attention is the mechanisms underlying phase separation of ΔNp63α-dependent SE in SCC cells. It has been shown that ΔNp63α interacts with the c-Rel subunit of nuclear factor-kappa B (NF-κB) in keratinocytes and HNSCC cells [192]. Given the central role of NF-κB in the immune system and cancer development, the cooperation between ΔNp63α and c-Rel may influence the tumor microenvironment [193]. Immunosuppressive tumor microenvironment facilitates SCC development, progression, and therapeutic resistance [194, 195]. A substantial proportion of SCC patients do not respond to immune checkpoint blockade therapy, including PD-1/PD-L1 mAb treatment [2, 196, 197]. To overcome barriers to effective immunotherapy, we believe that clarification is needed concerning how ΔNp63α cooperates with NF-κB and other immune-regulatory factors to alter the immune environment of SCC.
Change history
02 November 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00018-021-03982-9
Abbreviations
- SCC:
-
Squamous cell carcinoma
- HNSCC:
-
Head and neck squamous cell carcinoma
- TF:
-
Transcription factor
- ECM:
-
Extracellular matrix
- iPSCs:
-
Induced pluripotent stem cells
- RA:
-
Retinoic acid
- BMP4:
-
Bone morphogenetic protein 4
- hESCs:
-
Human embryonic stem cells
- KLF4:
-
Kruppel-like factor 4
- eRNAs:
-
Enhancer RNAs
- EpSCs:
-
Epidermal stem cells
- ChIP:
-
Chromatin immunoprecipitation
- PRC:
-
Polycomb repressive complex 1
- PC:
-
Pancreatic cancer
- PDA:
-
Pancreatic ductal adenocarcinoma
- SE:
-
Super enhancer
- COMPASS:
-
Complex of proteins associated with set1-like complex
- CBP:
-
CAMP response element-binding protein
- GCN5:
-
General control nonrepressed protein 5
- SWI/SNF:
-
Switch/sucrose nonfermentable
- EDC:
-
Epidermal differentiation complex
- Satb1:
-
Specialized adenine and thymine-rich binding protein 1
- Dnmt3a:
-
DNA methyltransferase 3A
- 5-hmC:
-
5-Hydroxymethylcytosine
- DNMT:
-
DNA methyltransferase
- LSH:
-
Lymphoid-specific helicase
- METTL3:
-
Methyltransferase-like 3
- NUP62:
-
Nucleoporin 62
- APC/C:
-
Anaphase-promoting complex/cyclosome
- RACK1:
-
Receptor for protein kinase C1
- WWP1:
-
WW domain containing E3 ubiquitin protein ligase 1
References
Sanchez-Danes A, Blanpain C (2018) Deciphering the cells of origin of squamous cell carcinomas. Nat Rev Cancer 18(9):549–561. https://doi.org/10.1038/s41568-018-0024-5
Dotto GP, Rustgi AK (2016) Squamous cell cancers: a unified perspective on biology and genetics. Cancer Cell 29(5):622–637. https://doi.org/10.1016/j.ccell.2016.04.004
Bailey P, Chang DK, Nones K, Johns AL, Patch AM, Gingras MC, Miller DK, Christ AN, Bruxner TJC, Quinn MC, Nourse C, Murtaugh LC, Harliwong I, Idrisoglu S, Manning S, Nourbakhsh E, Wani S, Fink L, Holmes O, Chin V, Anderson MJ, Kazakoff S, Leonard C, Newell F, Waddell N, Wood S, Xu QY, Wilson PJ, Cloonan N, Kassahn KS, Taylor D, Quek K, Robertson A, Pantano L, Mincarelli L, Sanchez LN, Evers L, Wu JM, Pinese M, Cowley MJ, Jones MD, Colvin EK, Nagrial AM, Humphrey ES, Chantrill LA, Mawson A, Humphris J, Chou A, Pajic M, Scarlett CJ, Pinho AV, Giry-Laterriere M, Rooman I, Samra JS, Kench JG, Lovell JA, Merrett ND, Toon CW, Epari K, Nguyen NQ, Barbour A, Zeps N, Moran-Jones K, Jamieson NB, Graham JS, Duthie F, Oien K, Hair J, Grutzmann R, Maitra A, Iacobuzio-Donahue CA, Wolfgang CL, Morgan RA, Lawlor RT, Corbo V, Bassi C, Rusev B, Capelli P, Salvia R, Tortora G, Mukhopadhyay D, Petersen GM, Munzy DM, Fisher WE, Karim SA, Eshleman JR, Hruban RH, Pilarsky C, Morton JR, Sansom OJ, Scarpa A, Musgrove EA, Bailey UMH, Hofmann O, Sutherland RL, Wheeler DA, Gill AJ, Gibbs RA, Pearson JV, Waddell N, Biankin AV, Grimmond SM, Genome APC (2016) Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531(7592):47. https://doi.org/10.1038/nature16965
Ntanasis-Stathopoulos I, Tsilimigras DI, Georgiadou D, Kanavidis P, Riccioni O, Salla C, Psaltopoulou T, Sergentanis TN (2017) Squamous cell carcinoma of the pancreas: a systematic review and pooled survival analysis. Eur J Cancer 79:193–204. https://doi.org/10.1016/j.ejca.2017.04.006
Moffitt RA, Marayati R, Flate EL, Volmar KE, Loeza SGH, Hoadley KA, Rashid NU, Williams LA, Eaton SC, Chung AH, Smyla JK, Anderson JM, Kim HJ, Bentrem DJ, Talamonti MS, Iacobuzio-Donahue CA, Hollingsworth MA, Yeh JJ (2015) Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat Genet 47(10):1168. https://doi.org/10.1038/ng.3398
Bernardo C, Eriksson P, Marzouka NA, Liedberg F, Sjodahl G, Hoglund M (2019) Molecular pathology of the luminal class of urothelial tumors. J Pathol. https://doi.org/10.1002/path.5318
Malik RD, Dakwar G, Hardee ME, Sanfilippo NJ, Rosenkrantz AB, Taneja SS (2011) Squamous cell carcinoma of the prostate. Rev Urol 13(1):56–60
Solomon B, Young RJ, Rischin D (2018) Head and neck squamous cell carcinoma: Genomics and emerging biomarkers for immunomodulatory cancer treatments. Semin Cancer Biol 52(Pt 2):228–240. https://doi.org/10.1016/j.semcancer.2018.01.008
Plzak J, Boucek J, Bandurova V, Kolar M, Hradilova M, Szabo P, Lacina L, Chovanec M, Smetana K Jr (2019) The head and neck squamous cell carcinoma microenvironment as a potential target for cancer therapy. Cancers. https://doi.org/10.3390/cancers11040440
Garraway LA, Sellers WR (2006) Lineage dependency and lineage-survival oncogenes in human cancer. Nat Rev Cancer 6(8):593–602. https://doi.org/10.1038/nrc1947
Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dotsch V, Andrews NC, Caput D, McKeon F (1998) p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol Cell 2(3):305–316
Duijf PH, Vanmolkot KR, Propping P, Friedl W, Krieger E, McKeon F, Dotsch V, Brunner HG, van Bokhoven H (2002) Gain-of-function mutation in ADULT syndrome reveals the presence of a second transactivation domain in p63. Hum Mol Genet 11(7):799–804
Mangiulli M, Valletti A, Caratozzolo MF, Tullo A, Sbisa E, Pesole G, D'Erchia AM (2009) Identification and functional characterization of two new transcriptional variants of the human p63 gene. Nucleic Acids Res 37(18):6092–6104. https://doi.org/10.1093/nar/gkp674
Serber Z, Lai HC, Yang A, Ou HD, Sigal MS, Kelly AE, Darimont BD, Duijf PH, Van Bokhoven H, McKeon F, Dotsch V (2002) A C-terminal inhibitory domain controls the activity of p63 by an intramolecular mechanism. Mol Cell Biol 22(24):8601–8611. https://doi.org/10.1128/mcb.22.24.8601-8611.2002
Wu G, Nomoto S, Hoque MO, Dracheva T, Osada M, Lee CC, Dong SM, Guo Z, Benoit N, Cohen Y, Rechthand P, Califano J, Moon CS, Ratovitski E, Jen J, Sidransky D, Trink B (2003) DeltaNp63alpha and TAp63alpha regulate transcription of genes with distinct biological functions in cancer and development. Cancer Res 63(10):2351–2357
Candi E, Rufini A, Terrinoni A, Dinsdale D, Ranalli M, Paradisi A, De Laurenzi V, Spagnoli LG, Catani MV, Ramadan S, Knight RA, Melino G (2006) Differential roles of p63 isoforms in epidermal development: selective genetic complementation in p63 null mice. Cell Death Differ 13(6):1037–1047. https://doi.org/10.1038/sj.cdd.4401926
Keyes WM, Vogel H, Koster MI, Guo XC, Qi Y, Petherbridge KM, Roop DR, Bradley A, Mills AA (2006) p63 heterozygous mutant mice are not prone to spontaneous or chemically induced tumors. Proc Natl Acad Sci USA 103(22):8435–8440. https://doi.org/10.1073/pnas.0602477103
Tordella L, Koch S, Salter V, Pagotto A, Doondeea JB, Feller SM, Ratnayaka I, Zhong S, Goldin RD, Lozano G, McKeon FD, Tavassoli M, Fritzsche F, Huber GF, Rossle M, Moch H, Lu X (2013) ASPP2 suppresses squamous cell carcinoma via RelA/p65-mediated repression of p63. Proc Natl Acad Sci USA 110(44):17969–17974. https://doi.org/10.1073/pnas.1309362110
Signoretti S, Waltregny D, Dilks J, Isaac B, Lin D, Garraway L, Yang A, Montironi R, McKeon F, Loda M (2000) p63 is a prostate basal cell marker and is required for prostate development. Am J Pathol 157(6):1769–1775. https://doi.org/10.1016/S0002-9440(10)64814-6
Cheng W, Jacobs WB, Zhang JJ, Moro A, Park JH, Kushida M, Qiu W, Mills AA, Kim PC (2006) DeltaNp63 plays an anti-apoptotic role in ventral bladder development. Development 133(23):4783–4792. https://doi.org/10.1242/dev.02621
