Abstract
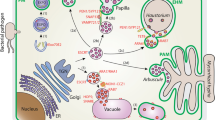
Autophagy is a highly conserved processing mechanism in eukaryotes whereby cytoplasmic components are engulfed in double-membrane vesicles called autophagosomes and are delivered into organelles such as lysosomes (mammal) or vacuoles (yeast/plant) for degradation and recycling of the resulting molecules. Isolation of yeastAUTOPHAGY (ATG) genes has facilitated the identification of correspondingArabidopsis ATG genes based on sequence similarity. Genetic and molecular analyses using knockout and/or knockdown mutants of those genes have unraveled the biological functions of autophagy during plant development, nutrient recycling, and environmental stress responses. Additional roles for autophagy have been suggested in the degradation of oxidized proteins during oxidative stress and the regulation of hypersensitive response (HR)-programmed cell death (PCD) during innate immunity. Our review summarizes knowledge about the structure and function of autophagic pathways andATG components, and the biological roles of autophagy in plants.
Similar content being viewed by others
Literature cited
Aubert S, Gout E, Bligny R, MartyMazars D, Barrieu F, Alabouvette J, Marty F, Douce R (1996) Ultrastructural and biochemical characterization of autophagy in higher plant cells subjected to carbon deprivation: control by the supply of mitochondria with respiratory substrates. J Cell Biol 133: 1251–1263
Baehrecke EH (2005) Autophagy: dual roles in life and death? Nat Rev Mol Cell 6: 505–510
Bassham DC (2007) Plant autophagy-more than a starvation response. Curr Opin Plant Biol 10: 587–593
Bassham DC, Laporte M, Marty F, Moriyasu Y, Ohsumi Y, Olsen LJ, Yoshimoto K (2006) Autophagy in development and stress responses of plants. Autophagy 2:2–11
Bursch W (2001) The autophagosomal-lysosomal compartment in programmed cell death. Cell Death Differ 8: 569–581
Chen MH, Liu LF, Chen YR, Wu HK, Yu SM (1994) Expression of áamylases, carbohydrate metabolism, and autophagy in cultured rice cells is coordinately regulated by sugar nutrient. Plant J 6: 625–636
Contento AL, Kim SJ, Bassham DC (2004) Transcription profiling of the response ofArabidopsis suspension culture cells to Suc starvation. Plant Physiol 135: 2330–2347
Contento AL, XiongY, Bassham DC (2005) Visualization of autophagy inArabidopsis using the fluorescent dye monodansylca-daverine and a GFP-AtATG8e fusion protein. Plant J 42: 598–608
Dangl JL, Jones JDG (2001) Plant pathogens and integrated defence responses to infection. Nature 411: 826–833
DoellingJH, Walker JM, Friedman EM, Thompson AR, Vierstra RD (2002) The APG8/12-activating enzyme APG7 is required for proper nutrient recycling and senescence inArabidopsis thaliana. J Biol Chem 277: 33105–33114
Ellis C, Turner JG, Devoto A (2002) Protein complexes mediate signaling in plant responses to hormones, light, sucrose, and pathogens. Plant Mol Biol 50: 971–980
Fujiki Y, Yoshimoto K, Ohsumi Y (2007) AnArabidopsis homolog of yeast ATG6/VPS30 is essential for pollen germination. Plant Physiol 143: 1132–1139
Fujioka Y, Noda NN, Fujii K, Yoshimoto K, Ohsumi Y, Inagaki F (2008) In vitro reconstitution of plant Atg8 and Atg12 conjugation systems essential for autophagy. J Biol Chem 283: 1921–1928
Fukuda H (2004) Signals that control plant vascular cell differentiation. Nat Rev Mol Cell Biol 5: 379–391
Gozuacik D, Kimchi A (2007) Autophagy and cell death. Curr Top Dev Biol 78: 217–245
Hanada T, Noda NN, Satomi Y, Ichimura Y, Fujioka Y, Takao T, Inagaki F, Ohsumi Y, (2007) The Atg12-Atg5 conjugate has a novel E3-like activity for protein lipidation in autophagy. J Biol Chem 282: 37298–37302
Hanaoka H, Noda T, Shirano Y, Kato T, Hayashi H, Shibata D, Tabata S, Ohsumi Y (2002) Leaf senescence and starvation-induced chlorosis are accelerated by the disruption of anArabidopsis autophagy gene. Plant Physiol 129: 1181 -1193
