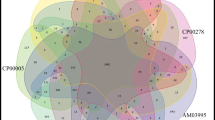
A detection method specific for Xanthomonas oryzae pv. oryzae, the pathogen responsible for bacterial blight of rice, was based on the polymerase chain reaction (PCR) and designed by amplifying the 16S–23S rDNA spacer region from this bacterium. The nucleotide sequence of the spacer region between the 16S and 23S rDNA, consisting of approximately 580-bp, from X. oryzae pv. oryzae, X. campestris pv. alfalfae, X. campestris pv. campestris, X. campestris pv. cannabis, X. campestris pv. citri, X. campestris pv. cucurbitae, X. campestris pv. pisi, X. campestris pv. pruni and X. campestris pv. vitians, was determined. The determined sequences had more than 95% identity. Therefore, a pair of primers, XOR-F (5′-GCATGACGTCATCGTCCTGT-3′) and XOR-R2 (5′-CTCGGAGCTATATGCCGTGC-3′) was designed and found to specifically amplify a 470-bp fragment from all strains of X. oryzae pv. oryzae isolated from diverse regions in Japan. No PCR product was amplified from X. campestris pathovars alfalfae, campestris, cannabis, carotae, cucurbitae, dieffenbachiae, glycines, pisi, pruni, vitians or zantedeschiae, except for pathovars citri, incanae and zinniae. The method could also detect the pathogen in infected rice leaves within 3 hr, at a detection limit of 4×101 cfu/ml.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received 17 December 1999/ Accepted in revised form 10 April 2000
Rights and permissions
About this article
Cite this article
ADACHI, N., OKU, T. PCR-mediated Detection of Xanthomonas oryzae pv. oryzae by Amplification of the 16S–23S rDNA Spacer Region Sequence. J Gen Plant Pathol 66, 303–309 (2000). https://doi.org/10.1007/PL00012969
Issue Date:
DOI: https://doi.org/10.1007/PL00012969