Crook T, Nicholls JM, Brooks L, O'Nions J, Allday MJ (2000) High level expression of deltaN-p63: a mechanism for the inactivation of p53 in undifferentiated nasopharyngeal carcinoma (NPC)? Oncogene 19(30):3439–3444. https://doi.org/10.1038/sj.onc.1203656
Holcakova J, Nekulova M, Orzol P, Nenutil R, Podhorec J, Svoboda M, Dvorakova P, Pjechova M, Hernychova L, Vojtesek B, Coates PJ (2017) Delta Np63 activates EGFR signaling to induce loss of adhesion in triple-negative basal-like breast cancer cells. Breast Cancer Res Ther 163(3):475–484. https://doi.org/10.1007/s10549-017-4216-6
Ramsey MR, Wilson C, Ory B, Rothenberg SM, Faquin W, Mills AA, Ellisen LW (2013) FGFR2 signaling underlies p63 oncogenic function in squamous cell carcinoma. J Clin Investig 123(8):3525–3538. https://doi.org/10.1172/JCI68899
Compagnone M, Gatti V, Presutti D, Ruberti G, Fierro C, Markert EK, Vousden KH, Zhou H, Mauriello A, Anemone L, Bongiorno-Borbone L, Melino G, Peschiaroli A (2017) DeltaNp63-mediated regulation of hyaluronic acid metabolism and signaling supports HNSCC tumorigenesis. Proc Natl Acad Sci USA 114(50):13254–13259. https://doi.org/10.1073/pnas.1711777114
Carroll DK, Carroll JS, Leong CO, Cheng F, Brown M, Mills AA, Brugge JS, Ellisen LW (2006) p63 regulates an adhesion programme and cell survival in epithelial cells. Nat Cell Biol 8(6):551–561. https://doi.org/10.1038/ncb1420
Beaudry VG, Jiang D, Dusek RL, Park EJ, Knezevich S, Ridd K, Vogel H, Bastian BC, Attardi LD (2010) Loss of the p53/p63 regulated desmosomal protein Perp promotes tumorigenesis. PLoS Genet 6(10):e1001168. https://doi.org/10.1371/journal.pgen.1001168
Smirnov A, Lena AM, Cappello A, Panatta E, Anemona L, Bischetti S, Annicchiarico-Petruzzelli M, Mauriello A, Melino G, Candi E (2019) ZNF185 is a p63 target gene critical for epidermal differentiation and squamous cell carcinoma development. Oncogene 38(10):1625–1638. https://doi.org/10.1038/s41388-018-0509-4
Hsieh MH, Choe JH, Gadhvi J, Kim YJ, Arguez MA, Palmer M, Gerold H, Nowak C, Do H, Mazambani S, Knighton JK, Cha M, Goodwin J, Kang MK, Jeong JY, Lee SY, Faubert B, Xuan Z, Abel ED, Scafoglio C, Shackelford DB, Minna JD, Singh PK, Shulaev V, Bleris L, Hoyt K, Kim J, Inoue M, DeBerardinis RJ, Kim TH, Kim JW (2019) p63 and SOX2 dictate glucose reliance and metabolic vulnerabilities in squamous cell carcinomas. Cell Rep 28(7):1860–1878.e1869. https://doi.org/10.1016/j.celrep.2019.07.027
Viticchie G, Agostini M, Lena AM, Mancini M, Zhou H, Zolla L, Dinsdale D, Saintigny G, Melino G, Candi E (2015) p63 supports aerobic respiration through hexokinase II. Proc Natl Acad Sci USA 112(37):11577–11582. https://doi.org/10.1073/pnas.1508871112
Latina A, Viticchie G, Lena AM, Piro MC, Annicchiarico-Petruzzelli M, Melino G, Candi E (2016) DeltaNp63 targets cytoglobin to inhibit oxidative stress-induced apoptosis in keratinocytes and lung cancer. Oncogene 35(12):1493–1503. https://doi.org/10.1038/onc.2015.222
Arcidiacono P, Webb CM, Brooke MA, Zhou HQ, Delaney PJ, Ng KE, Blaydon DC, Tinker A, Kelsell DP, Chikh A (2018) p63 is a key regulator of iRHOM2 signalling in the keratinocyte stress response. Nat Commun. https://doi.org/10.1038/S41467-018-03470-Y
Bretz AC, Gittler MP, Charles JP, Gremke N, Eckhardt I, Mernberger M, Mandic R, Thomale J, Nist A, Wanzel M, Stiewe T (2016) DeltaNp63 activates the Fanconi anemia DNA repair pathway and limits the efficacy of cisplatin treatment in squamous cell carcinoma. Nucleic Acids Res 44(7):3204–3218. https://doi.org/10.1093/nar/gkw036
Lin YL, Sengupta S, Gurdziel K, Bell GW, Jacks T, Flores ER (2009) p63 and p73 transcriptionally regulate genes involved in DNA repair. PLoS Genet 5(10):e1000680. https://doi.org/10.1371/journal.pgen.1000680
Candi E, Rufini A, Terrinoni A, Giamboi-Miraglia A, Lena AM, Mantovani R, Knight R, Melino G (2007) Delta Np63 regulates thymic development through enhanced expression of FgfR2 and Jag2. Proc Natl Acad Sci USA 104(29):11999–12004. https://doi.org/10.1073/pnas.0703458104
Yugawa T, Narisawa-Saito M, Yoshimatsu Y, Haga K, Ohno S, Egawa N, Fujita M, Kiyono T (2010) DeltaNp63alpha repression of the Notch1 gene supports the proliferative capacity of normal human keratinocytes and cervical cancer cells. Cancer Res 70(10):4034–4044. https://doi.org/10.1158/0008-5472.CAN-09-4063
Lu H, Yang X, Duggal P, Allen CT, Yan B, Cohen J, Nottingham L, Romano RA, Sinha S, King KE, Weinberg WC, Chen Z, Van Waes C (2011) TNF-alpha promotes c-REL/DeltaNp63alpha interaction and TAp73 dissociation from key genes that mediate growth arrest and apoptosis in head and neck cancer. Cancer Res 71(21):6867–6877. https://doi.org/10.1158/0008-5472.CAN-11-2460
Yang X, Lu H, Yan B, Romano RA, Bian Y, Friedman J, Duggal P, Allen C, Chuang R, Ehsanian R, Si H, Sinha S, Van Waes C, Chen Z (2011) DeltaNp63 versatilely regulates a Broad NF-kappaB gene program and promotes squamous epithelial proliferation, migration, and inflammation. Cancer Res 71(10):3688–3700. https://doi.org/10.1158/0008-5472.CAN-10-3445
Bid HK, Roberts RD, Cam M, Audino A, Kurmasheva RT, Lin J, Houghton PJ, Cam H (2014) DeltaNp63 promotes pediatric neuroblastoma and osteosarcoma by regulating tumor angiogenesis. Cancer Res 74(1):320–329. https://doi.org/10.1158/0008-5472.CAN-13-0894
Ratovitski EA (2011) DeltaNp63alpha/IRF6 interplay activates NOS2 transcription and induces autophagy upon tobacco exposure. Arch Biochem Biophys 506(2):208–215. https://doi.org/10.1016/j.abb.2010.11.020
Grabocka E, Pylayeva-Gupta Y, Jones MJK, Lubkov V, Yennanaberhan E, Taylor L, Jeng HH, Bar-Sagi D (2014) Wild-type H- and N-Ras promote mutant K-Ras-driven tumorigenesis by modulating the DNA damage response. Cancer Cell 25(2):243–256. https://doi.org/10.1016/j.ccr.2014.01.005
Suarez-Carmona M, Hubert P, Gonzalez A, Duray A, Roncarati P, Erpicum C, Boniver J, Castronovo V, Noel A, Saussez S, Peulen O, Delvenne P, Herfs M (2014) DeltaNp63 isoform-mediated beta-defensin family up-regulation is associated with (lymph)angiogenesis and poor prognosis in patients with squamous cell carcinoma. Oncotarget 5(7):1856–1868. https://doi.org/10.18632/oncotarget.1819
Latil M, Nassar D, Beck B, Boumahdi S, Wang L, Brisebarre A, Dubois C, Nkusi E, Lenglez S, Checinska A, Vercauteren Drubbel A, Devos M, Declercq W, Yi R, Blanpain C (2017) Cell-type-specific chromatin states differentially prime squamous cell carcinoma tumor-initiating cells for epithelial to mesenchymal transition. Cell Stem Cell 20(2):191–204.e195. https://doi.org/10.1016/j.stem.2016.10.018
Higashikawa K, Yoneda S, Tobiume K, Taki M, Shigeishi H, Kamata N (2007) Snail-induced down-regulation of DeltaNp63alpha acquires invasive phenotype of human squamous cell carcinoma. Cancer Res 67(19):9207–9213. https://doi.org/10.1158/0008-5472.CAN-07-0932
Yoh KE, Regunath K, Guzman A, Lee SM, Pfister NT, Akanni O, Kaufman LJ, Prives C, Prywes R (2016) Repression of p63 and induction of EMT by mutant Ras in mammary epithelial cells. Proc Natl Acad Sci USA 113(41):E6107–E6116. https://doi.org/10.1073/pnas.1613417113
Hu L, Liang S, Chen H, Lv T, Wu J, Chen D, Wu M, Sun S, Zhang H, You H, Ji H, Zhang Y, Bergholz J, Xiao ZJ (2017) DeltaNp63alpha is a common inhibitory target in oncogenic PI3K/Ras/Her2-induced cell motility and tumor metastasis. Proc Natl Acad Sci USA 114(20):E3964–E3973. https://doi.org/10.1073/pnas.1617816114
Di Giacomo V, Tian TV, Mas A, Pecoraro M, Batlle-Morera L, Noya L, Martin-Caballero J, Ruberte J, Keyes WM (2017) Delta Np63 alpha promotes adhesion of metastatic prostate cancer cells to the bone through regulation of CD82. Oncogene 36(31):4381–4392. https://doi.org/10.1038/onc.2017.42
Helton ES, Zhu J, Chen X (2006) The unique NH2-terminally deleted (DeltaN) residues, the PXXP motif, and the PPXY motif are required for the transcriptional activity of the DeltaN variant of p63. J Biol Chem 281(5):2533–2542. https://doi.org/10.1074/jbc.M507964200
Ortt K, Raveh E, Gat U, Sinha S (2008) A chromatin immunoprecipitation screen in mouse keratinocytes reveals Runx1 as a direct transcriptional target of Delta Np63. J Cell Biochem 104(4):1204–1219. https://doi.org/10.1002/jcb.21700