Harding TM, Morano KA, Scott SV, Klionsky DJ (1995) Isolation and characterization of yeast mutants in the cytoplasm to vacuole protein targeting pathway. J Cell Biol 131: 591–602
Harding TM, Hefner-Gravink A, Thumm M, Klionsky DJ (1996) Genetic and phenotypic overlap between autophagy and the cytoplasm to vacuole protein targeting pathway. J Biol Chem 271: 17621–17624
Harrison-Lowe NJ, Olsen LJ (2008) Autophagy protein 6 (ATG6) is required for pollen germination inArabidopsis thaliana. Autophagy 4: 339–348
Ichimura Y, Kirisako T, Takao T, Satomi Y, Shimonishi Y, Ishihara N, Mizushima N, Tanida I, Kominami E, Ohsumi M, Noda T, Ohsumi Y (2000) A ubiquitin-like system mediates protein lipidation. Nature 408: 488–492
Inoue Y, Suzuki T, Hattori M, Yoshimoto K, Ohsumi Y, Moriyasu Y (2006) AtATG genes, homologs of yeast autophagy genes, are involved in constitutive autophagy inArabidopsis root tip cells. Plant Cell Physiol 47: 1641–1652
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444: 323–329
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T, Kominami E, Ohsumi Y, Yoshimori T (2000) LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 19: 5720–5728
Kamada Y, Funakoshi T, Shintani T, Nagano K, Ohsumi M, Ohsumi Y (2000) Tor-mediated induction of autophagy via an Apg1 protein kinase complex. J Cell Biol 150: 1507–1513
Kang C, You Y, Avery L (2007) Dual roles of autophagy in the survival ofCaenorhabditis elegans during starvation. Genes Dev 21: 2161–2171
Kirisako T, Baba M, Ishihara N, Miyazawa K, Ohsumi M, Yoshimori T, Noda T, Ohsumi Y (1999) Formation process of autophagosome is traced with Apg8/Aut7p in yeast. J Cell Biol 147: 435–446
Klionsky DJ (2005) The molecular machinery of autophagy: Unanswered questions. J Cell Sci 118: 7–18
Klionsky DJ (2007) Autophagy; from phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol 8: 931–937
Klionsky DJ etal. (2008) Guidelines for the use and interpretation of assays for monitoring autophagy in higher eukaryotes. Autophagy 4: 151–175
KumaA, Mizushima N, Ishihara N, Ohsumi Y (2002) Formation of the approximately 350-kDa Apg12-Apg5.Apg16 multimeric complex, mediated by Apg16 oligomerization, is essential for autophagy in yeast. J Biol Chem 277: 18619–18625
Lamb E, Kato N, Lawton M (2001) Programmed cell death, mitochondria and the plant hypersensitive response. Nature 411: 848–853
Levine B, Klionsky DJ (2004) Development by self-digestion: Molecular mechanisms and biological functions of autophagy. Dev Cell 6: 463–477
Levine B, Kroemer G (2008) Autophagy in the pathogenesis of disease. Cell 132: 27–42
Levine B, Yuan J (2005) Autophagy in cell death: An innocent convict? J Clin Invest 115: 2679–2688
Liu Y, Schiff M, Czymmek K, Talloczy Z, Levine B, Dinesh-Kumar SP (2005) Autophagy regulates programmed cell death during the plant innate immune response. Cell 121: 567–577
Maiuri MC, Zalckvar E, Kimchi A, Kroemer G (2007) Self-eating and self-killing: crosstalk between autophagy and apoptosis. Nat Rev Mol Cell Biol 8: 741–752
Massey AC, Zhang C, Cuervo AM (2006) Chaperone-mediated autophagy in aging and disease. Curr Top Dev Biol 73: 205–235
Menand B, Desnos T, Nussaume L, Berger F, Bouchez D, Meyer C, Robaglia C (2002) Expression and disruption of theArabidopsis TOR (target of rapamycin) gene. Proc Natl Acad Sci USA 99: 6422–6427
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F. (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9: 490–498
Mizushima N (2007) Autophagy: process and function. Genes Dev 21:2861–2873
Mizushima N, Levine B, Cuervo AM, Klionsky DJ (2008) Autophagy fights disease through cellular self-digestion. Nature 451: 1069–1075
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD, Klionsky DJ, Ohsumi M, Ohsumi Y (1998) A protein conjugation system essential for autophagy. Nature 395: 395–398
