Abstract
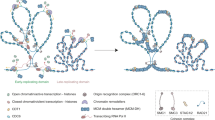
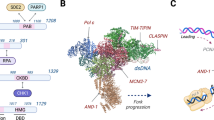
Although INO80 chromatin remodeling enzyme has been shown in yeast to play roles in non-transcriptional nuclear processes such as DNA replication, its cellular functions in higher eukaryotes have remained largely unexplored. Here, we provide evidence that human INO80 (hINO80) participates in both DNA replication and chromosome segregation during the normal cell division cycle. hINO80 binds to chromatin localizing at replication forks during the S-phase, and is required for efficient DNA synthesis and S-phase progression. Unexpectedly, hINO80 associates with spindle microtubule during mitosis, and its deficiency leads to defective microtubule assembly and abnormal chromosome segregation. Consistent with these results, hINO80 is critical for suppressing aneuploidy and structural chromosome abnormalities. This work therefore not only emphasizes the evolutionary importance of INO80 in DNA replication but also reveals a new role for this remodeler in chromosome segregation, both of which likely come into play in maintaining the genome integrity.









Similar content being viewed by others
References
Aguilera A, Gómez-González B (2008) Genome instability: a mechanistic view of its causes and consequences. Nat Rev Genet 9:204–217
Draviam VM, Xie S, Sorger PK (2004) Chromosome segregation and genomic stability. Curr Opin Genet Dev 14:120–125
Groth A, Rocha W, Verreault A, Almouzni G (2007) Chromatin challenges during DNA replication and repair. Cell 128:721–733
Swedlow JR, Hirano T (2003) The making of the mitotic chromosome: modern insights into classical questions. Mol Cell 11:557–569
Flaus A, Martin DM, Barton GJ, Owen-Hughes T (2006) Identification of multiple distinct Snf2 subfamilies with conserved structural motifs. Nucleic Acids Res 34:2887–2905
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705
Smith CL, Peterson CL (2005) ATP-dependent chromatin remodeling. Curr Top Dev Biol 65:115–148
Shen X, Mizuguchi G, Hamiche A, Wu C (2000) A chromatin remodelling complex involved in transcription and DNA processing. Nature 406:541–544
Jin J, Cai Y, Yao T, Gottschalk AJ, Florens L, Swanson SK, Gutiérrez JL, Coleman MK, Workman JL, Mushegian A, Washburn MP, Conaway RC, Conaway JW (2005) A mammalian chromatin remodeling complex with similarities to the yeast INO80 complex. J Biol Chem 280:41207–41212
Shen X, Xiao H, Ranallo R, Wu WH, Wu C (2003) Modulation of ATP-dependent chromatin-remodeling complexes by inositol polyphosphates. Science 299:112–114
Bao Y, Shen X (2007) INO80 subfamily of chromatin remodeling complexes. Mutat Res 618:18–29
Conaway RC, Conaway JW (2009) The INO80 chromatin remodeling complex in transcription, replication and repair. Trends Biochem Sci 34:71–77
Morrison AJ, Shen X (2009) Chromatin remodelling beyond transcription: the INO80 and SWR1 complexes. Nat Rev Mol Cell Biol 10:373–384
Mizuguchi G, Shen X, Landry J, Wu WH, Sen S, Wu C (2004) ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 303:343–348
van Attikum H, Fritsch O, Hohn B, Gasser SM (2004) Recruitment of the INO80 complex by H2A phosphorylation links ATP-dependent chromatin remodeling with DNA double-strand break repair. Cell 119:777–788
Morrison AJ, Highland J, Krogan NJ, Arbel-Eden A, Greenblatt JF, Haber JE, Shen X (2004) INO80 and γ-H2AX interaction links ATP-dependent chromatin remodeling to DNA damage repair. Cell 119:767–775
Downs JA, Allard S, Jobin-Robitaille O, Javaheri A, Auger A, Bouchard N, Kron SJ, Jackson SP, Cote J (2004) Binding of chromatin-modifying activities to phosphorylated histone H2A at DNA damage sites. Mol Cell 16:979–990
Tsukuda T, Fleming AB, Nickoloff JA, Osley MA (2005) Chromatin remodelling at a DNA double-strand break site in Saccharomyces cerevisiae. Nature 438:379–383
Tsukuda T, Lo YC, Krishna S, Sterk R, Osley MA, Nickoloff JA (2009) INO80-dependent chromatin remodeling regulates early and late stages of mitotic homologous recombination. DNA Repair (Amst) 8:360–369