Birkaya B, Ortt K, Sinha S (2007) Novel in vivo targets of DeltaNp63 in keratinocytes identified by a modified chromatin immunoprecipitation approach. BMC Mol Biol 8:43. https://doi.org/10.1186/1471-2199-8-43
Kouwenhoven EN, Oti M, Niehues H, van Heeringen SJ, Schalkwijk J, Stunnenberg HG, van Bokhoven H, Zhou H (2015) Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation. EMBO Rep 16(7):863–878. https://doi.org/10.15252/embr.201439941
Bao X, Rubin AJ, Qu K, Zhang J, Giresi PG, Chang HY, Khavari PA (2015) A novel ATAC-seq approach reveals lineage-specific reinforcement of the open chromatin landscape via cooperation between BAF and p63. Genome Biol 16:284. https://doi.org/10.1186/s13059-015-0840-9
Gatti V, Fierro C, Annicchiarico-Petruzzelli M, Melino G, Peschiaroli A (2019) DeltaNp63 in squamous cell carcinoma: defining the oncogenic routes affecting epigenetic landscape and tumour microenvironment. Mol Oncol 13(5):981–1001. https://doi.org/10.1002/1878-0261.12473
Celli J, Duijf P, Hamel BC, Bamshad M, Kramer B, Smits AP, Newbury-Ecob R, Hennekam RC, Van Buggenhout G, van Haeringen A, Woods CG, van Essen AJ, de Waal R, Vriend G, Haber DA, Yang A, McKeon F, Brunner HG, van Bokhoven H (1999) Heterozygous germline mutations in the p53 homolog p63 are the cause of EEC syndrome. Cell 99(2):143–153. https://doi.org/10.1016/s0092-8674(00)81646-3
Senoo M, Pinto F, Crum CP, McKeon F (2007) p63 is essential for the proliferative potential of stem cells in stratified epithelia. Cell 129(3):523–536. https://doi.org/10.1016/j.cell.2007.02.045
Yang A, Schweitzer R, Sun DQ, Kaghad M, Walker N, Bronson RT, Tabin C, Sharpe A, Caput D, Crum C, McKeon F (1999) p63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature 398(6729):714–718
Mills AA, Zheng B, Wang XJ, Vogel H, Roop DR, Bradley A (1999) p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature 398(6729):708–713. https://doi.org/10.1038/19531
Romano RA, Smalley K, Magraw C, Serna VA, Kurita T, Raghavan S, Sinha S (2012) Delta Np63 knockout mice reveal its indispensable role as a master regulator of epithelial development and differentiation. Development 139(4):772–782. https://doi.org/10.1242/dev.071191
Chen G, Gulbranson DR, Hou Z, Bolin JM, Ruotti V, Probasco MD, Smuga-Otto K, Howden SE, Diol NR, Propson NE, Wagner R, Lee GO, Antosiewicz-Bourget J, Teng JM, Thomson JA (2011) Chemically defined conditions for human iPSC derivation and culture. Nat Methods 8(5):424–429. https://doi.org/10.1038/nmeth.1593
Soares E, Xu Q, Li Q, Qu J, Zheng Y, Raeven HHM, Brandao KO, Petit I, van den Akker WMR, van Heeringen SJ, Aberdam D, Tang F, Zhou H (2019) Single-cell RNA-seq identifies a reversible mesodermal activation in abnormally specified epithelia of p63 EEC syndrome. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1908180116
Pattison JM, Melo SP, Piekos SN, Torkelson JL, Bashkirova E, Mumbach MR, Rajasingh C, Zhen HH, Li L, Liaw E, Alber D, Rubin AJ, Shankar G, Bao X, Chang HY, Khavari PA, Oro AE (2018) Retinoic acid and BMP4 cooperate with p63 to alter chromatin dynamics during surface epithelial commitment. Nat Genet. https://doi.org/10.1038/s41588-018-0263-0
Sammons MA, Zhu J, Drake AM, Berger SL (2015) TP53 engagement with the genome occurs in distinct local chromatin environments via pioneer factor activity. Genome Res 25(2):179–188. https://doi.org/10.1101/gr.181883.114
Rhie SK, Hazelett DJ, Coetzee SG, Yan CL, Noushmehr H, Coetzee GA (2014) Nucleosome positioning and histone modifications define relationships between regulatory elements and nearby gene expression in breast epithelial cells. BMC Genomics. https://doi.org/10.1186/1471-2164-15-331
Sethi I, Sinha S, Buck MJ (2014) Role of chromatin and transcriptional co-regulators in mediating p63-genome interactions in keratinocytes. BMC Genomics 15:1042. https://doi.org/10.1186/1471-2164-15-1042
Qu J, Yi G, Zhou H (2019) p63 cooperates with CTCF to modulate chromatin architecture in skin keratinocytes. Epigenetics Chromatin 12(1):31. https://doi.org/10.1186/s13072-019-0280-y
Fan X, Wang D, Burgmaier JE, Teng Y, Romano RA, Sinha S, Yi R (2018) Single cell and open chromatin analysis reveals molecular origin of epidermal cells of the skin. Dev Cell 47(1):21–37.e25. https://doi.org/10.1016/j.devcel.2018.08.010
Lin-Shiao E, Lan YM, Welzenbach J, Alexander KA, Zhang Z, Knapp M, Mangold E, Sammons M, Ludwig KU, Berger SL (2019) p63 establishes epithelial enhancers at critical craniofacial development genes. Sci Adv. https://doi.org/10.1126/sciadv.aaw0946
Santos-Pereira JM, Gallardo-Fuentes L, Neto A, Acemel RD, Tena JJ (2019) Pioneer and repressive functions of p63 during zebrafish embryonic ectoderm specification. Nat Commun 10(1):3049. https://doi.org/10.1038/s41467-019-11121-z
Li L, Wang Y, Torkelson JL, Shankar G, Pattison JM, Zhen HH, Fang F, Duren Z, Xin J, Gaddam S, Melo SP, Piekos SN, Li J, Liaw EJ, Chen L, Li R, Wernig M, Wong WH, Chang HY, Oro AE (2019) TFAP2C- and p63-dependent networks sequentially rearrange chromatin landscapes to drive human epidermal lineage commitment. Cell Stem Cell 24(2):271–284.e278. https://doi.org/10.1016/j.stem.2018.12.012
Chakravarti D, Su X, Cho MS, Bui NH, Coarfa C, Venkatanarayan A, Benham AL, Flores Gonzalez RE, Alana J, Xiao W, Leung ML, Vin H, Chan IL, Aquino A, Muller N, Wang H, Cooney AJ, Parker-Thornburg J, Tsai KY, Gunaratne PH, Flores ER (2014) Induced multipotency in adult keratinocytes through down-regulation of DeltaNp63 or DGCR8. Proc Natl Acad Sci USA 111(5):E572–581. https://doi.org/10.1073/pnas.1319743111
Smith E, Shilatifard A (2014) Enhancer biology and enhanceropathies. Nat Struct Mol Biol 21(3):210–219. https://doi.org/10.1038/nsmb.2784
Herz HM, Mohan M, Garruss AS, Liang K, Takahashi YH, Mickey K, Voets O, Verrijzer CP, Shilatifard A (2012) Enhancer-associated H3K4 monomethylation by Trithorax-related, the Drosophila homolog of mammalian Mll3/Mll4. Genes Dev 26(23):2604–2620. https://doi.org/10.1101/gad.201327.112
Shen Y, Yue F, McCleary DF, Ye Z, Edsall L, Kuan S, Wagner U, Dixon J, Lee L, Lobanenkov VV, Ren B (2012) A map of the cis-regulatory sequences in the mouse genome. Nature 488(7409):116–120. https://doi.org/10.1038/nature11243
Hon GC, Rajagopal N, Shen Y, McCleary DF, Yue F, Dang MD, Ren B (2013) Epigenetic memory at embryonic enhancers identified in DNA methylation maps from adult mouse tissues. Nat Genet 45(10):1198–1206. https://doi.org/10.1038/ng.2746
Kim TK, Shiekhattar R (2015) Architectural and functional commonalities between enhancers and promoters. Cell 162(5):948–959. https://doi.org/10.1016/j.cell.2015.08.008
Rinaldi L, Datta D, Serrat J, Morey L, Solanas G, Avgustinova A, Blanco E, Pons JI, Matallanas D, Von Kriegsheim A, Di Croce L, Benitah SA (2016) Dnmt3a and Dnmt3b associate with enhancers to regulate human epidermal stem cell homeostasis. Cell Stem Cell 19(4):491–501. https://doi.org/10.1016/j.stem.2016.06.020
Kouwenhoven EN, van Heeringen SJ, Tena JJ, Oti M, Dutilh BE, Alonso ME, de la Calle-Mustienes E, Smeenk L, Rinne T, Parsaulian L, Bolat E, Jurgelenaite R, Huynen MA, Hoischen A, Veltman JA, Brunner HG, Roscioli T, Oates E, Wilson M, Manzanares M, Gomez-Skarmeta JL, Stunnenberg HG, Lohrum M, van Bokhoven H, Zhou HQ (2010) Genome-wide profiling of p63 DNA-binding sites identifies an element that regulates gene expression during limb development in the 7q21 SHFM1 locus. PLoS Genet. https://doi.org/10.1371/journal.pgen.1001065
Qu J, Tanis SEJ, Smits JPH, Kouwenhoven EN, Oti M, van den Bogaard EH, Logie C, Stunnenberg HG, van Bokhoven H, Mulder KW, Zhou H (2018) Mutant p63 affects epidermal cell identity through rewiring the enhancer landscape. Cell Rep 25(12):3490–3503.e3494. https://doi.org/10.1016/j.celrep.2018.11.039
Shalom-Feuerstein R, Lena AM, Zhou H, De La Forest DS, Van Bokhoven H, Candi E, Melino G, Aberdam D (2011) DeltaNp63 is an ectodermal gatekeeper of epidermal morphogenesis. Cell Death Differ 18(5):887–896. https://doi.org/10.1038/cdd.2010.159
Mardaryev AN, Liu B, Rapisarda V, Poterlowicz K, Malashchuk I, Rudolf J, Sharov AA, Jahoda CA, Fessing MY, Benitah SA, Xu GL, Botchkarev VA (2016) Cbx4 maintains the epithelial lineage identity and cell proliferation in the developing stratified epithelium. J Cell Biol 212(1):77–89. https://doi.org/10.1083/jcb.201506065