Moriyasu Y, Hattori M, Jauh GY, Rogers JC (2003) Alpha tonoplast intrinsic protein is specifically associated with vacuole membrane involved in an autophagic process. Plant Cell Physiol 44: 795–802
Moriyasu Y, Ohsumi Y (1996) Autophagy in tobacco suspension-cultured cells in response to sucrose starvation. Plant Physiol 111: 1233–1241
Mukaiyama H, Oku M, Baba M, Samizo T, Hammond AT, Glick BS, Kato N, Sakai Y (2002) Paz2 and 13 otherPAZ gene products regulate vacuolar engulfment of peroxisomes during micropexophagy. Genes Cells 7: 75–90
Nair U, Klionsky DJ (2005) Molecular mechanisms and regulation of specific and nonspecific autophagy pathways in yeast. J Biol Chem 280: 41785–41788
Nakatogawa H, Ichimura Y, Ohsumi Y (2007) Atg8, a Ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell 130: 165–178
Ogawa M, Sasakawa C (2006) Bacterial evasion of the autophagic defense system. Curr Opin Microbiol 9: 62–68
Ohsumi Y (2001) Molecular dissection of autophagy: Two ubiquitin-like systems. Nat Rev Mol Cell Biol 2: 211–216
Patel S, Caplan J, Dinesh-Kumar SP (2006) Autophagy in the control of programmed cell death. Curr Opin Plant Biol 9: 391–396
Patel S, Dinesh-Kumar SP (2008)Arabidopsis ATG6 is required to limit the pathogen-associated cell death response. Autophagy 4: 20–27
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N, Packer M, Schneider MD, Levine B (2005) Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 122: 927–939
Pennell RI, Lamb C (1997) Programmed cell death. Plant Cell 9: 1157–1168
Phillips BA, Suttangkakul A, Vierstra RD (2008) The ATG12 conjugating enzyme ATG10 is essential for autophagic vesicle formation inArabidopsis thaliana. Genetics 178: 1339–1353
Qin G, Ma Z, Zhang L, Xing S, Hou X, Deng J, Liu J, Chen Z, Qu LJ, Gu H (2007)Arabidopsis AtBECLIN 1/AtAtg6/AtVps30 is essential for pollen germination and plant development. Cell Res 17: 249–263
Rose TL, Bonneau L, Der C, Marty-Mazars D, Marty F (2006) Starvation-induced expression of autophagy-related genes inArabidopsis. Biol Cell 98: 53–67
Sakai Y, Koller A, Rangell LK, Keller GA, Subramani S (1998) Peroxisome degradation by microautophagy inPichia pastoris: identification of specific steps and morphological intermediates. J Cell Biol 141: 625–636
Sanmartin M, Ordonez A, Sohn EJ, Robert S, Sanchez-Serrano JJ, Surpin MA, Raikhel NV, Rojo E (2007) Divergent functions of VTI12 and VTI11 in trafficking to storage and lytic vacuoles inArabidopsis. Proc Natl Acad Sci USA 104: 3645–3650
Scherz-Shouval R, Elazar Z (2007) ROS, mitochondria and the regulation of autophagy. Trends Cell Biol 17: 422–427
Scherz-Shouval R, Shvets E, Fass E, Shorer H, Gil L, Elazar Z (2007) Reactive oxygen species are essential for autophagy and specifically regulate the activity of Atg4. EMBO J 26: 1749–1760
Schmelzle T, Hall MN (2000) TOR, a central controller of cell growth. Cell 103: 253–262
Scott RC, Juhasz G, Neufeld TP (2007) Direct induction of autophagy by Atg1 inhibits cell growth and induces apoptotic cell death. Curr Biol 17: 1–11
Seay MD, Dinesh-Kumar SP (2005) Life after death: are autophagy genes involved in cell death and survival during plant innate immune responses? Autophagy 1: 185–186
Seay MD, Patel S, Dinesh-Kumar SP (2006) Autophagy and plant innate immunity. Cellular Microbiology 8: 899–906
Slavikova S, Shy G, Yao YL, Giozman R, Levanony H, Pietrokovski S, Elazar Z, Galili G (2005) The autophagy-associated Atg8 gene family operates both under favourable growth conditions and under starvation stresses inArabidopsis plants. J Exp Bot 56: 2839–2849
Smalle J, Vierstra RD (2004) The ubiquitin 26S proteasome proteolytic pathway. Annu Rev Plant Biol 55: 555–590
Su W, Ma H, Liu C, Wu J, Yang J (2006) Identification and characterization of two rice autophagy associated genes, OsAtg8 and OsAtg4. Mol Biol Rep 33: 273–278