Papamichos-Chronakis M, Krebs JE, Peterson CL (2006) Interplay between Ino80 and Swr1 chromatin remodeling enzymes regulates cell cycle checkpoint adaptation in response to DNA damage. Genes Dev 20:2437–2449
Morrison AJ, Kim JA, Person MD, Highland J, Xiao J, Wehr TS, Hensley S, Bao Y, Shen J, Collins SR, Weissman JS, Delrow J, Krogan NJ, Haber JE, Shen X (2007) Mec1/Tel1 phosphorylation of the INO80 chromatin remodeling complex influences DNA damage checkpoint responses. Cell 130:499–511
Papamichos-Chronakis M, Peterson CL (2008) The Ino80 chromatin-remodeling enzyme regulates replisome function and stability. Nat Struct Mol Biol 15:338–345
Shimada K, Oma Y, Schleker T, Kugou K, Ohta K, Harata M, Gasser SM (2008) Ino80 chromatin remodeling complex promotes recovery of stalled replication forks. Curr Biol 18:566–575
Vincent JA, Kwong TJ, Tsukiyama T (2008) ATP-dependent chromatin remodeling shapes the DNA replication landscape. Nat Struct Mol Biol 15:477–484
Park JH, Park EJ, Hur SK, Kim S, Kwon J (2009) Mammalian SWI/SNF chromatin remodeling complexes are required to prevent apoptosis after DNA damage. DNA Repair (Amst) 8:29–39
Park JH, Park EJ, Lee HS, Kim SJ, Hur SK, Imbalzano AN, Kwon J (2006) Mammalian SWI/SNF complexes facilitate DNA double-strand break repair by promoting gamma-H2AX induction. EMBO J 25:3986–3997
Shaffer LG, Tommerrup N (2005) An international system for human cytogenetic nomenclature. S. Karger Publishers, Inc., Basel
Falbo KB, Alabert C, Katou Y, Wu S, Han J, Wehr T, Xiao J, He X, Zhang Z, Shi Y, Shirahige K, Pasero P, Shen X (2009) Involvement of a chromatin remodeling complex in damage tolerance during DNA replication. Nat Struct Mol Biol 16:1167–1172
Kops GJ, Weaver BA, Cleveland DW (2005) On the road to cancer: aneuploidy and the mitotic checkpoint. Nat Rev Cancer 5:773–785
Glotzer M (2005) The molecular requirements for cytokinesis. Science 307:1735–1739
Ogiwara H, Ui A, Kawashima S, Kugou K, Onoda F, Iwahashi H, Harata M, Ohta K, Enomoto T, Seki M (2007) Actin-related protein Arp4 functions in kinetochore assembly. Nucleic Acids Res 35:3109–3117
Ducat D, Kawaguchi S, Liu H, Yates JR 3rd, Zheng Y (2008) Regulation of microtubule assembly and organization in mitosis by the AAA+ ATPase Pontin. Mol Biol Cell 19:3097–3110
Murnane JP (2006) Telomeres and chromosome instability. DNA Repair (Amst) 5:1082–1092
Wu S, Shi Y, Mulligan P, Gay F, Landry J, Liu H, Lu J, Qi HH, Wang W, Nickoloff JA, Wu C, Shi Y (2007) A YY1-INO80 complex regulates genomic stability through homologous recombination-based repair. Nat Struct Mol Biol 14:1165–1172
Acknowledgments
We thank Jae-Ho Lee (Ajou University, Korea), Dae-Sik Lim (KAIST, Korea), and the Kazusa DNA Research Institute (Japan) for kindly providing GFP-H2B HeLa cells, pYFP-α-tubulin, and the full-length hINO80 cDNA clone, respectively. This work was supported by the Molecular and Cellular BioDiscovery Research Program (M10748000334-08N4800-33410) grant to J.K. from the Korea Science and Engineering Foundation (KOSEF) funded by the Korea Ministry of Education, Science and Technology (MEST), and also supported in part by the grant to J.K. (R01-2007-000-10571-0) from KOSEF funded by MEST, and by grant No. R15-2006-020 from the National Core Research Center (NCRC) program of MEST and KOSEF through the Center for Cell Signaling and Drug Discovery Research at Ewha Womans University.
Author information
Authors and Affiliations
Corresponding author
Additional information
S.-K. Hur and E.-J. Park contributed equally to this work.
Rights and permissions
About this article
Cite this article
Hur, SK., Park, EJ., Han, JE. et al. Roles of human INO80 chromatin remodeling enzyme in DNA replication and chromosome segregation suppress genome instability. Cell. Mol. Life Sci. 67, 2283–2296 (2010). https://doi.org/10.1007/s00018-010-0337-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-010-0337-3