Herz HM, Hu DQ, Shilatifard A (2014) Enhancer malfunction in cancer. Mol Cell 53(6):859–866. https://doi.org/10.1016/j.molcel.2014.02.033
Somerville TDD, Xu YL, Miyabayashi K, Tiriac H, Cleary CR, Maia-Silva D, Milazzo JP, Tuveson DA, Vakoc CR (2018) TP63-mediated enhancer reprogramming drives the squamous subtype of pancreatic ductal adenocarcinoma. Cell Rep 25(7):1741. https://doi.org/10.1016/j.celrep.2018.10.051
Hamdan FH, Johnsen SA (2018) DeltaNp63-dependent super enhancers define molecular identity in pancreatic cancer by an interconnected transcription factor network. Proc Natl Acad Sci USA 115(52):E12343–E12352. https://doi.org/10.1073/pnas.1812915116
Basturk O, Khanani F, Sarkar F, Levi E, Cheng JD, Adsay NV (2005) DeltaNp63 expression in pancreas and pancreatic neoplasia. Mod Pathol 18(9):1193–1198. https://doi.org/10.1038/modpathol.3800401
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-Andre V, Sigova AA, Hoke HA, Young RA (2013) Super-enhancers in the control of cell identity and disease. Cell 155(4):934–947. https://doi.org/10.1016/j.cell.2013.09.053
Sengupta S, George RE (2017) Super-enhancer-driven transcriptional dependencies in cancer. Trends Cancer 3(4):269–281. https://doi.org/10.1016/j.trecan.2017.03.006
Yi M, Cai J, Li J, Chen S, Zeng Z, Peng Q, Ban Y, Zhou Y, Li X, Xiong W, Li G, Xiang B (2018) Rediscovery of NF-kappaB signaling in nasopharyngeal carcinoma: how genetic defects of NF-kappaB pathway interplay with EBV in driving oncogenesis? J Cell Physiol 233(8):5537–5549. https://doi.org/10.1002/jcp.26410
Li J, Wang W, Chen S, Cai J, Ban Y, Peng Q, Zhou Y, Zeng Z, Li X, Xiong W, Li G, Yi M, Xiang B (2019) FOXA1 reprograms the TGF-beta-stimulated transcriptional program from a metastasis promoter to a tumor suppressor in nasopharyngeal carcinoma. Cancer Lett 442:1–14. https://doi.org/10.1016/j.canlet.2018.10.036
Chiang CT, Chu WK, Chow SE, Chen JK (2009) Overexpression of delta Np63 in a human nasopharyngeal carcinoma cell line downregulates CKIs and enhances cell proliferation. J Cell Physiol 219(1):117–122. https://doi.org/10.1002/jcp.21656
Yip YL, Tsao SW (2008) Regulation of p63 expression in primary and immortalized nasopharyngeal epithelial cells. Int J Oncol 33(4):713–724. https://doi.org/10.3892/ijo_00000057
Cai J, Chen S, Yi M, Tan Y, Peng Q, Ban Y, Yang J, Li X, Zeng Z, Xiong W, McCarthy JB, Li G, Li X, Xiang B (2019) DeltaNp63alpha is a super enhancer-enriched master factor controlling the basal-to-luminal differentiation transcriptional program and gene regulatory networks in nasopharyngeal carcinoma. Carcinogenesis. https://doi.org/10.1093/carcin/bgz203
Chen S, Youhong T, Tan Y, He Y, Ban Y, Cai J, Li X, Xiong W, Zeng Z, Li G, Yi M, Liu W, Xiang B (2019) EGFR-PKM2 signaling promotes the metastatic potential of nasopharyngeal carcinoma through induction of FOSL1 and ANTXR2. Carcinogenesis. https://doi.org/10.1093/carcin/bgz180
Jiang Y, Jiang YY, Xie JJ, Mayakonda A, Hazawa M, Chen L, Xiao JF, Li CQ, Huang ML, Ding LW, Sun QY, Xu L, Kanojia D, Jeitany M, Deng JW, Liao LD, Soukiasian HJ, Berman BP, Hao JJ, Xu LY, Li EM, Wang MR, Bi XG, Lin DC, Koeffler HP (2018) Co-activation of super-enhancer-driven CCAT1 by TP63 and SOX2 promotes squamous cancer progression. Nat Commun. https://doi.org/10.1038/S41467-018-06081-9
Xie JJ, Jiang YY, Jiang Y, Li CQ, Lim MC, An O, Mayakonda A, Ding LW, Long L, Sun C, Lin LH, Chen L, Wu JY, Wu ZY, Cao Q, Fang WK, Yang W, Soukiasian H, Meltzer SJ, Yang H, Fullwood M, Xu LY, Li EM, Lin DC, Koeffler HP (2018) Super-enhancer-driven long non-coding RNA LINC01503, regulated by TP63, is over-expressed and oncogenic in squamous cell carcinoma. Gastroenterology 154(8):2137. https://doi.org/10.1053/j.gastro.2018.02.018
Zhang E, Han L, Yin D, He X, Hong L, Si X, Qiu M, Xu T, De W, Xu L, Shu Y, Chen J (2017) H3K27 acetylation activated-long non-coding RNA CCAT1 affects cell proliferation and migration by regulating SPRY4 and HOXB13 expression in esophageal squamous cell carcinoma. Nucleic Acids Res 45(6):3086–3101. https://doi.org/10.1093/nar/gkw1247
Hahn S (2018) Phase separation, protein disorder, and enhancer function. Cell 175(7):1723–1725. https://doi.org/10.1016/j.cell.2018.11.034
Sabari BR, Dall'Agnese A, Boija A, Klein IA, Coffey EL, Shrinivas K, Abraham BJ, Hannett NM, Zamudio AV, Manteiga JC, Li CH, Guo YE, Day DS, Schuijers J, Vasile E, Malik S, Hnisz D, Lee TI, Cisse II, Roeder RG, Sharp PA, Chakraborty AK, Young RA (2018) Coactivator condensation at super-enhancers links phase separation and gene control. Science. https://doi.org/10.1126/science.aar3958
Boija A, Klein IA, Sabari BR, Dall'Agnese A, Coffey EL, Zamudio AV, Li CH, Shrinivas K, Manteiga JC, Hannett NM, Abraham BJ, Afeyan LK, Guo YE, Rimel JK, Fant CB, Schuijers J, Lee TI, Taatjes DJ, Young RA (2018) Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175(7):1842–1855.e1816. https://doi.org/10.1016/j.cell.2018.10.042
Nair SJ, Yang L, Meluzzi D, Oh S, Yang F, Friedman MJ, Wang S, Suter T, Alshareedah I, Gamliel A, Ma Q, Zhang J, Hu YR, Tan YL, Ohgi KA, Jayani RS, Banerjee PR, Aggarwal AK, Rosenfeld MG (2019) Phase separation of ligand-activated enhancers licenses cooperative chromosomal enhancer assembly. Nat Struct Mol Biol 26(3):193. https://doi.org/10.1038/s41594-019-0190-5
Cermakova K, Hodges HC (2018) Next-generation drugs and probes for chromatin biology: from targeted protein degradation to phase separation. Molecules. https://doi.org/10.3390/molecules23081958
Watanabe H, Ma QP, Peng SY, Adelmant G, Swain D, Song WY, Fox C, Francis JM, Pedamallu CS, DeLuca DS, Brooks AN, Wang S, Que JW, Rustgi AK, Wong KK, Ligon KL, Liu XS, Marto JA, Meyerson M, Bass AJ (2014) SOX2 and p63 colocalize at genetic loci in squamous cell carcinomas. J Clin Investig 124(4):1636–1645. https://doi.org/10.1172/JCI71545
Bass AJ, Watanabe H, Mermel CH, Yu SY, Perner S, Verhaak RG, Kim SY, Wardwell L, Tamayo P, Gat-Viks I, Ramos AH, Woo MS, Weir BA, Getz G, Beroukhim R, O'Kelly M, Dutt A, Rozenblatt-Rosen O, Dziunycz P, Komisarof J, Chirieac LR, LaFargue CJ, Scheble V, Wilbertz T, Ma CQ, Rao S, Nakagawa H, Stairs DB, Lin L, Giordano TJ, Wagner P, Minna JD, Gazdar AF, Zhu CQ, Brose MS, Cecconello I, Ribeiro U, Marie SK, Dahl O, Shivdasani RA, Tsao MS, Rubin MA, Wong KK, Regev A, Hahn WC, Beer DG, Rustgi AK, Meyerson M (2009) SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat Genet 41(11):1238–U1105. https://doi.org/10.1038/ng.465
Hussenet T, Dali S, Exinger J, Monga B, Jost B, Dembele D, Martinet N, Thibault C, Huelsken J, Brambilla E, du Manoir S (2010) SOX2 is an oncogene activated by recurrent 3q26.3 amplifications in human lung squamous cell carcinomas. PLoS ONE. https://doi.org/10.1371/journal.pone.0008960
Gen Y, Yasui K, Zen Y, Zen K, Dohi O, Endo M, Tsuji K, Wakabayashi N, Itoh Y, Naito Y, Taniwaki M, Nakanuma Y, Okanoue T, Yoshikawa T (2010) SOX2 identified as a target gene for the amplification at 3q26 that is frequently detected in esophageal squamous cell carcinoma. Cancer Genet Cytogenet 202(2):82–93. https://doi.org/10.1016/j.cancergencyto.2010.01.023
Maier S, Wilbertz T, Braun M, Scheble V, Reischl M, Mikut R, Menon R, Nikolov P, Petersen K, Beschorner C, Moch H, Kakies C, Protzel C, Bauer J, Soltermann A, Fend F, Staebler A, Lengerke C, Perner S (2011) SOX2 amplification is a common event in squamous cell carcinomas of different organ sites. Hum Pathol 42(8):1078–1088. https://doi.org/10.1016/j.humpath.2010.11.010
Hu DQ, Gao X, Morgan MA, Herz HM, Smith ER, Shilatifard A (2013) The MLL3/MLL4 branches of the COMPASS family function as major histone H3K4 monomethylases at enhancers. Mol Cell Biol 33(23):4745–4754. https://doi.org/10.1128/Mcb.01181-13
Shilatifard A (2012) The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu Rev Biochem 81:65–95. https://doi.org/10.1146/annurev-biochem-051710-134100
Lin-Shiao E, Lan Y, Coradin M, Anderson A, Donahue G, Simpson CL, Sen P, Saffie R, Busino L, Garcia BA, Berger SL, Capell BC (2018) KMT2D regulates p63 target enhancers to coordinate epithelial homeostasis. Genes Dev 32(2):181–193. https://doi.org/10.1101/gad.306241.117