Surpin M, Zheng H, Moria MT, Saito C, Avila E, Blakeslee JJ, Bandyopadhyay A, Kovaleva V, Carter D, Murphy A, Tasaka M, Raikhel N (2003) The VTI family of SNARE proteins is nessary for plant viability and mediates different protein transport pathways. Plant Cell 15: 2885–2899
Suzuki NN, Yoshimoto K, Fujioka Y, Ohsumi Y, Inagaki F (2005) The crystal structure of plant ATG12 and its biological implication in autophagy. Autophagy 1: 119–126
Takatsuka C, Inoue Y, Matsuoka K, Moriyasu Y (2004) 3-Methyladenine inhibits autophagy in tobacco culture cells under sucrose starvation conditions. Plant Cell Physiol 45: 265–274
Thompson AR, Doelling JH, Suttangkakul A, Vierstra RD (2005) Autophagic nutrient recycling inArabidopsis directed by the ATG8 and ATG12 conjugation pathways. Plant Physiol 138: 2097–2110
Thompson AR, Vierstra RD (2005) Autophagic recycling: lessons from yeast help define the process in plants. Curr Opin Plant Biol 8: 165–173
Thumm M, Egner R, Koch M, Schlumpberger M, Straub M, Veenhuis M, Wolf DH (1994) Isolation of autophagocytosis mutants ofSaccharomyces cerevisiae. FEBS Lett 349: 275–280
Titorenko VI, Keizer I, Harder W, Veenhuis M (1995) Isolation and characterization of mutants impaired in the selective degradation of peroxisomes in the yeastHansenula polymorpha. J Bacteriol 177: 357–363
Toyooka K, Moriyasu Y, Goto Y, Takeuchi M, Fukuda H, Matsuoka K (2006) Protein aggregates are transported to vacuoles by a macroautophagic mechanism in nutrient-starved plant cells. Autophagy 2: 96–106
Toyooka K, Okamoto T, Minamikawa T (2001) Cotyledon cells ofVigna mungo seedlings use at least two distinct autophagic machineries for degradation of starch granules and cellular components. J Cell Biol 154: 973–982
Tsukada M, Ohsumi Y (1993) Isolation and characterization of autophagy-defective mutants ofSaccharomyces cerevisiae. FEBS Lett 333: 169–174
Van der Wilden W, Herman EM, Chrispeels MJ (1980) Protein bodies of mung bean cotyledons as autophagic organelles. Proc Natl Acad Sci USA 77: 428–432
Van Doorn WG, Woltering EJ (2005) Many ways to exit? Cell death categories in plants. Trends Plant Sci 10: 117–122
Vierstra RD (2003) The ubiquitin/26S proteasome pathway, the complex last chapter in the life of many plant proteins. Trends Plant Sci 8: 135–142
Wang CW, Kim J, Huang WP, Abeliovich H, Stromhaug PE, Dunn WA, Klionsky DJ (2001) Apg2 is a novel protein required for the cytoplasm to vacuole targeting, autophagy, and pexophagy. J Biol Chem 276: 30442–30451
Xiong Y, Contento AL, Bassham DC (2005) AtATG18a is required for the formation of autophagosomes during nutrient stress and senescence inArabidopsis thaliana. Plant J 42: 535–546
Xiong Y, Contento AL, Bassham DC (2007a) Disruption of autophagy results in constitutive oxidative stress in Arabidopsis. Autophagy 3: 257–258
Xiong Y, Contento AL, Nguyen PQ, Bassham DC (2007b) Degradation of oxidized proteins by autophagy during oxidative stress in Arabidopsis. Plant Physiol 143: 291–299
Yano K, Suzuki T, Moriyasu Y (2007) Constitutive autophagy in plant root cells. Autophagy 3: 360–362
Yoshimori T (2007) Autophagy: Paying Charon’s toll. Cell 128: 833–836
Yoshimoto K, Hanaoka H, Sato S, Kato T, Tabata S, Noda T, Ohsumi Y (2004) Processing of ATG8s, ubiquitin-like proteins, and their deconjugation by ATG4s are essential for plant autophagy. Plant Cell 16: 2967–2983
Yuan W, Tuttle DL, Shi YJ, Ralph GS, Dunn Jr. WA (1997) Glucose-induced microautophagy inPichia pastoris requires the α-sub- unit of phosphofructokinase. J Cell Sci 110: 1935–1945
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kwon, S.I., Park, O.K. Autophagy in plants. J. Plant Biol. 51, 313–320 (2008). https://doi.org/10.1007/BF03036132
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF03036132