Lin DC, Hao JJ, Nagata Y, Xu L, Shang L, Meng X, Sato Y, Okuno Y, Varela AM, Ding LW, Garg M, Liu LZ, Yang H, Yin D, Shi ZZ, Jiang YY, Gu WY, Gong T, Zhang Y, Xu X, Kalid O, Shacham S, Ogawa S, Wang MR, Koeffler HP (2014) Genomic and molecular characterization of esophageal squamous cell carcinoma. Nat Genet 46(5):467–473. https://doi.org/10.1038/ng.2935
Gao YB, Chen ZL, Li JG, Hu XD, Shi XJ, Sun ZM, Zhang F, Zhao ZR, Li ZT, Liu ZY, Zhao YD, Sun J, Zhou CC, Yao R, Wang SY, Wang P, Sun N, Zhang BH, Dong JS, Yu Y, Luo M, Feng XL, Shi SS, Zhoul F, Tan FW, Qiu B, Li N, Shao K, Zhang LJ, Zhang LJ, Xue Q, Gao SG, He J (2014) Genetic landscape of esophageal squamous cell carcinoma. Nat Genet 46(10):1097–1102. https://doi.org/10.1038/ng.3076
Song YM, Li L, Ou YW, Gao ZB, Li EM, Li XC, Zhang WM, Wang JQ, Xu LY, Zhou Y, Ma XJ, Liu LY, Zhao ZT, Huang XL, Fan J, Dong LJ, Chen G, Ma LY, Yang J, Chen LY, He MH, Li M, Zhuang XH, Huang K, Qiu KL, Yin GL, Guo GW, Feng Q, Chen PS, Wu ZY, Wu JY, Ma L, Zhao JY, Luo LH, Fu M, Xu BN, Chen B, Li YR, Tong T, Wang MR, Liu ZH, Lin DX, Zhang XQ, Yang HM, Wang J, Zhan QM (2014) Identification of genomic alterations in oesophageal squamous cell cancer. Nature 509(7498):91. https://doi.org/10.1038/nature13176
Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, Hanna J, Lodato MA, Frampton GM, Sharp PA, Boyer LA, Young RA, Jaenisch R (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci USA 107(50):21931–21936. https://doi.org/10.1073/pnas.1016071107
MacPartlin M, Zeng S, Lee H, Stauffer D, Jin Y, Thayer M, Lu H (2005) p300 regulates p63 transcriptional activity. J Biol Chem 280(34):30604–30610. https://doi.org/10.1074/jbc.M503352200
Restelli M, Molinari E, Marinari B, Conte D, Gnesutta N, Costanzo A, Merlo GR, Guerrini L (2015) FGF8, c-Abl and p300 participate in a pathway that controls stability and function of the DeltaNp63alpha protein. Hum Mol Genet 24(15):4185–4197. https://doi.org/10.1093/hmg/ddv151
Katoh I, Maehata Y, Moriishi K, Hata RI, Kurata S (2019) C-terminal alpha domain of p63 binds to p300 to coactivate beta-catenin. Neoplasia 21(5):494–503. https://doi.org/10.1016/j.neo.2019.03.010
Selvi RB, Swaminathan A, Chatterjee S, Shanmugam MK, Li F, Ramakrishnan GB, Siveen KS, Chinnathambi A, Zayed ME, Alharbi SA, Basha J, Bhat A, Vasudevan M, Dharmarajan A, Sethi G, Kundu TK (2015) Inhibition of p300 lysine acetyltransferase activity by luteolin reduces tumor growth in head and neck squamous cell carcinoma (HNSCC) xenograft mouse model. Oncotarget 6(41):43806–43818. https://doi.org/10.18632/oncotarget.6245
Xie X, Piao L, Bullock BN, Smith A, Su T, Zhang M, Teknos TN, Arora PS, Pan Q (2014) Targeting HPV16 E6–p300 interaction reactivates p53 and inhibits the tumorigenicity of HPV-positive head and neck squamous cell carcinoma. Oncogene 33(8):1037–1046. https://doi.org/10.1038/onc.2013.25
Ichise T, Yoshida N, Ichise H (2019) CBP/p300 antagonises EGFR-Ras-Erk signalling and suppresses increased Ras-Erk signalling-induced tumour formation in mice. J Pathol 249(1):39–51. https://doi.org/10.1002/path.5279
Hu C, Liu M, Zhang W, Xu Q, Ma K, Chen L, Wang Z, He S, Zhu H, Xu N (2015) Upregulation of KLF4 by methylseleninic acid in human esophageal squamous cell carcinoma cells: Modification of histone H3 acetylation through HAT/HDAC interplay. Mol Carcinog 54(10):1051–1059. https://doi.org/10.1002/mc.22174
Qiu H, Chereji RV, Hu C, Cole HA, Rawal Y, Clark DJ, Hinnebusch AG (2016) Genome-wide cooperation by HAT Gcn5, remodeler SWI/SNF, and chaperone Ydj1 in promoter nucleosome eviction and transcriptional activation. Genome Res 26(2):211–225. https://doi.org/10.1101/gr.196337.115
Sanz AB, Garcia R, Rodriguez-Pena JM, Nombela C, Arroyo J (2016) Cooperation between SAGA and SWI/SNF complexes is required for efficient transcriptional responses regulated by the yeast MAPK Slt2. Nucleic Acids Res 44(15):7159–7172. https://doi.org/10.1093/nar/gkw324
Bennett G, Peterson CL (2015) SWI/SNF recruitment to a DNA double-strand break by the NuA4 and Gcn5 histone acetyltransferases. DNA Repair 30:38–45. https://doi.org/10.1016/j.dnarep.2015.03.006
Son EY, Crabtree GR (2014) The role of BAF (mSWI/SNF) complexes in mammalian neural development. Am J Med Genet C Semin Med Genet 166C(3):333–349. https://doi.org/10.1002/ajmg.c.31416
Ho L, Crabtree GR (2010) Chromatin remodelling during development. Nature 463(7280):474–484. https://doi.org/10.1038/nature08911
Narayanan R, Pirouz M, Kerimoglu C, Pham L, Wagener RJ, Kiszka KA, Rosenbusch J, Seong RH, Kessel M, Fischer A, Stoykova A, Staiger JF, Tuoc T (2015) Loss of BAF (mSWI/SNF) complexes causes global transcriptional and chromatin state changes in forebrain development. Cell Rep 13(9):1842–1854. https://doi.org/10.1016/j.celrep.2015.10.046
Alver BH, Kim KH, Lu P, Wang XF, Manchester HE, Wang WS, Haswell JR, Park PJ, Roberts CWM (2017) The SWI/SNF chromatin remodelling complex is required for maintenance of lineage specific enhancers. Nat Commun. https://doi.org/10.1038/Ncomms14648
Mardaryev AN, Gdula MR, Yarker JL, Emelianov VU, Poterlowicz K, Sharov AA, Sharova TY, Scarpa JA, Joffe B, Solovei I, Chambon P, Botchkarev VA, Fessing MY (2014) p63 and Brg1 control developmentally regulated higher-order chromatin remodelling at the epidermal differentiation complex locus in epidermal progenitor cells. Development 141(1):101–111. https://doi.org/10.1242/dev.103200
Fessing MY, Mardaryev AN, Gdula MR, Sharov AA, Sharova TY, Rapisarda V, Gordon KB, Smorodchenko AD, Poterlowicz K, Ferone G, Kohwi Y, Missero C, Kohwi-Shigematsu T, Botchkarev VA (2011) p63 regulates Satb1 to control tissue-specific chromatin remodeling during development of the epidermis. J Cell Biol 194(6):825–839. https://doi.org/10.1083/jcb.201101148
Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo QM, Edsall L, Antosiewicz-Bourget J, Stewart R, Ruotti V, Millar AH, Thomson JA, Ren B, Ecker JR (2009) Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462(7271):315–322. https://doi.org/10.1038/nature08514
Stadler MB, Murr R, Burger L, Ivanek R, Lienert F, Scholer A, Wirbelauer C, Oakeley EJ, Gaidatzis D, Tiwari VK, Schubeler D (2011) DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480(7378):490–495. https://doi.org/10.1038/nature10716
Dunham I, Kundaje A, Aldred SF, Collins PJ, Davis C, Doyle F, Epstein CB, Frietze S, Harrow J, Kaul R, Khatun J, Lajoie BR, Landt SG, Lee BK, Pauli F, Rosenbloom KR, Sabo P, Safi A, Sanyal A, Shoresh N, Simon JM, Song L, Trinklein ND, Altshuler RC, Birney E, Brown JB, Cheng C, Djebali S, Dong XJ, Dunham I, Ernst J, Furey TS, Gerstein M, Giardine B, Greven M, Hardison RC, Harris RS, Herrero J, Hoffman MM, Iyer S, Kellis M, Khatun J, Kheradpour P, Kundaje A, Lassmann T, Li QH, Lin X, Marinov GK, Merkel A, Mortazavi A, Parker SCJ, Reddy TE, Rozowsky J, Schlesinger F, Thurman RE, Wang J, Ward LD, Whitfield TW, Wilder SP, Wu W, Xi HLS, Yip KY, Zhuang JL, Bernstein BE, Birney E, Dunham I, Green ED, Gunter C, Snyder M, Pazin MJ, Lowdon RF, Dillon LAL, Adams LB, Kelly CJ, Zhang J, Wexler JR, Green ED, Good PJ, Feingold EA, Bernstein BE, Birney E, Crawford GE, Dekker J, Elnitski L, Farnham PJ, Gerstein M, Giddings MC, Gingeras TR, Green ED, Guigo R, Hardison RC, Hubbard TJ, Kellis M, Kent WJ, Lieb JD, Margulies EH, Myers RM, Snyder M, Stamatoyannopoulos JA, Tenenbaum SA, Weng ZP, White KP, Wold B, Khatun J, Yu Y, Wrobel J, Risk BA, Gunawardena HP, Kuiper HC, Maier CW, Xie L, Chen X, Giddings MC, Bernstein BE, Epstein CB, Shoresh N, Ernst J, Kheradpour P, Mikkelsen TS, Gillespie S, Goren A, Ram O, Zhang XL, Wang L, Issner R, Coyne MJ, Durham T, Ku M, Truong T, Ward LD, Altshuler RC, Eaton ML, Kellis M, Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, Tanzer A, Lagarde J, Lin W, Schlesinger F, Xue CH, Marinov GK, Khatun J, Williams BA, Zaleski C, Rozowsky J, Roeder M, Kokocinski F, Abdelhamid RF, Alioto T, Antoshechkin I, Baer MT, Batut P, Bell I, Bell K, Chakrabortty S, Chen X, Chrast J, Curado J, Derrien T, Drenkow J, Dumais E, Dumais J, Duttagupta R, Fastuca M, Fejes-Toth K, Ferreira P, Foissac S, Fullwood MJ, Gao H, Gonzalez D, Gordon A, Gunawardena HP, Howald C, Jha S, Johnson R, Kapranov P, King B, Kingswood C, Li GL, Luo OJ, Park E, Preall JB, Presaud K, Ribeca P, Risk BA, Robyr D, Ruan XA, Sammeth M, Sandhu KS, Schaeffer L, See LH, Shahab A, Skancke J, Suzuki AM, Takahashi H, Tilgner H, Trout D, Walters N, Wang HE, Wrobel J, Yu YB, Hayashizaki Y, Harrow J, Gerstein M, Hubbard TJ, Reymond A, Antonarakis SE, Hannon GJ, Giddings MC, Ruan YJ, Wold B, Carninci P, Guigo R, Gingeras TR, Rosenbloom KR, Sloan CA, Learned K, Malladi VS, Wong MC, Barber G, Cline MS, Dreszer TR, Heitner SG, Karolchik D, Kent WJ, Kirkup VM, Meyer LR, Long JC, Maddren M, Raney BJ, Furey TS, Song LY, Grasfeder LL, Giresi PG, Lee BK, Battenhouse A, Sheffield NC, Simon JM, Showers KA, Safi A, London D, Bhinge AA, Shestak C, Schaner MR, Kim SK, Zhang ZZZ, Mieczkowski PA, Mieczkowska JO, Liu Z, McDaniell RM, Ni YY, Rashid NU, Kim MJ, Adar S, Zhang ZC, Wang TY, Winter D, Keefe D, Birney E, Iyer VR, Lieb JD, Crawford GE, Li GL, Sandhu KS, Zheng MZ, Wang P, Luo OJ, Shahab A, Fullwood MJ, Ruan XA, Ruan YJ, Myers RM, Pauli F, Williams BA, Gertz J, Marinov GK, Reddy TE, Vielmetter J, Partridge EC, Trout D, Varley KE, Gasper C, Bansal A, Pepke S, Jain P, Amrhein H, Bowling KM, Anaya M, Cross MK, King B, Muratet MA, Antoshechkin I, Newberry KM, Mccue K, Nesmith AS, Fisher-Aylor KI, Pusey B, DeSalvo G, Parker SL, Balasubramanian S, Davis NS, Meadows SK, Eggleston T, Gunter C, Newberry JS, Levy SE, Absher DM, Mortazavi A, Wong WH, Wold B, Blow MJ, Visel A, Pennachio LA, Elnitski L, Margulies EH, Parker SCJ, Petrykowska HM, Abyzov A, Aken B, Barrell D, Barson G, Berry A, Bignell A, Boychenko V, Bussotti G, Chrast J, Davidson C, Derrien T, Despacio-Reyes G, Diekhans M, Ezkurdia I, Frankish A, Gilbert J, Gonzalez JM, Griffiths E, Harte R, Hendrix DA, Howald C, Hunt T, Jungreis I, Kay M, Khurana E, Kokocinski F, Leng J, Lin MF, Loveland J, Lu Z, Manthravadi D, Mariotti M, Mudge J, Mukherjee G, Notredame C, Pei BK, Rodriguez JM, Saunders G, Sboner A, Searle S, Sisu C, Snow C, Steward C, Tanzer A, Tapanari E, Tress ML, van Baren MJ, Walters N, Washietl S, Wilming L, Zadissa A, Zhang ZD, Brent M, Haussler D, Kellis M, Valencia A, Gerstein M, Reymond A, Guigo R, Harrow J, Hubbard TJ, Landt SG, Frietze S, Abyzov A, Addleman N, Alexander RP, Auerbach RK, Balasubramanian S, Bettinger K, Bhardwaj N, Boyle AP, Cao AR, Cayting P, Charos A, Cheng Y, Cheng C, Eastman C, Euskirchen G, Fleming JD, Grubert F, Habegger L, Hariharan M, Harmanci A, Iyengar S, Jin VX, Karczewski KJ, Kasowski M, Lacroute P, Lam H, Lamarre-Vincent N, Leng J, Lian J, Lindahl-Allen M, Min RQ, Miotto B, Monahan H, Moqtaderi Z, Mu XMJ, O'Geen H, Ouyang ZQ, Patacsil D, Pei BK, Raha D, Ramirez L, Reed B, Rozowsky J, Sboner A, Shi MY, Sisu C, Slifer T, Witt H, Wu LF, Xu XQ, Yan KK, Yang XQ, Yip KY, Zhang ZD, Struhl K, Weissman SM, Gerstein M, Farnham PJ, Snyder M, Tenenbaum SA, Penalva LO, Doyle F, Karmakar S, Landt SG, Bhanvadia RR, Choudhury A, Domanus M, Ma LJ, Moran J, Patacsil D, Slifer T, Victorsen A, Yang XQ, Snyder M, White KP, Auer T, Centanin L, Eichenlaub M, Gruhl F, Heermann S, Hoeckendorf B, Inoue D, Kellner T, Kirchmaier S, Mueller C, Reinhardt R, Schertel L, Schneider S, Sinn R, Wittbrodt B, Wittbrodt J, Weng ZP, Whitfield TW, Wang J, Collins PJ, Aldred SF, Trinklein ND, Partridge EC, Myers RM, Dekker J, Jain G, Lajoie BR, Sanyal A, Balasundaram G, Bates DL, Byron R, Canfield TK, Diegel MJ, Dunn D, Ebersol AK, Frum T, Garg K, Gist E, Hansen RS, Boatman L, Haugen E, Humbert R, Jain G, Johnson AK, Johnson EM, Kutyavin TV, Lajoie BR, Lee K, Lotakis D, Maurano MT, Neph SJ, Neri FV, Nguyen ED, Qu HZ, Reynolds AP, Roach V, Rynes E, Sabo P, Sanchez ME, Sandstrom RS, Sanyal A, Shafer AO, Stergachis AB, Thomas S, Thurman RE, Vernot B, Vierstra J, Vong S, Wang H, Weaver MA, Yan YQ, Zhang MH, Akey JM, Bender M, Dorschner MO, Groudine M, MacCoss MJ, Navas P, Stamatoyannopoulos G, Kaul R, Dekker J, Stamatoyannopoulos JA, Dunham I, Beal K, Brazma A, Flicek P, Herrero J, Johnson N, Keefe D, Lukk M, Luscombe NM, Sobral D, Vaquerizas JM, Wilder SP, Batzoglou S, Sidow A, Hussami N, Kyriazopoulou-Panagiotopoulou S, Libbrecht MW, Schaub MA, Kundaje A, Hardison RC, Miller W, Giardine B, Harris RS, Wu W, Bickel PJ, Banfai B, Boley NP, Brown JB, Huang HY, Li QH, Li JJ, Noble WS, Bilmes JA, Buske OJ, Hoffman MM, Sahu AD, Kharchenko PV, Park PJ, Baker D, Taylor J, Weng ZP, Iyer S, Dong XJ, Greven M, Lin XY, Wang J, Xi HLS, Zhuang JL, Gerstein M, Alexander RP, Balasubramanian S, Cheng C, Harmanci A, Lochovsky L, Min R, Mu XMJ, Rozowsky J, Yan KK, Yip KY, Birney E, EP Consortium (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489(7414):57–74. https://doi.org/10.1038/nature11247
Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, Heravi-Moussavi A, Kheradpour P, Zhang Z, Wang J, Ziller MJ, Amin V, Whitaker JW, Schultz MD, Ward LD, Sarkar A, Quon G, Sandstrom RS, Eaton ML, Wu YC, Pfenning AR, Wang X, Claussnitzer M, Liu Y, Coarfa C, Harris RA, Shoresh N, Epstein CB, Gjoneska E, Leung D, Xie W, Hawkins RD, Lister R, Hong C, Gascard P, Mungall AJ, Moore R, Chuah E, Tam A, Canfield TK, Hansen RS, Kaul R, Sabo PJ, Bansal MS, Carles A, Dixon JR, Farh KH, Feizi S, Karlic R, Kim AR, Kulkarni A, Li D, Lowdon R, Elliott G, Mercer TR, Neph SJ, Onuchic V, Polak P, Rajagopal N, Ray P, Sallari RC, Siebenthall KT, Sinnott-Armstrong NA, Stevens M, Thurman RE, Wu J, Zhang B, Zhou X, Beaudet AE, Boyer LA, De Jager PL, Farnham PJ, Fisher SJ, Haussler D, Jones SJM, Li W, Marra MA, McManus MT, Sunyaev S, Thomson JA, Tlsty TD, Tsai LH, Wang W, Waterland RA, Zhang MQ, Chadwick LH, Bernstein BE, Costello JF, Ecker JR, Hirst M, Meissner A, Milosavljevic A, Ren B, Stamatoyannopoulos JA, Wang T, Kellis M, Consortium RE (2015) Integrative analysis of 111 reference human epigenomes. Nature 518(7539):317–330. https://doi.org/10.1038/nature14248
Szulwach KE, Li XK, Li YJ, Song CX, Han JW, Kim S, Namburi S, Hermetz K, Kim JJ, Rudd MK, Yoon YS, Ren B, He C, Jin P (2011) Integrating 5-hydroxymethylcytosine into the epigenomic landscape of human embryonic stem cells. PLoS Genet. https://doi.org/10.1371/journal.pgen.1002154
Stroud H, Feng SH, Kinney SM, Pradhan S, Jacobsen SE (2011) 5-Hydroxymethylcytosine is associated with enhancers and gene bodies in human embryonic stem cells. Genome Biol. https://doi.org/10.1186/Gb-2011-12-6-R54
Sun ZY, Terragni J, Borgaro JG, Liu YW, Yu L, Guan SX, Wang H, Sun DP, Cheng XD, Zhu ZY, Pradhan S, Zheng Y (2013) High-resolution enzymatic mapping of genomic 5-hydroxymethylcytosine in mouse embryonic stem cells. Cell Rep 3(2):567–576. https://doi.org/10.1016/j.celrep.2013.01.001
Pulakanti K, Pinello L, Stelloh C, Blinka S, Allred J, Milanovich S, Kiblawi S, Peterson J, Wang A, Yuan GC, Rao S (2013) Enhancer transcribed RNAs arise from hypomethylated, Tet-occupied genomic regions. Epigenetics-US 8(12):1303–1320. https://doi.org/10.4161/epi.26597
Bhattacharyya S, Pradhan K, Campbell N, Mazdo J, Vasantkumar A, Maqbool S, Bhagat TD, Gupta S, Suzuki M, Yu YT, Greally JM, Steidl U, Bradner J, Dawlaty M, Godley L, Maitra A, Verma A (2017) Altered hydroxymethylation is seen at regulatory regions in pancreatic cancer and regulates oncogenic pathways. Genome Res 27(11):1830–1842. https://doi.org/10.1101/gr.222794.117
Michalak EM, Burr ML, Bannister AJ, Dawson MA (2019) The roles of DNA, RNA and histone methylation in ageing and cancer. Nat Rev Mol Cell Biol. https://doi.org/10.1038/s41580-019-0143-1
Lyko F (2018) The DNA methyltransferase family: a versatile toolkit for epigenetic regulation. Nat Rev Genet 19(2):81–92. https://doi.org/10.1038/nrg.2017.80
Rondelet G, Wouters J (2017) Human DNA (cytosine-5)-methyltransferases: a functional and structural perspective for epigenetic cancer therapy. Biochimie 139:137–147. https://doi.org/10.1016/j.biochi.2017.06.003
Charostad J, Astani A, Goudarzi H, Faghihloo E (2019) DNA methyltransferases in virus-associated cancers. Rev Med Virol 29(2):e2022. https://doi.org/10.1002/rmv.2022
Ratovitski EA (2014) Phospho-Delta Np63 alpha/microRNA network modulates epigenetic regulatory enzymes in squamous cell carcinomas. Cell Cycle 13(5):749–761. https://doi.org/10.4161/cc.27676
Ramsey MR, He L, Forster N, Ory B, Ellisen LW (2011) Physical Association of HDAC1 and HDAC2 with p63 mediates transcriptional repression and tumor maintenance in squamous cell carcinoma. Cancer Res 71(13):4373–4379. https://doi.org/10.1158/0008-5472.CAN-11-0046
LeBoeuf M, Terrell A, Trivedi S, Sinha S, Epstein JA, Olson EN, Morrisey EE, Millar SE (2010) Hdac1 and Hdac2 act redundantly to control p63 and p53 functions in epidermal progenitor cells. Dev Cell 19(6):807–818. https://doi.org/10.1016/j.devcel.2010.10.015
Urvalek AM, Gudas LJ (2014) Retinoic acid and histone deacetylases regulate epigenetic changes in embryonic stem cells. J Biol Chem 289(28):19519–19530. https://doi.org/10.1074/jbc.M114.556555
Pietrzak J, Ploszaj T, Pulaski L, Robaszkiewicz A (2019) EP300-HDAC1-SWI/SNF functional unit defines transcription of some DNA repair enzymes during differentiation of human macrophages. Biochim Biophys Acta 1862(2):198–208. https://doi.org/10.1016/j.bbagrm.2018.10.019
Xu Z, Meng X, Cai Y, Koury MJ, Brandt SJ (2006) Recruitment of the SWI/SNF protein Brg1 by a multiprotein complex effects transcriptional repression in murine erythroid progenitors. Biochem J 399(2):297–304. https://doi.org/10.1042/BJ20060873
Pal S, Yun R, Datta A, Lacomis L, Erdjument-Bromage H, Kumar J, Tempst P, Sif S (2003) mSin3A/histone deacetylase 2- and PRMT5-containing Brg1 complex is involved in transcriptional repression of the Myc target gene cad. Mol Cell Biol 23(21):7475–7487. https://doi.org/10.1128/mcb.23.21.7475-7487.2003
Bao X, Tang J, Lopez-Pajares V, Tao S, Qu K, Crabtree GR, Khavari PA (2013) ACTL6a enforces the epidermal progenitor state by suppressing SWI/SNF-dependent induction of KLF4. Cell Stem Cell 12(2):193–203. https://doi.org/10.1016/j.stem.2012.12.014
Lu W, Fang L, Ouyang B, Zhang X, Zhan S, Feng X, Bai Y, Han X, Kim H, He Q, Wan M, Shi FT, Feng XH, Liu D, Huang J, Songyang Z (2015) Actl6a protects embryonic stem cells from differentiating into primitive endoderm. Stem Cells 33(6):1782–1793. https://doi.org/10.1002/stem.2000
Zhang Y, Cui P, Li Y, Feng G, Tong M, Guo L, Li T, Liu L, Li W, Zhou Q (2018) Mitochondrially produced ATP affects stem cell pluripotency via Actl6a-mediated histone acetylation. FASEB J 32(4):1891–1902. https://doi.org/10.1096/fj.201700626RR
Park J, Wood MA, Cole MD (2002) BAF53 forms distinct nuclear complexes and functions as a critical c-Myc-interacting nuclear cofactor for oncogenic transformation. Mol Cell Biol 22(5):1307–1316. https://doi.org/10.1128/mcb.22.5.1307-1316.2002
Taulli R, Foglizzo V, Morena D, Coda DM, Ala U, Bersani F, Maestro N, Ponzetto C (2014) Failure to downregulate the BAF53a subunit of the SWI/SNF chromatin remodeling complex contributes to the differentiation block in rhabdomyosarcoma. Oncogene 33(18):2354–2362. https://doi.org/10.1038/onc.2013.188
Saladi SV, Ross K, Karaayvaz M, Tata PR, Mou H, Rajagopal J, Ramaswamy S, Ellisen LW (2017) ACTL6A is co-amplified with p63 in squamous cell carcinoma to drive YAP activation, regenerative proliferation, and poor prognosis. Cancer Cell 31(1):35–49. https://doi.org/10.1016/j.ccell.2016.12.001
Alfert A, Moreno N, Kerl K (2019) The BAF complex in development and disease. Epigenetics Chromatin 12(1):19. https://doi.org/10.1186/s13072-019-0264-y
Kadoch C, Hargreaves DC, Hodges C, Elias L, Ho L, Ranish J, Crabtree GR (2013) Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat Genet 45(6):592. https://doi.org/10.1038/ng.2628
Simon JA, Kingston RE (2013) Occupying chromatin: polycomb mechanisms for getting to genomic targets, stopping transcriptional traffic, and staying put. Mol Cell 49(5):808–824. https://doi.org/10.1016/j.molcel.2013.02.013
Liu B, Liu YF, Du YR, Mardaryev AN, Yang W, Chen H, Xu ZM, Xu CQ, Zhang XR, Botchkarev VA, Zhang Y, Xu GL (2013) Cbx4 regulates the proliferation of thymic epithelial cells and thymus function. Development 140(4):780–788. https://doi.org/10.1242/dev.085035
Endoh M, Endo TA, Endoh T, Isono K, Sharif J, Ohara O, Toyoda T, Ito T, Eskeland R, Bickmore WA, Vidal M, Bernstein BE, Koseki H (2012) Histone H2A mono-ubiquitination is a crucial step to mediate PRC1-dependent repression of developmental genes to maintain ES cell identity. PLoS Genet. https://doi.org/10.1371/journal.pgen.1002774
Zhang L, Zhou Y, Cheng C, Cui H, Cheng L, Kong P, Wang J, Li Y, Chen W, Song B, Wang F, Jia Z, Li L, Li Y, Yang B, Liu J, Shi R, Bi Y, Zhang Y, Wang J, Zhao Z, Hu X, Yang J, Li H, Gao Z, Chen G, Huang X, Yang X, Wan S, Chen C, Li B, Tan Y, Chen L, He M, Xie S, Li X, Zhuang X, Wang M, Xia Z, Luo L, Ma J, Dong B, Zhao J, Song Y, Ou Y, Li E, Xu L, Wang J, Xi Y, Li G, Xu E, Liang J, Yang X, Guo J, Chen X, Zhang Y, Li Q, Liu L, Li Y, Zhang X, Yang H, Lin D, Cheng X, Guo Y, Wang J, Zhan Q, Cui Y (2015) Genomic analyses reveal mutational signatures and frequently altered genes in esophageal squamous cell carcinoma. Am J Hum Genet 96(4):597–611. https://doi.org/10.1016/j.ajhg.2015.02.017
Menon DU, Shibata Y, Mu W, Magnuson T (2019) Mammalian SWI/SNF collaborates with a polycomb-associated protein to regulate male germline transcription in the mouse. Development. https://doi.org/10.1242/dev.174094
Kia SK, Gorski MM, Giannakopoulos S, Verrijzer CP (2008) SWI/SNF mediates polycomb eviction and epigenetic reprogramming of the INK4b-ARF-INK4a locus. Mol Cell Biol 28(10):3457–3464. https://doi.org/10.1128/MCB.02019-07
Wong MM, Cox LK, Chrivia JC (2007) The chromatin remodeling protein, SRCAP, is critical for deposition of the histone variant H2A.Z at promoters. J Biol Chem 282(36):26132–26139. https://doi.org/10.1074/jbc.M703418200
Ruhl DD, Jin JJ, Cai Y, Swanson S, Florens L, Washburn MP, Conaway RC, Conaway JW, Chrivia JC (2006) Purification of a human SRCAP complex that remodels chromatin by incorporating the histone variant H2A.Z into nucleosomes. Biochemistry-US 45(17):5671–5677. https://doi.org/10.1021/bi060043d
Gallant-Behm CL, Ramsey MR, Bensard CL, Nojek I, Tran J, Liu M, Ellisen LW, Espinosa JM (2012) DeltaNp63alpha represses anti-proliferative genes via H2A.Z deposition. Genes Dev 26(20):2325–2336. https://doi.org/10.1101/gad.198069.112
Dennis K, Fan T, Geiman T, Yan QS, Muegge K (2001) Lsh, a member of the SNF2 family, is required for genome-wide methylation. Genes Dev 15(22):2940–2944. https://doi.org/10.1101/Gad.929101
Liu XL, Ding J, Meng LH (2018) Oncogene-induced senescence: a double edged sword in cancer. Acta Pharmacol Sin 39(10):1553–1558. https://doi.org/10.1038/aps.2017.198
Keyes WM, Wu Y, Vogel H, Guo X, Lowe SW, Mills AA (2005) p63 deficiency activates a program of cellular senescence and leads to accelerated aging. Genes Dev 19(17):1986–1999. https://doi.org/10.1101/gad.342305
Keyes WM, Pecoraro M, Aranda V, Vernersson-Lindahl E, Li W, Vogel H, Guo X, Garcia EL, Michurina TV, Enikolopov G, Muthuswamy SK, Mills AA (2011) DeltaNp63alpha is an oncogene that targets chromatin remodeler Lsh to drive skin stem cell proliferation and tumorigenesis. Cell Stem Cell 8(2):164–176. https://doi.org/10.1016/j.stem.2010.12.009
Yu W, McIntosh C, Lister R, Zhu I, Han Y, Ren J, Landsman D, Lee E, Briones V, Terashima M, Leighty R, Ecker JR, Muegge K (2014) Genome-wide DNA methylation patterns in LSH mutant reveals de-repression of repeat elements and redundant epigenetic silencing pathways. Genome Res 24(10):1613–1623. https://doi.org/10.1101/gr.172015.114
Huang J, Fan T, Yan Q, Zhu H, Fox S, Issaq HJ, Best L, Gangi L, Munroe D, Muegge K (2004) Lsh, an epigenetic guardian of repetitive elements. Nucleic Acids Res 32(17):5019–5028. https://doi.org/10.1093/nar/gkh821
Ren J, Finney R, Ni K, Cam M, Muegge K (2019) The chromatin remodeling protein Lsh alters nucleosome occupancy at putative enhancers and modulates binding of lineage specific transcription factors. Epigenetics-US 14(3):277–293. https://doi.org/10.1080/15592294.2019.1582275
Yu W, Briones V, Lister R, McIntosh C, Han Y, Lee EY, Ren J, Terashima M, Leighty RM, Ecker JR, Muegge K (2014) CG hypomethylation in Lsh−/− mouse embryonic fibroblasts is associated with de novo H3K4me1 formation and altered cellular plasticity. Proc Natl Acad Sci USA 111(16):5890–5895. https://doi.org/10.1073/pnas.1320945111
He X, Yan B, Liu S, Jia J, Lai W, Xin X, Tang CE, Luo D, Tan T, Jiang Y, Shi Y, Liu Y, Xiao D, Chen L, Liu S, Mao C, Yin G, Cheng Y, Fan J, Cao Y, Muegge K, Tao Y (2016) Chromatin remodeling factor LSH drives cancer progression by suppressing the activity of fumarate hydratase. Cancer Res 76(19):5743–5755. https://doi.org/10.1158/0008-5472.CAN-16-0268
Xiao DS, Huang J, Pan Y, Li H, Fu CY, Mao C, Cheng Y, Shi Y, Chen L, Jiang YQ, Yang R, Liu YT, Zhou JH, Cao Y, Liu S, Tao YG (2017) Chromatin remodeling factor LSH is upregulated by the LRP6-GSK3(beta)-E2F1 axis linking reversely with survival in gliomas. Theranostics 7(1):132–143. https://doi.org/10.7150/thno.17032
Yang X, Miao BS, Wei CY, Dong RZ, Gao PT, Zhang XY, Lu JC, Gao C, Wang XY, Sun HC, Zhou J, Fan J, Ke AW, Shi GM, Cai JB (2019) Lymphoid-specific helicase promotes the growth and invasion of hepatocellular carcinoma by transcriptional regulation of centromere protein F expression. Cancer Sci 110(7):2133–2144. https://doi.org/10.1111/cas.14037
Zhou R, Gao Y, Lv D, Wang C, Wang D, Li Q (2019) METTL3 mediated m(6)A modification plays an oncogenic role in cutaneous squamous cell carcinoma by regulating DeltaNp63. Biochem Biophys Res Commun 515(2):310–317. https://doi.org/10.1016/j.bbrc.2019.05.155
Li XC, Jin F, Wang BY, Yin XJ, Hong W, Tian FJ (2019) The m6A demethylase ALKBH5 controls trophoblast invasion at the maternal–fetal interface by regulating the stability of CYR61 mRNA. Theranostics 9(13):3853–3865. https://doi.org/10.7150/thno.31868
Hazawa M, Lin DC, Kobayashi A, Jiang YY, Xu L, Dewi FRP, Mohamed MS, Hartono NM, Meguro-Horike M, Horike SI, Koeffler HP, Wong RW (2018) ROCK-dependent phosphorylation of NUP62 regulates p63 nuclear transport and squamous cell carcinoma proliferation. EMBO Rep 19(1):73–88. https://doi.org/10.15252/embr.201744523
Armstrong SR, Wu H, Wang B, Abuetabh Y, Sergi C, Leng RP (2016) The regulation of tumor suppressor p63 by the ubiquitin–proteasome system. Int J Mol Sci. https://doi.org/10.3390/ijms17122041
Rokudai S, Li Y, Otaka Y, Fujieda M, Owens DM, Christiano AM, Nishiyama M, Prives C (2018) STXBP4 regulates APC/C-mediated p63 turnover and drives squamous cell carcinogenesis. Proc Natl Acad Sci USA 115(21):E4806–E4814. https://doi.org/10.1073/pnas.1718546115
Li Y, Peart MJ, Prives C (2009) Stxbp4 regulates DeltaNp63 stability by suppression of RACK1-dependent degradation. Mol Cell Biol 29(14):3953–3963. https://doi.org/10.1128/MCB.00449-09
Otaka Y, Rokudai S, Kaira K, Fujieda M, Horikoshi I, Iwakawa-Kawabata R, Yoshiyama S, Yokobori T, Ohtaki Y, Shimizu K, Oyama T, Tamura J, Prives C, Nishiyama M (2017) STXBP4 drives tumor growth and is associated with poor prognosis through PDGF receptor signaling in lung squamous cell carcinoma. Clin Cancer Res 23(13):3442–3452. https://doi.org/10.1158/1078-0432.CCR-16-1815
Harvey KF, Kumar S (1999) Nedd4-like proteins: an emerging family of ubiquitin-protein ligases implicated in diverse cellular functions. Trends Cell Biol 9(5):166–169
Chen C, Matesic LE (2007) The Nedd4-like family of E3 ubiquitin ligases and cancer. Cancer Metastasis Rev 26(3–4):587–604. https://doi.org/10.1007/s10555-007-9091-x
Rossi M, Aqeilan RI, Neale M, Candi E, Salomoni P, Knight RA, Croce CM, Melino G (2006) The E3 ubiquitin ligase Itch controls the protein stability of p63. Proc Natl Acad Sci USA 103(34):12753–12758. https://doi.org/10.1073/pnas.0603449103
Melino S, Bellomaria A, Nepravishta R, Paci M, Melino G (2014) p63 threonine phosphorylation signals the interaction with the WW domain of the E3 ligase Itch. Cell Cycle 13(20):3207–3217. https://doi.org/10.4161/15384101.2014.951285
Lu KP, Zhou XZ (2007) The prolyl isomerase PIN1: a pivotal new twist in phosphorylation signalling and disease. Nat Rev Mol Cell Biol 8(11):904–916. https://doi.org/10.1038/nrm2261
Li Y, Zhou Z, Chen C (2008) WW domain-containing E3 ubiquitin protein ligase 1 targets p63 transcription factor for ubiquitin-mediated proteasomal degradation and regulates apoptosis. Cell Death Differ 15(12):1941–1951. https://doi.org/10.1038/cdd.2008.134
Li C, Chang DL, Yang Z, Qi J, Liu R, He H, Li D, Xiao ZX (2013) Pin1 modulates p63alpha protein stability in regulation of cell survival, proliferation and tumor formation. Cell Death Dis 4:e943. https://doi.org/10.1038/cddis.2013.468
Yi Y, Chen D, Ao J, Sun S, Wu M, Li X, Bergholz J, Zhang Y, Xiao ZX (2017) Metformin promotes AMP-activated protein kinase-independent suppression of DeltaNp63alpha protein expression and inhibits cancer cell viability. J Biol Chem 292(13):5253–5261. https://doi.org/10.1074/jbc.M116.769141
Fomenkov A, Zangen R, Huang YP, Osada M, Guo Z, Fomenkov T, Trink B, Sidransky D, Ratovitski EA (2004) RACK1 and stratifin target DeltaNp63alpha for a proteasome degradation in head and neck squamous cell carcinoma cells upon DNA damage. Cell Cycle 3(10):1285–1295. https://doi.org/10.4161/cc.3.10.1155
King KE, Ponnamperuma RM, Allen C, Lu H, Duggal P, Chen Z, Van Waes C, Weinberg WC (2008) The p53 homologue DeltaNp63alpha interacts with the nuclear factor-kappaB pathway to modulate epithelial cell growth. Cancer Res 68(13):5122–5131. https://doi.org/10.1158/0008-5472.CAN-07-6123
King KE, George AL, Sakakibara N, Mahmood K, Moses MA, Weinberg WC (2019) Intersection of the p63 and NF-kappaB pathways in epithelial homeostasis and disease. Mol Carcinog 58(9):1571–1580. https://doi.org/10.1002/mc.23081
Hirata E, Sahai E (2017) Tumor microenvironment and differential responses to therapy. Cold Spring Harb Perspect Med. https://doi.org/10.1101/cshperspect.a026781
Wu T, Dai Y (2017) Tumor microenvironment and therapeutic response. Cancer Lett 387:61–68. https://doi.org/10.1016/j.canlet.2016.01.043
Migden MR, Rischin D, Schmults CD, Guminski A, Hauschild A, Lewis KD, Chung CH, Hernandez-Aya L, Lim AM, Chang ALS, Rabinowits G, Thai AA, Dunn LA, Hughes BGM, Khushalani NI, Modi B, Schadendorf D, Gao B, Seebach F, Li S, Li J, Mathias M, Booth J, Mohan K, Stankevich E, Babiker HM, Brana I, Gil-Martin M, Homsi J, Johnson ML, Moreno V, Niu J, Owonikoko TK, Papadopoulos KP, Yancopoulos GD, Lowy I, Fury MG (2018) PD-1 blockade with cemiplimab in advanced cutaneous squamous-cell carcinoma. N Engl J Med 379(4):341–351. https://doi.org/10.1056/NEJMoa1805131
Davis RJ, Van Waes C, Allen CT (2016) Overcoming barriers to effective immunotherapy: MDSCs, TAMs, and Tregs as mediators of the immunosuppressive microenvironment in head and neck cancer. Oral Oncol 58:59–70. https://doi.org/10.1016/j.oraloncology.2016.05.002
Acknowledgements
We apologize to those researchers whose work in the field could not be cited in the present review due to space constraints.
Funding
This study was supported in part by grants from The National Natural Science Foundation of China (81572667, 81772902, 81872278, 81703131), the National “111” Project (Project #111-2-12), The Natural Science Foundation of Hunan Province, China (2018JJ1040), Research Program of Hunan Health Commission, China (20201067, 20201040), and Hunan Provincial Key Research and Development Program (2018SK2130, 2018SK2131).
Author information
Authors and Affiliations
Contributions
BX, PQT, and XYL designed the idea for the article. MY, YXT, and BX performed the literature search and data analysis. MY, YXT, and XYL drafted the manuscript. MY and YXT prepared the illustrations in the article. LW, JC, XLL, ZZY, WX, GYL, XYL, PQT, and BX revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Consent for publication
All authors read and approved the final manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yi, M., Tan, Y., Wang, L. et al. TP63 links chromatin remodeling and enhancer reprogramming to epidermal differentiation and squamous cell carcinoma development. Cell. Mol. Life Sci. 77, 4325–4346 (2020). https://doi.org/10.1007/s00018-020-03539-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-020-03539-2